![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_B_1_CGATGT_L008_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 24596630 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
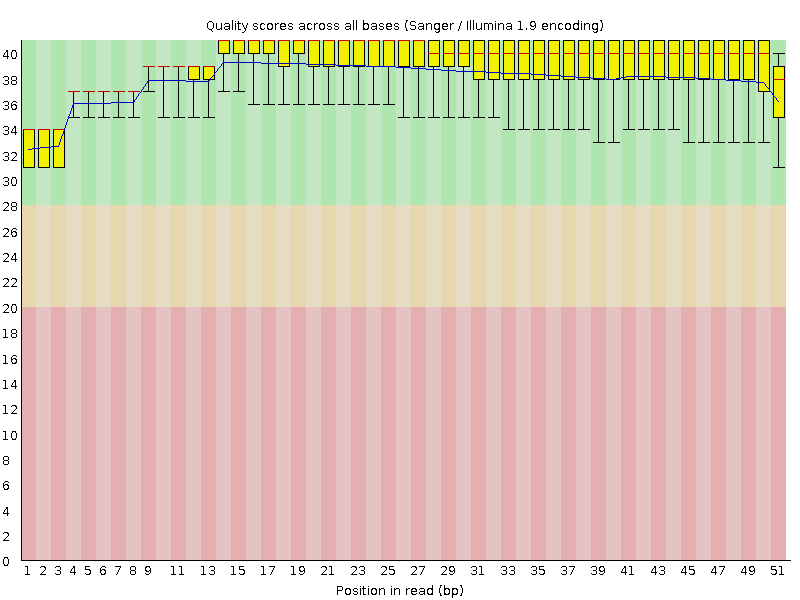
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
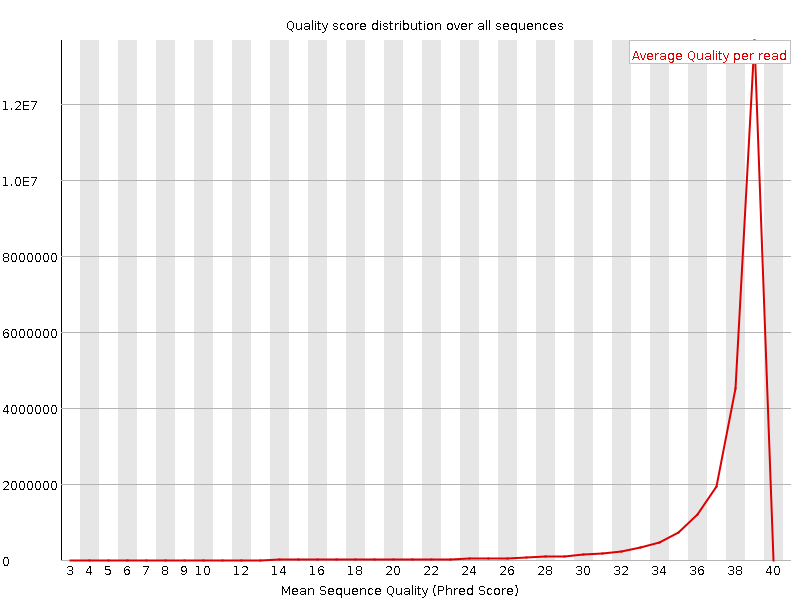
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
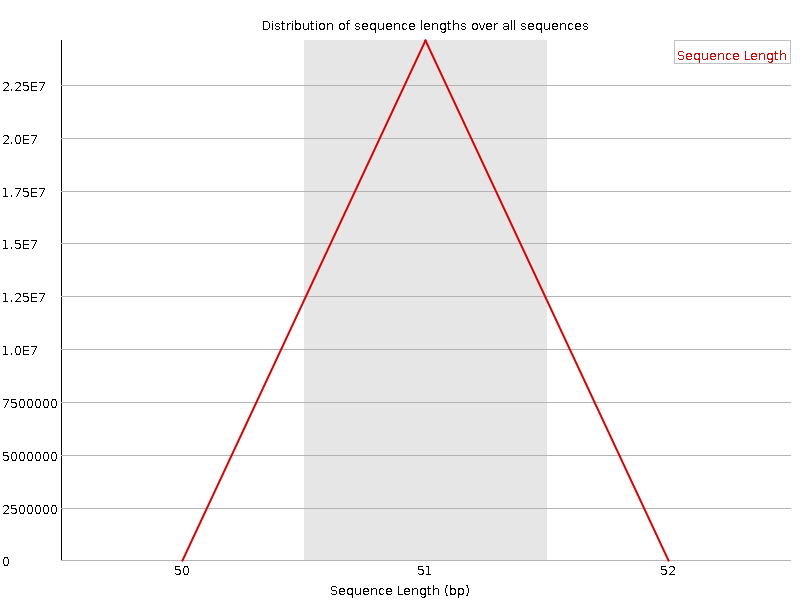
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
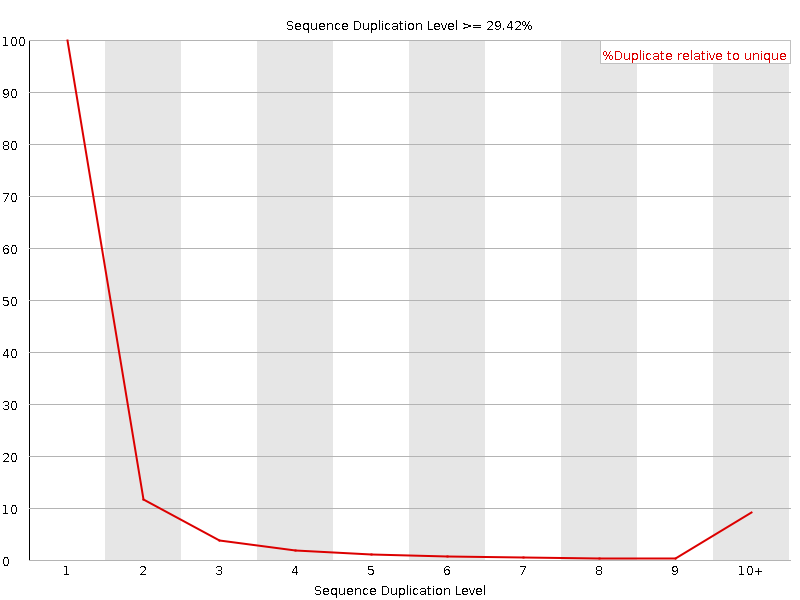
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 50793 | 0.20650389911138234 | No Hit |
| TTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 33711 | 0.13705536083601697 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
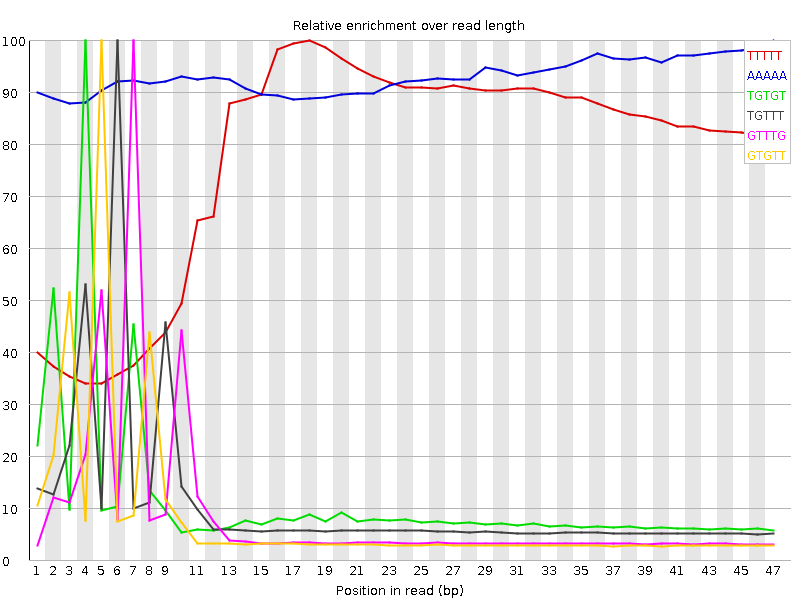
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 13129140 | 5.8676686 | 7.5431147 | 18 |
