![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_B_12_GGCTAC_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 35734689 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
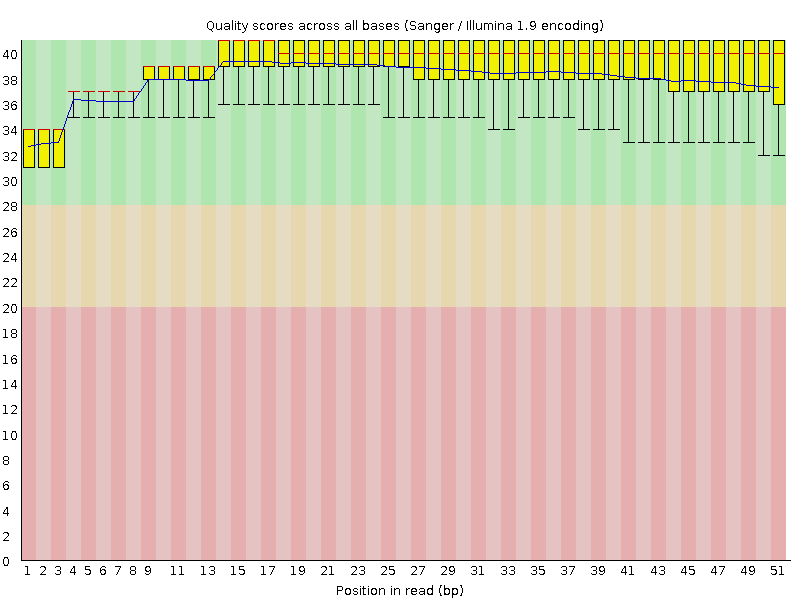
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
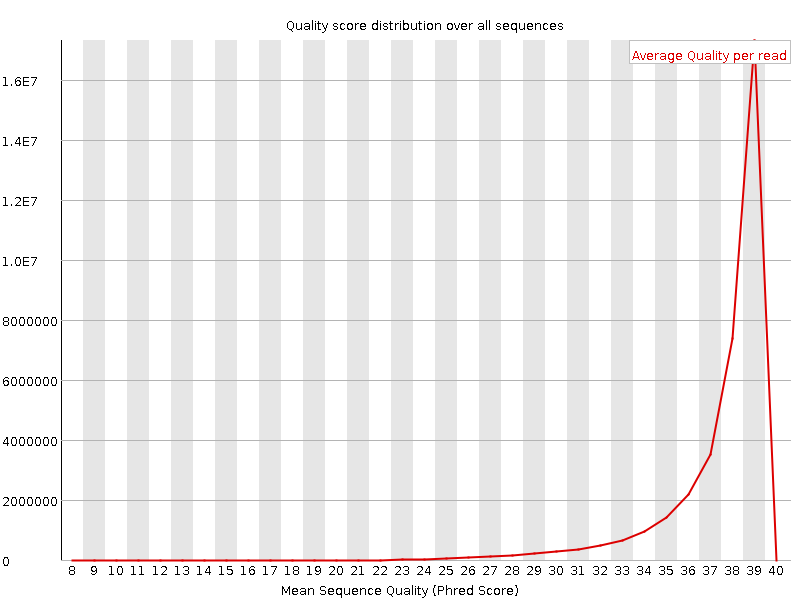
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
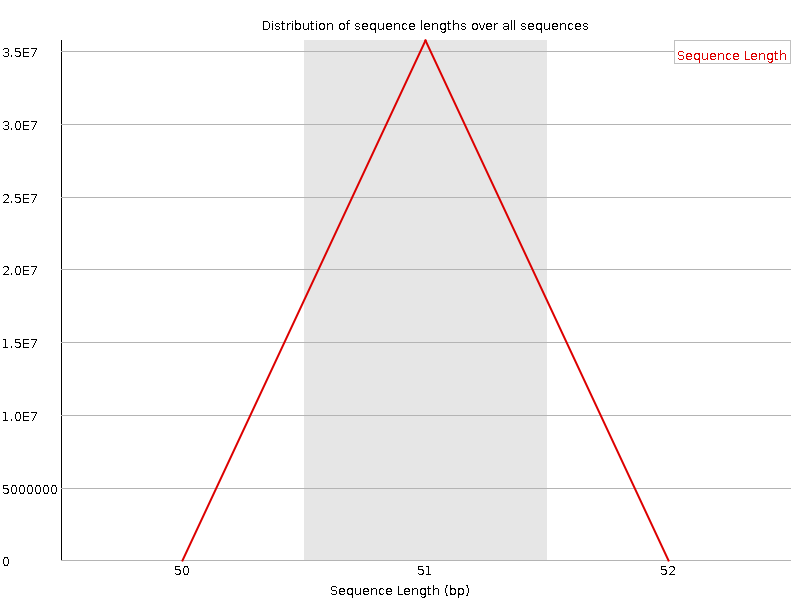
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
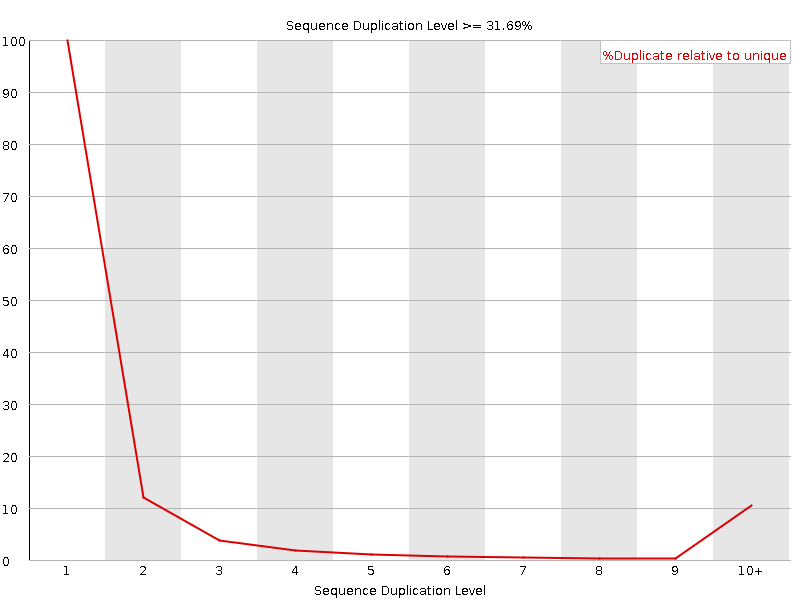
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACGGCTACATCTCGTATGCC | 651909 | 1.824302990296068 | TruSeq Adapter, Index 11 (100% over 51bp) |
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 104853 | 0.2934207710608591 | No Hit |
| AAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAA | 44424 | 0.12431617916137455 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
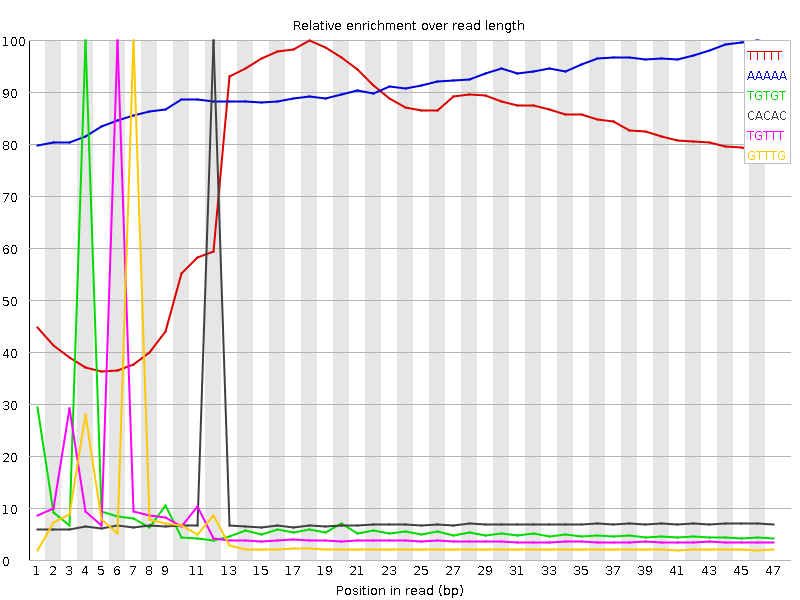
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 19504675 | 6.290242 | 8.193788 | 18 |
| AAAAA | 10176965 | 4.325102 | 4.7362037 | 46 |
| TGTGT | 7469810 | 3.7194426 | 45.929306 | 4 |
| CACAC | 3447980 | 2.7149303 | 30.742428 | 12 |
| TGTTT | 6742925 | 2.7020712 | 36.964066 | 6 |
| GTTTG | 5290515 | 2.6343062 | 45.186687 | 7 |
| CTTTG | 4848455 | 2.5217018 | 47.34745 | 1 |
| GTGTT | 5047255 | 2.5131798 | 45.17905 | 5 |
| TTTGA | 5834290 | 2.4706225 | 38.664074 | 8 |
| CTCCA | 3177330 | 2.3674805 | 28.522581 | 24 |
