![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 140701_JD_B_10_GATCAG_L002_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 40487517 |
| Filtered Sequences | 0 |
| Sequence length | 51 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
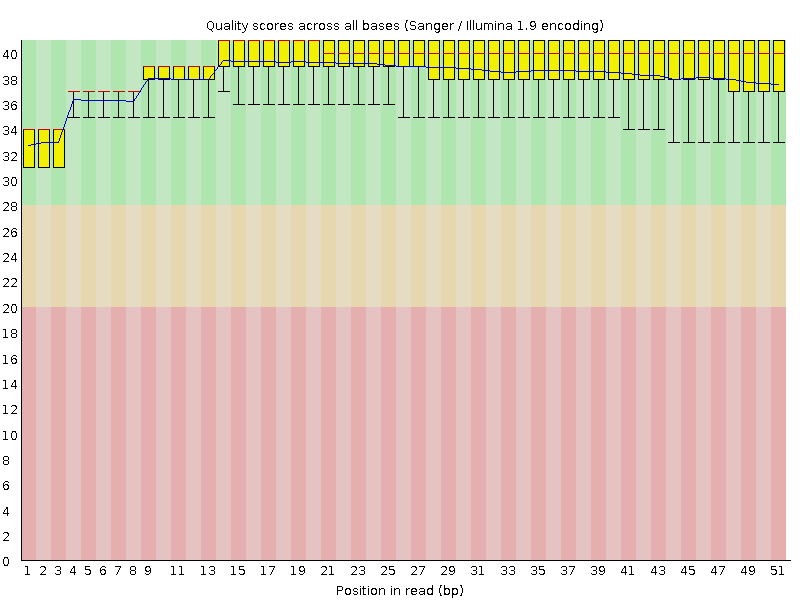
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
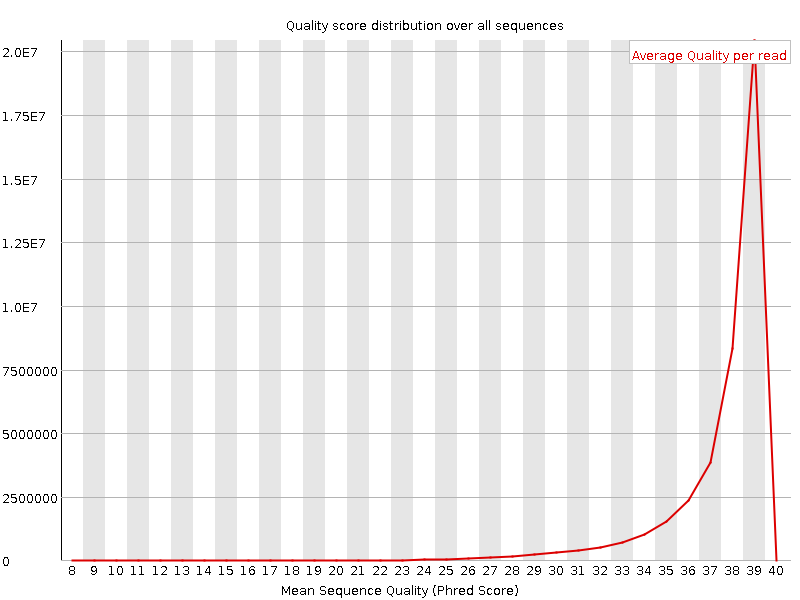
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content

![[OK]](Icons/tick.png) Per base GC content
Per base GC content

![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
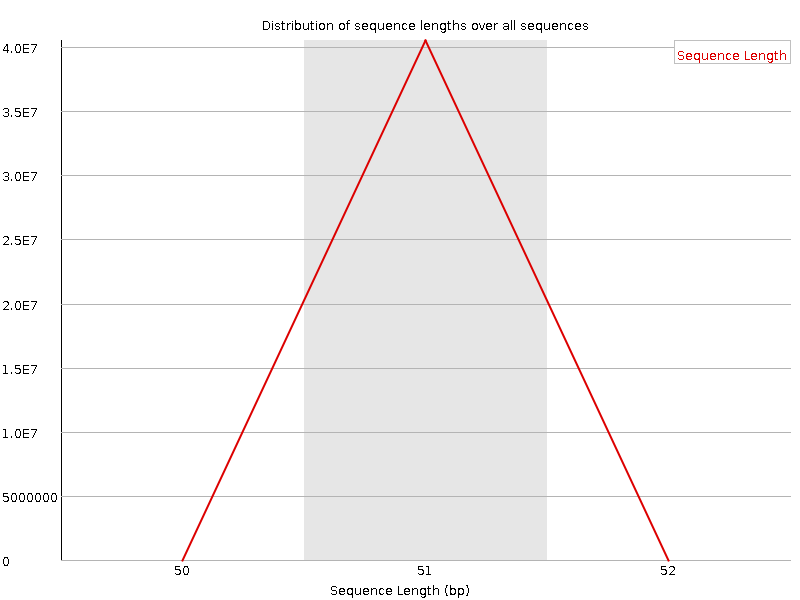
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
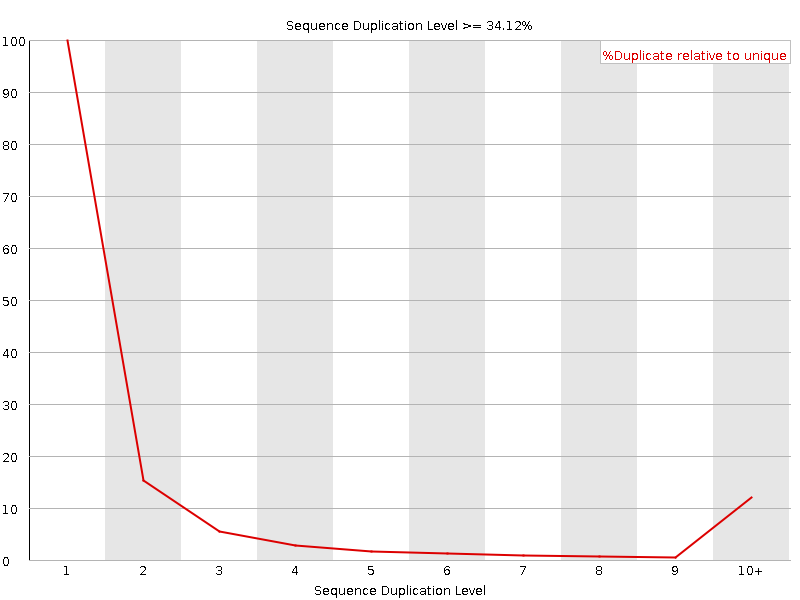
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTTGTGTTTGATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 118267 | 0.2921073179172731 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
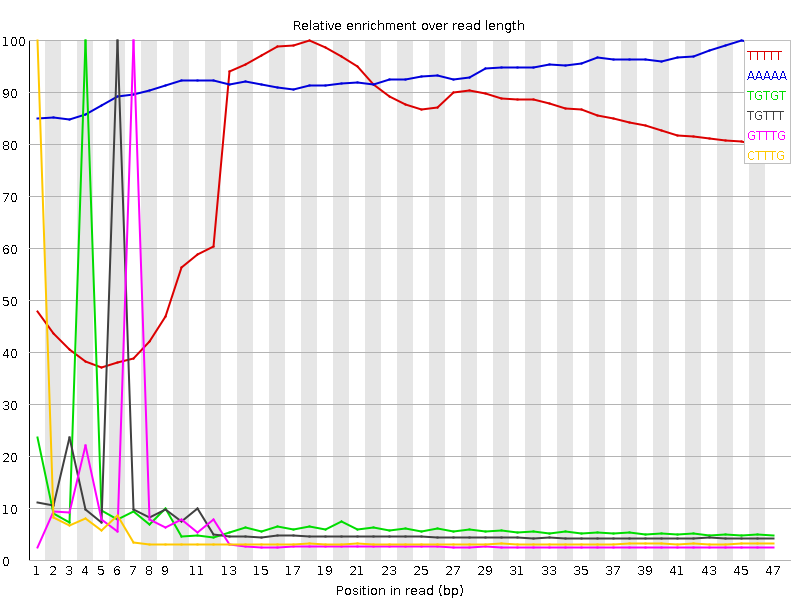
