![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | 20S15C_CTTGTA_L004_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 12211451 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 44 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
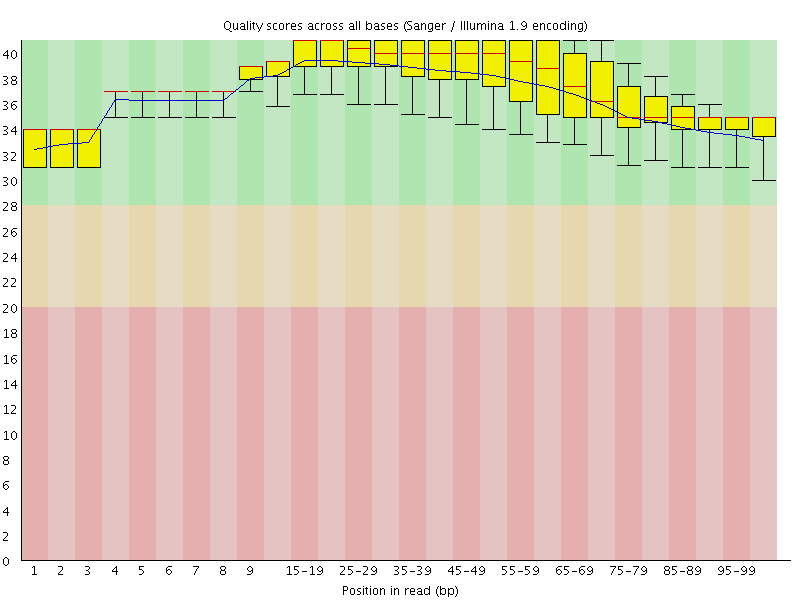
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
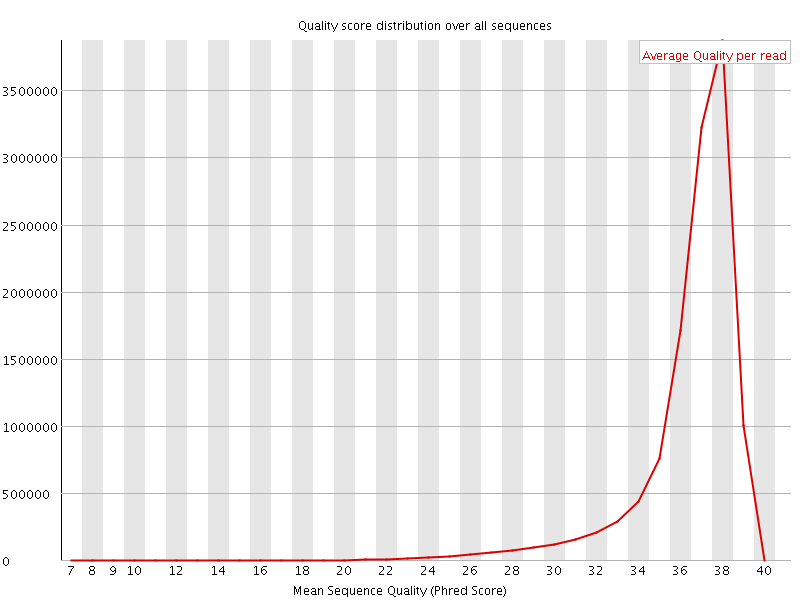
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
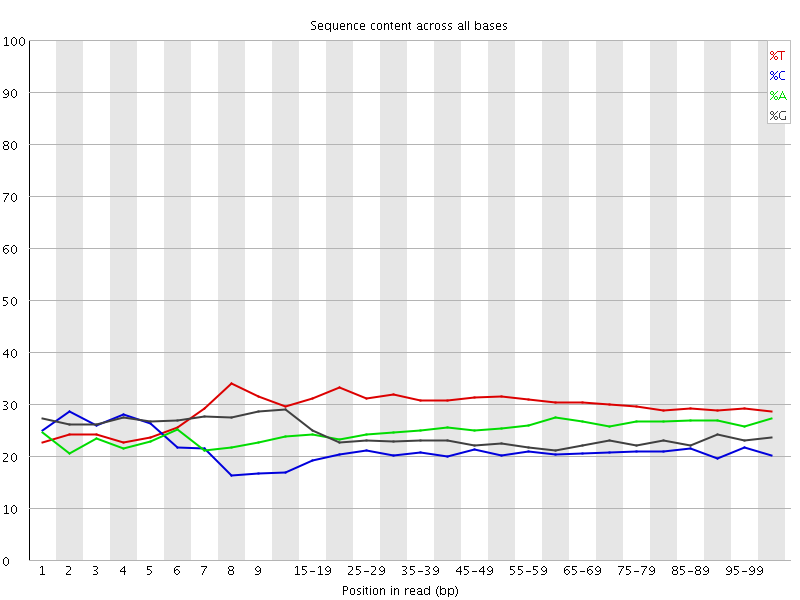
![[WARN]](Icons/warning.png) Per base GC content
Per base GC content
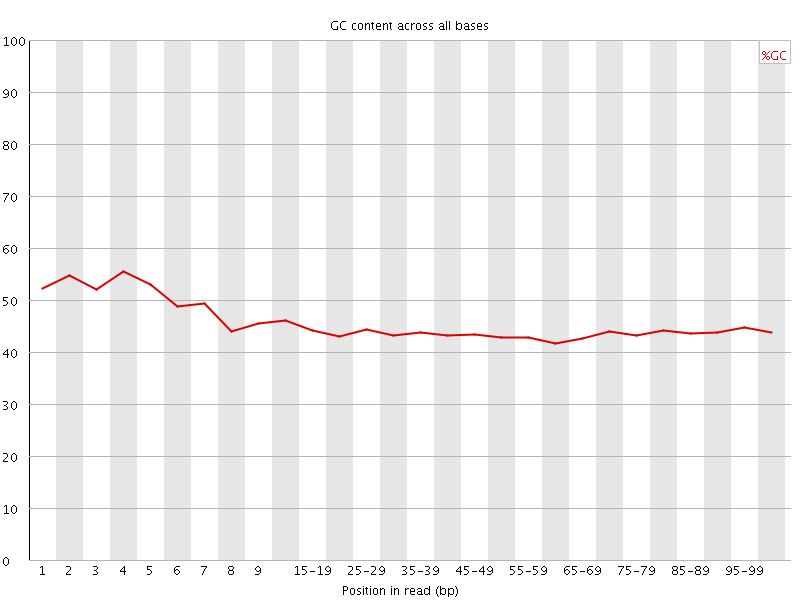
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
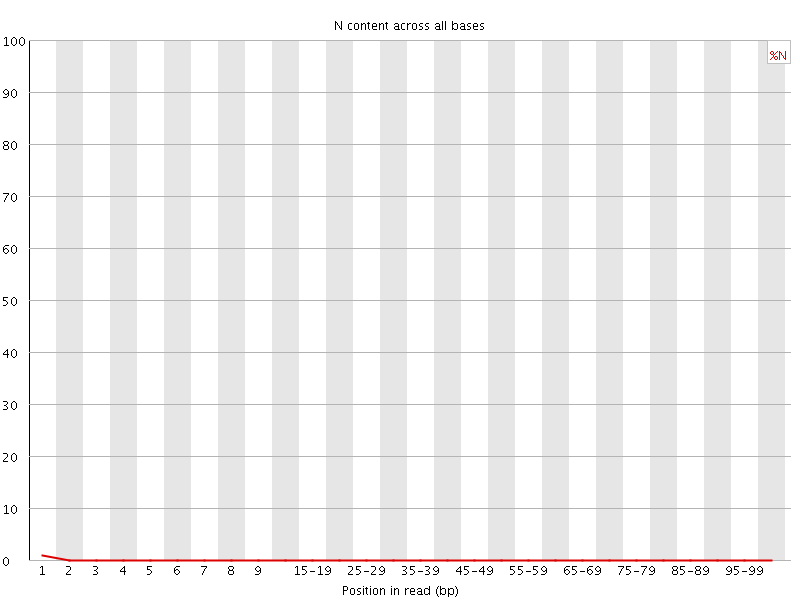
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
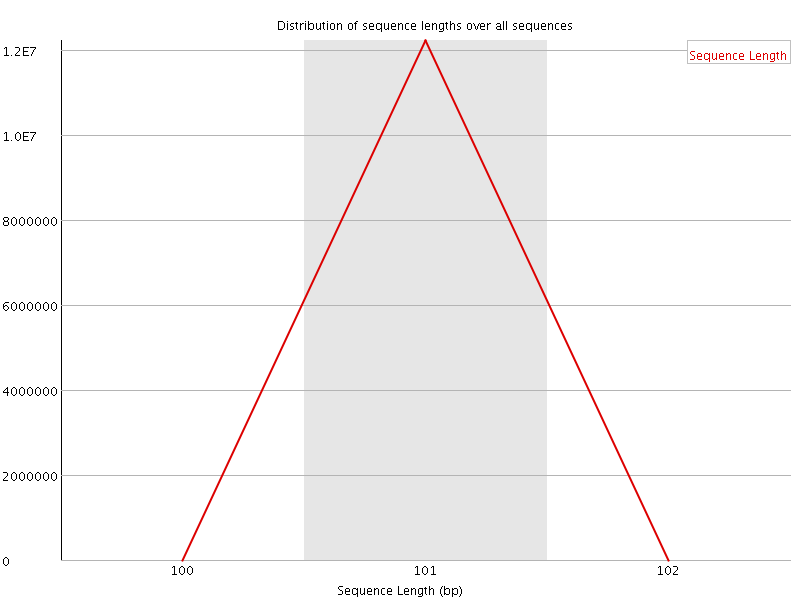
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
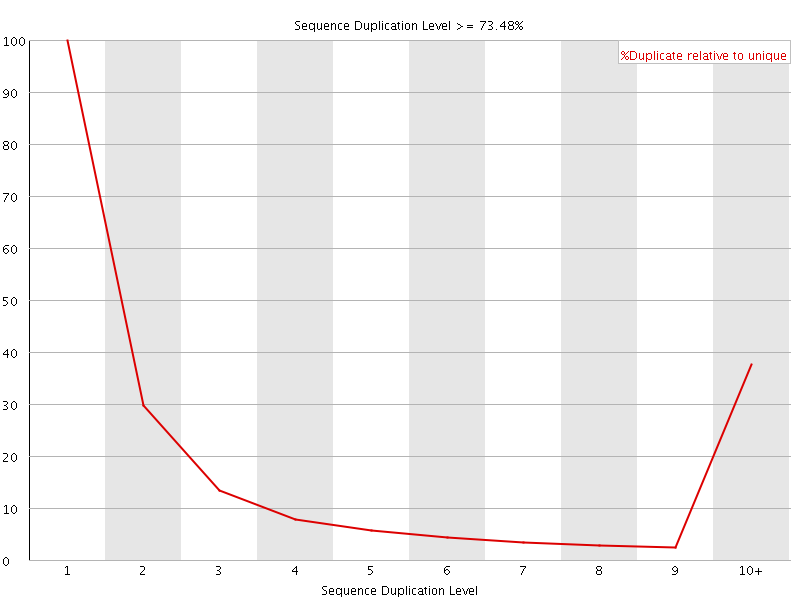
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| ACCCAATTGGTCAAGGAAGCTTCTCTGACGGTATGCCTTTAGGAATCTCT | 183110 | 1.4994942042514032 | No Hit |
| CTGGTACTTTCAACTTCATGATTGTGTTCCAAGCTGAACACAACATCCTT | 99309 | 0.81324487974443 | No Hit |
| GGTGTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTTCCTTAGTAAC | 83212 | 0.6814259828745987 | No Hit |
| GTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTTCCTTAGTAACTTC | 68470 | 0.5607032284697371 | No Hit |
