![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_B_12_GGCTAC_L002_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 35734689 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 45 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
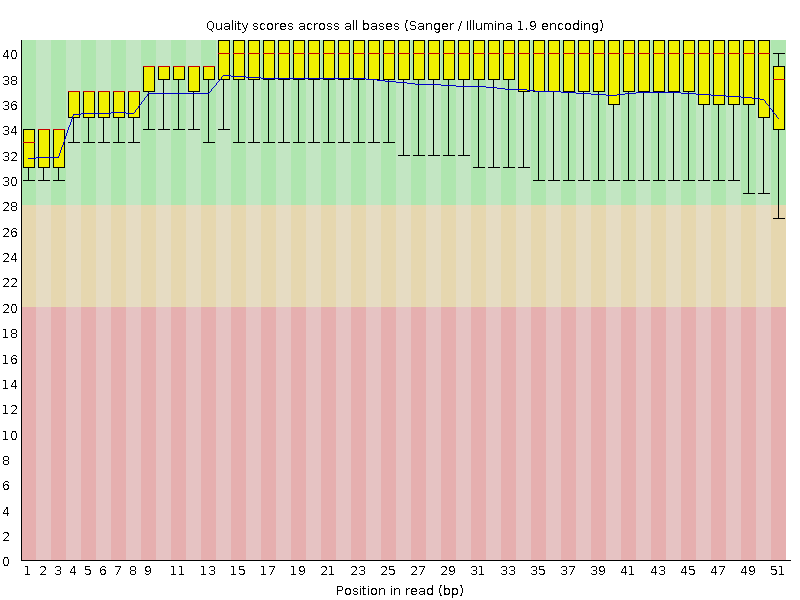
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
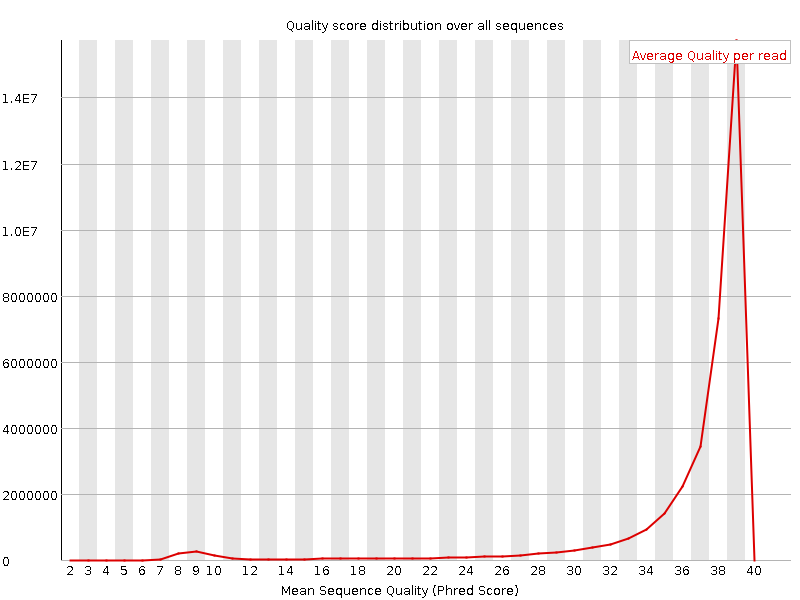
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[WARN]](Icons/warning.png) Per base GC content
Per base GC content

![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
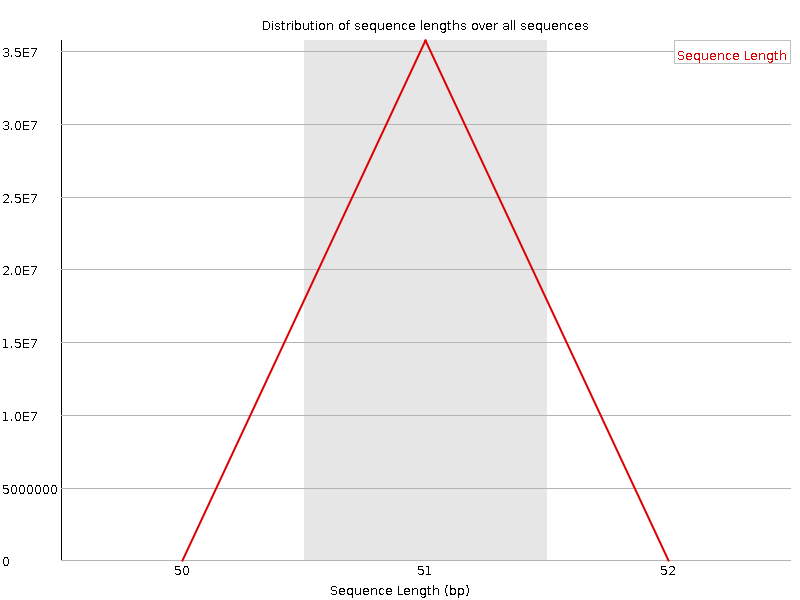
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
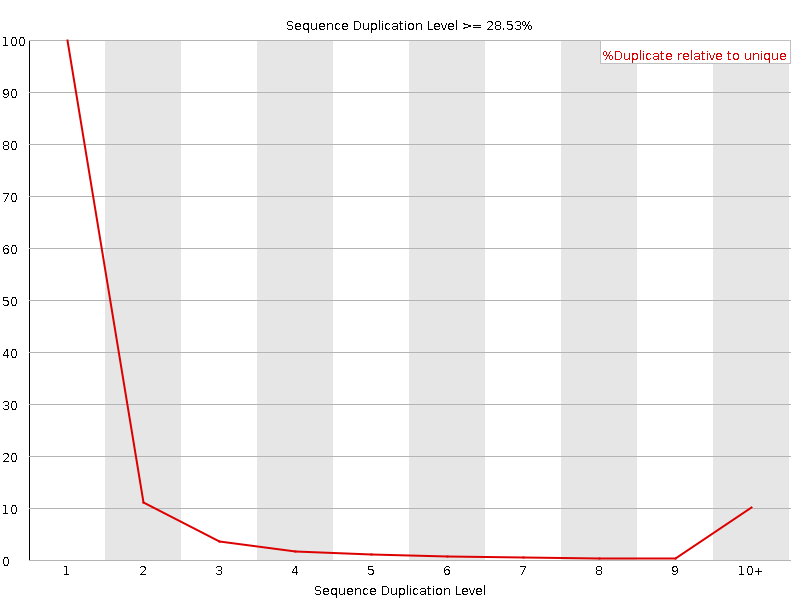
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 86327 | 0.2415775886562214 | No Hit |
| TTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 83891 | 0.23476068309983053 | No Hit |
| TAACTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 43661 | 0.12218099897273486 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
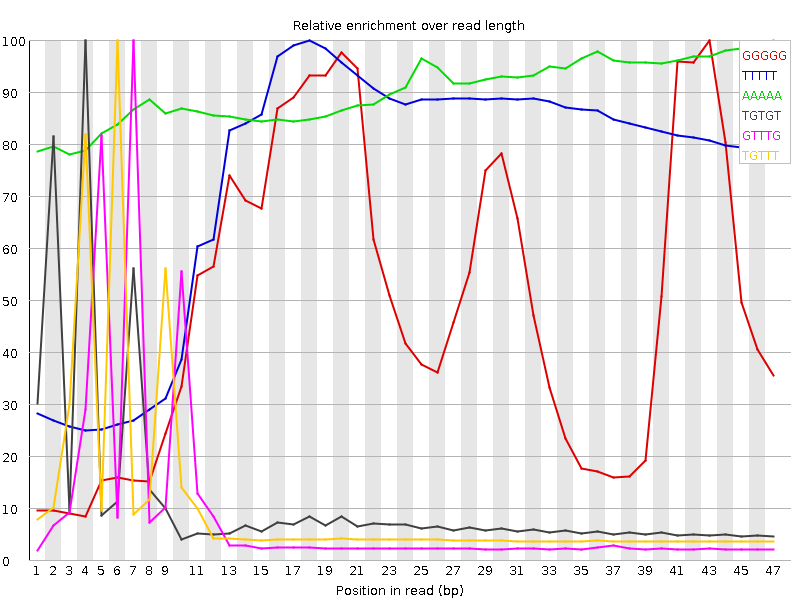
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GGGGG | 13133545 | 9.378768 | 18.957727 | 43 |
| TTTTT | 24621550 | 7.489593 | 10.153844 | 18 |
| AAAAA | 9251810 | 4.452207 | 4.933497 | 47 |
| TGTGT | 10402190 | 4.4515986 | 38.244446 | 4 |
| GTTTG | 7808530 | 3.341647 | 37.610886 | 7 |
| TGTTT | 9191235 | 3.3162014 | 32.22524 | 6 |