| TCCAG | 3305265 | 2.3578074 | 27.428463 | 25 |
| GGAAG | 3356595 | 2.3191395 | 27.20655 | 5 |
| TTTGT | 5560560 | 2.228266 | 36.76229 | 2 |
| TTGTG | 4460135 | 2.220835 | 45.22489 | 3 |
| GAAGA | 3676585 | 2.1603422 | 23.268906 | 6 |
| GAGGG | 2646375 | 2.1499546 | 11.977611 | 11 |
| GGGGG | 2177835 | 2.0804288 | 5.065935 | 13 |
| AAGAG | 3495755 | 2.0540876 | 23.078548 | 7 |
| AGAGC | 2841150 | 2.050426 | 27.910269 | 8 |
| GAGCA | 2735100 | 1.9738909 | 27.807446 | 9 |
| GCACA | 2519075 | 1.8989478 | 28.78609 | 11 |
| TTGAG | 3599330 | 1.8939123 | 26.705853 | 9 |
| TGAGG | 2877205 | 1.8811736 | 15.985786 | 10 |
| AGCAC | 2446825 | 1.8444837 | 28.760405 | 10 |
| CTGAA | 3151905 | 1.8306412 | 22.467346 | 19 |
| CCAGT | 2557995 | 1.8247433 | 26.893082 | 26 |
| TCTGA | 3147470 | 1.7299032 | 21.238577 | 18 |
| GTCTG | 2670645 | 1.7259425 | 24.717758 | 17 |
| CATCT | 2961355 | 1.7000929 | 20.777195 | 39 |
| CAGTC | 2314905 | 1.6513351 | 26.680534 | 27 |
| ATGCC | 2214350 | 1.5796043 | 25.592587 | 47 |
| ACTCC | 2110020 | 1.5722103 | 27.807863 | 23 |
| AGGGG | 1924580 | 1.5635575 | 6.6229444 | 12 |
| TGAAC | 2661975 | 1.5460875 | 22.108229 | 20 |
| AGTCA | 2655970 | 1.5425998 | 21.800657 | 28 |
| GTCAC | 2152410 | 1.5354195 | 26.442186 | 29 |
| GAACT | 2593710 | 1.5064389 | 22.079704 | 21 |
| AACTC | 2430640 | 1.474595 | 22.906443 | 22 |
| TGAGA | 2543190 | 1.4141221 | 5.089182 | 10 |