| GGTACTTTCAACTTCATGATTGTGTTCCAAGCTGAACACAACATCCTTAT | 63383 | 0.5190456072746801 | No Hit |
| TGGTGTAGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTT | 60761 | 0.49757395742733607 | No Hit |
| CTGACGGTATGCCTTTAGGAATCTCTGGTACTTTCAACTTCATGATTGTG | 58523 | 0.4792468970313192 | No Hit |
| ACATGGGTCGTGAGTGGGAACTTAGCTACCGTTTAGGTATGCGTCCTTGG | 56820 | 0.4653009703760839 | No Hit |
| GCTACTGCTGTTTTCTTGATCTACCCAATTGGTCAAGGAAGCTTCTCTGA | 51513 | 0.4218417614745373 | No Hit |
| GTAGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTTCCTT | 49797 | 0.40778937736391846 | No Hit |
| CCTGTAGATATTGATGGTATCCGTGAGCCTGTTTCTGGTTCTCTTCTTTA | 38781 | 0.3175789674789671 | No Hit |
| GGTGTAGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTTC | 36896 | 0.3021426364483631 | No Hit |
| AGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTTCCTTAG | 36637 | 0.3000216763757231 | No Hit |
| ATGCATGGTTCCTTAGTAACTTCAAGTTTAATCCGTGAAACTACTGAGAA | 33588 | 0.27505330857078325 | No Hit |
| CCACATGCTTGGTGTAGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTA | 32744 | 0.2681417630058869 | No Hit |
| GCTACTTTAGAAAGACGCGAAAGCGCAAGCCTATGGGGTCGCTTCTGCGA | 32726 | 0.2679943603753559 | No Hit |
| CCTGTTGCGGCTGCTACTGCTGTTTTCTTGATCTACCCAATTGGTCAAGG | 31103 | 0.2547035565224804 | No Hit |
| GAACACAACATCCTTATGCACCCATTCCACATGCTTGGTGTAGCTGGTGT | 27818 | 0.22780257645057905 | No Hit |