| TTTGA | 8244430 | 3.2603848 | 34.95876 | 8 |
| GTGTT | 7573140 | 3.240912 | 37.700966 | 5 |
| TTGTG | 6939395 | 2.969702 | 37.788467 | 3 |
| CTTTG | 5810215 | 2.8918464 | 44.121418 | 1 |
| GAGGG | 4050585 | 2.672998 | 12.207174 | 9 |
| TTTGT | 6579330 | 2.373825 | 32.130455 | 2 |
| TTGAG | 5022685 | 2.3559623 | 23.676779 | 9 |
| GGGGA | 3541565 | 2.3370934 | 8.619806 | 22 |
| AGGGG | 3504825 | 2.3128486 | 10.760539 | 10 |
| GGGAG | 3407825 | 2.248838 | 5.294396 | 23 |
| GGGGC | 2528425 | 2.0999343 | 8.501048 | 45 |
| TGAGG | 3553460 | 1.9770063 | 13.445593 | 10 |
| GGCCC | 1681040 | 1.8885009 | 5.016051 | 46 |
| ACTTT | 4087920 | 1.8801928 | 22.253134 | 3 |
| AACTT | 3655280 | 1.8427303 | 24.994726 | 2 |
| GGGCC | 1876730 | 1.8127972 | 6.4240713 | 45 |
| GGAGA | 2885210 | 1.7594428 | 5.0336328 | 6 |
| GGGAA | 2874335 | 1.7528113 | 5.8199253 | 22 |
| CGGGG | 1933395 | 1.6057439 | 7.159655 | 40 |
| TGAGC | 2232785 | 1.4447585 | 5.6245155 | 10 |
| TAACT | 2835440 | 1.4294257 | 24.772512 | 1 |
| GGCGG | 1563345 | 1.2984059 | 7.3870535 | 13 |
| GGGCG | 1561655 | 1.2970023 | 6.4429255 | 12 |
| TTGAT | 3223950 | 1.2749598 | 8.64355 | 9 |
| TGAGT | 2580135 | 1.2102494 | 6.546221 | 10 |
| GAGTG | 2105760 | 1.1715626 | 5.039163 | 11 |
| GCGGG | 1375270 | 1.1422039 | 6.6181574 | 14 |
| TTGAC | 1984330 | 1.0825254 | 7.0659914 | 9 |
| GTAAC | 1284085 | 0.7678194 | 6.1630454 | 1 |