| TGAGC | 2052555 | 1.4017639 | 6.047561 | 10 |
| CTACA | 2295515 | 1.3926189 | 22.334406 | 36 |
| ACATC | 2273885 | 1.3794965 | 21.512093 | 38 |
| ATCTC | 2296595 | 1.3184589 | 20.218483 | 40 |
| GCTAC | 1840275 | 1.3127583 | 26.199495 | 35 |
| GGCTA | 1895305 | 1.2943722 | 25.142632 | 34 |
| CACGG | 1443860 | 1.2798144 | 32.15089 | 31 |
| CGGCT | 1502845 | 1.2605677 | 30.658264 | 33 |
| GAGTG | 1872890 | 1.2245325 | 5.669888 | 11 |
| ACGGC | 1355580 | 1.2015644 | 32.10364 | 32 |
| TGAGT | 2260460 | 1.1894196 | 7.3204317 | 10 |
| TACAT | 2456375 | 1.1481665 | 16.910025 | 37 |
| TTGAT | 2663405 | 1.1278609 | 10.030157 | 9 |
| GTATG | 2005785 | 1.0554134 | 18.950033 | 45 |
| CGGAA | 1461430 | 1.0546976 | 27.114248 | 4 |
| TGATT | 2400455 | 1.0165106 | 6.0003414 | 10 |
| ACACG | 1332775 | 1.0046823 | 27.887108 | 13 |
| TATGC | 1781415 | 0.9790961 | 19.37145 | 46 |
| CGTCT | 1415285 | 0.9553799 | 24.959507 | 16 |
| CTCGT | 1415115 | 0.95526516 | 23.619797 | 42 |
| CACGT | 1328555 | 0.94772345 | 26.330215 | 14 |
| TCTCG | 1375660 | 0.92863125 | 23.362553 | 41 |
| TTGAC | 1685520 | 0.9263906 | 7.5737143 | 9 |
| TCGGA | 1340385 | 0.91539735 | 25.608784 | 3 |
| TCACG | 1245000 | 0.8881196 | 25.710129 | 30 |
| GATCG | 1270045 | 0.8673595 | 25.569933 | 1 |
| ACGTC | 1148215 | 0.81907797 | 26.225441 | 15 |
| ATCGG | 1167900 | 0.79760104 | 25.517838 | 2 |
| TCGTA | 1157455 | 0.63615704 | 19.249704 | 43 |
| CGTAT | 1149925 | 0.6320184 | 19.225384 | 44 |