| GCTTGGCCTGTAGTAGGTATCTGGTTCACTGCATTAGGTATCAGCACTAT | 26361 | 0.21587115241260027 | No Hit |
| TGCTACATGGGTCGTGAGTGGGAACTTAGCTACCGTTTAGGTATGCGTCC | 25612 | 0.20973756517550618 | No Hit |
| CTATTCAGTGCTATGCATGGTTCCTTAGTAACTTCAAGTTTAATCCGTGA | 25533 | 0.2090906314081758 | No Hit |
| GACTGCTACTTTAGAAAGACGCGAAAGCGCAAGCCTATGGGGTCGCTTCT | 25351 | 0.20760022703280714 | No Hit |
| TGGCCTGTAGTAGGTATCTGGTTCACTGCATTAGGTATCAGCACTATGGC | 25075 | 0.20534005336466568 | No Hit |
| ACATGCTTGGTGTAGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATG | 24387 | 0.19970599726437097 | No Hit |
| GCAACTTCTGTATTTATTATTGCTTTCATTGCAGCTCCTCCTGTAGATAT | 23965 | 0.19625022448192275 | No Hit |
| ATGGTTCGGTGTTTTAATGATTCCTACTCTATTAACTGCAACTTCTGTAT | 23612 | 0.1933594951165099 | No Hit |
| GACGGTATGCCTTTAGGAATCTCTGGTACTTTCAACTTCATGATTGTGTT | 22464 | 0.18395848290264605 | No Hit |
| CTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTTCCTTAGTA | 21780 | 0.17835718294246933 | No Hit |
| AGCCTATGGGGTCGCTTCTGCGACTGGGTTACCAGCACTGAAAACCGCCT | 21733 | 0.1779722982960829 | No Hit |
| AAGCCTATGGGGTCGCTTCTGCGACTGGGTTACCAGCACTGAAAACCGCC | 21052 | 0.17239556544099469 | No Hit |
| GCTTGGTGTAGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATGCATG | 20635 | 0.16898073783369397 | No Hit |
| ACTTTAGAAAGACGCGAAAGCGCAAGCCTATGGGGTCGCTTCTGCGACTG | 19697 | 0.16129942297602473 | No Hit |