| AAAAA | 6192525 | 3.6667833 | 3.9353468 | 47 |
| TGTGT | 5061035 | 3.589233 | 31.303028 | 4 |
| TGTTT | 4772030 | 2.6865752 | 25.047045 | 6 |
| GTTTG | 3746125 | 2.6567125 | 30.732166 | 7 |
| GTGTT | 3542175 | 2.5120733 | 30.793987 | 5 |
| TTTGA | 4111255 | 2.4485438 | 26.03405 | 8 |
| CTTTG | 3230485 | 2.4281723 | 32.84628 | 1 |
| TTGTG | 3314090 | 2.350318 | 30.75568 | 3 |
| GAGGG | 1842755 | 2.193818 | 9.268312 | 11 |
| TTTGT | 3710975 | 2.0892184 | 24.819674 | 2 |
| TTGAG | 2577115 | 1.9334536 | 18.993214 | 9 |
| TGAGG | 2028800 | 1.9173691 | 11.5638485 | 10 |
| AGGGG | 1374240 | 1.6360463 | 5.179037 | 12 |
| ACTTT | 2386620 | 1.5064898 | 12.077374 | 3 |
| AACTT | 2238585 | 1.494838 | 13.118069 | 2 |
| TGAGT | 1582360 | 1.1871492 | 5.38544 | 10 |
| TTGAT | 1874615 | 1.1164662 | 6.5887246 | 9 |
| TAACT | 1636365 | 1.0926993 | 12.869976 | 1 |
| TTGAC | 1191750 | 0.94762063 | 5.171499 | 9 |