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| TTTTT | 22224850 | 5.9540434 | 7.656367 | 18 |
| AAAAA | 11158730 | 3.8994997 | 4.1863093 | 45 |
| TGTGT | 7732875 | 3.3885794 | 39.950283 | 4 |
| TGTTT | 7240215 | 2.4807122 | 31.344193 | 6 |
| GTTTG | 5478360 | 2.400641 | 39.26574 | 7 |
| CTTTG | 5186260 | 2.3724613 | 41.164158 | 1 |
| GTGTT | 5130845 | 2.2483582 | 39.237892 | 5 |
| TTTGA | 6090240 | 2.200612 | 32.613586 | 8 |
| GAGGG | 2841580 | 2.1479502 | 11.3781 | 11 |
| TTTGT | 6069095 | 2.0794516 | 31.138437 | 2 |
| TTGTG | 4705610 | 2.0620184 | 39.295086 | 3 |
| TGAGG | 3162485 | 1.8691366 | 14.369916 | 10 |
| TTGAG | 3823120 | 1.7667648 | 23.438133 | 9 |
| AGGGG | 2132515 | 1.6119679 | 6.392594 | 12 |
| TGAGC | 2331380 | 1.438447 | 5.530727 | 10 |
| GAGTG | 2070940 | 1.2239962 | 5.3953853 | 11 |
| TGAGT | 2550875 | 1.1788268 | 6.7447796 | 10 |
| TTGAT | 2958560 | 1.069029 | 8.845723 | 9 |
| TGATT | 2769520 | 1.0007223 | 5.638653 | 10 |
| TTGAC | 1867855 | 0.90109843 | 6.260845 | 9 |