| CCTCCTGTAGATATTGATGGTATCCGTGAGCCTGTTTCTGGTTCTCTTCT | 19492 | 0.1596206707949776 | No Hit |
| ATTAACTGCAACTTCTGTATTTATTATTGCTTTCATTGCAGCTCCTCCTG | 19329 | 0.1582858580851694 | No Hit |
| GGTATCCGTGAGCCTGTTTCTGGTTCTCTTCTTTACGGAAACAACATCAT | 19014 | 0.15570631205087748 | No Hit |
| CAAGCCTATGGGGTCGCTTCTGCGACTGGGTTACCAGCACTGAAAACCGC | 19008 | 0.1556571778407005 | No Hit |
| GGCCTGTAGTAGGTATCTGGTTCACTGCATTAGGTATCAGCACTATGGCT | 17901 | 0.14659191606304608 | No Hit |
| CCCCGGTTGTTACCCGACTACCAATAGGGGTGCTTCGTGCACCCCCAGAA | 17871 | 0.14634624501216112 | No Hit |
| TTACATCGGATGGTTCGGTGTTTTAATGATTCCTACTCTATTAACTGCAA | 17738 | 0.14525710335323788 | No Hit |
| ATTCAGTGCTATGCATGGTTCCTTAGTAACTTCAAGTTTAATCCGTGAAA | 17279 | 0.14149833627469824 | No Hit |
| CCACGTAGTTGGTTAGCTACTTCTCACTTTGTATTAGGATTTTTCTTCTT | 16344 | 0.13384158852211747 | No Hit |
| GTATTTATTATTGCTTTCATTGCAGCTCCTCCTGTAGATATTGATGGTAT | 16119 | 0.1319990556404804 | No Hit |
| GATGGTTCGGTGTTTTAATGATTCCTACTCTATTAACTGCAACTTCTGTA | 16045 | 0.13139306704829753 | No Hit |
| TGACCAAACTTGTAACCTGCGTTAGCAGACTCATTCTCAGTAGTTTCACG | 15783 | 0.12924753987056903 | No Hit |
| GCAAGCCTATGGGGTCGCTTCTGCGACTGGGTTACCAGCACTGAAAACCG | 15043 | 0.1231876539487404 | No Hit |
| GGCTCTCTATTCAGTGCTATGCATGGTTCCTTAGTAACTTCAAGTTTAAT | 14955 | 0.12246701886614458 | No Hit |
| GTAGATATTGATGGTATCCGTGAGCCTGTTTCTGGTTCTCTTCTTTACGG | 14866 | 0.12173819474851923 | No Hit |
| TTCATGCGTTGGAGAGACCGTTTCTTATTCTGTGCAGAAGCTATTTACAA | 14671 | 0.12014133291776709 | No Hit |
| TAGCTGCTTGGCCTGTAGTAGGTATCTGGTTCACTGCATTAGGTATCAGC | 14408 | 0.11798761670500908 | No Hit |
| ACTCTATTAACTGCAACTTCTGTATTTATTATTGCTTTCATTGCAGCTCC | 14119 | 0.11562098558148413 | No Hit |
| CATGCTTGGTGTAGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATGC | 13937 | 0.11413058120611547 | No Hit |
| ATGCCTTTAGGAATCTCTGGTACTTTCAACTTCATGATTGTGTTCCAAGC | 13858 | 0.11348364743878513 | No Hit |
| CTCTATTCAGTGCTATGCATGGTTCCTTAGTAACTTCAAGTTTAATCCGT | 13851 | 0.11342632419357862 | No Hit |
| GCCCTAGGTAACTTCTGTATTATAGTTGCTCCAGTTTTAAACTGAAAAAA | 13088 | 0.10717809046607155 | No Hit |
| TGACTGCTACTTTAGAAAGACGCGAAAGCGCAAGCCTATGGGGTCGCTTC | 12977 | 0.10626910757779726 | No Hit |
| TACCAGCACTGAAAACCGCCTTTACATCGGATGGTTCGGTGTTTTAATGA | 12655 | 0.10363223829829887 | No Hit |
| CCTACTCTATTAACTGCAACTTCTGTATTTATTATTGCTTTCATTGCAGC | 12643 | 0.10353396987794489 | No Hit |
| TACATCGGATGGTTCGGTGTTTTAATGATTCCTACTCTATTAACTGCAAC | 12611 | 0.10327192075700095 | No Hit |
| TGACCGAACTTGTAACCTGCGTTAGCAGACTCATTCTCAGTAGTTTCACG | 12450 | 0.10195348611725175 | No Hit |
| GCTCACGGTTACTTTGGTAGATTAATTTTCCAATACGCTAGCTTTAACAA | 12439 | 0.10186340673192729 | No Hit |
| GCGACTGGGTTACCAGCACTGAAAACCGCCTTTACATCGGATGGTTCGGT | 12237 | 0.10020922165596864 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
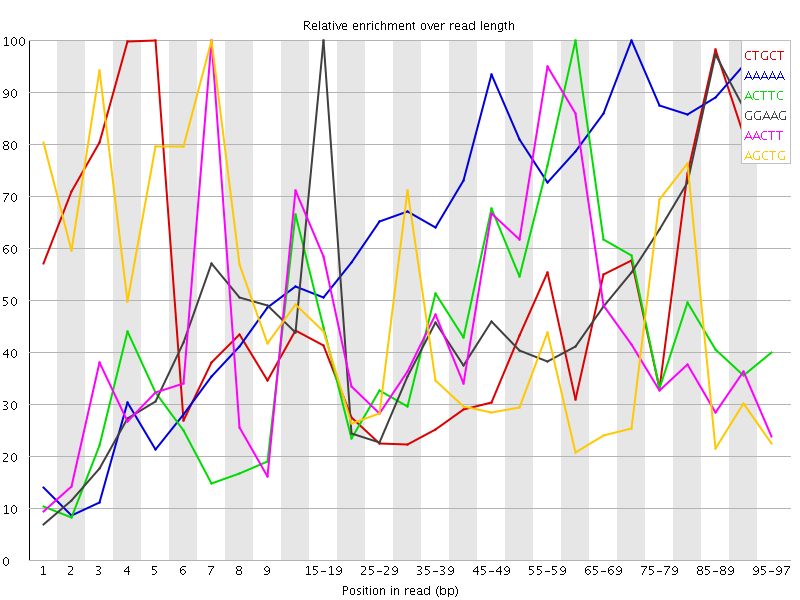
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CTGCT | 3747660 | 3.4121647 | 7.102334 | 5 |
| AAAAA | 4273925 | 3.3769503 | 4.6622167 | 70-74 |
| ACTTC | 3957935 | 3.3381705 | 6.9393578 | 60-64 |
| GGAAG | 3161285 | 3.1423273 | 5.987529 | 15-19 |
| AACTT | 4497385 | 3.094837 | 6.5269246 | 7 |
| AGCTG | 3196085 | 3.0405147 | 7.4076815 | 7 |
| AGCAC | 2364840 | 3.0300786 | 6.73196 | 80-84 |
| GAAGA | 3232310 | 2.9762595 | 5.8539624 | 90-94 |
| CTGGT | 3444290 | 2.762087 | 12.844238 | 1 |
| CTACT | 3265430 | 2.7541034 | 9.814975 | 2 |
| TCAAG | 3118315 | 2.7480128 | 5.2360525 | 10-14 |
| TCTCT | 3851490 | 2.7382774 | 8.250918 | 20-24 |
| CTTCA | 3165340 | 2.6696858 | 7.2735105 | 60-64 |
| CTTCT | 3693340 | 2.6258383 | 5.524653 | 20-24 |
| CGGAA | 2298745 | 2.5942435 | 7.156334 | 95-97 |
| TTCAA | 3746965 | 2.5784419 | 8.078571 | 9 |
| CAACT | 2535270 | 2.5366187 | 9.64419 | 9 |
| GCTGC | 2163775 | 2.5229177 | 7.1906147 | 3 |
| CTGAA | 2858540 | 2.5190861 | 6.328597 | 80-84 |
| TGGTG | 3550420 | 2.5077558 | 7.873068 | 6 |
| ACAAC | 2105730 | 2.499337 | 7.2321963 | 85-89 |
| AGGAA | 2665935 | 2.4547503 | 6.069616 | 40-44 |
| GGTGT | 3468900 | 2.450176 | 9.712043 | 1 |
| CAAGC | 1895400 | 2.4285831 | 7.3652287 | 75-79 |
| CAACA | 2025555 | 2.4041755 | 6.971527 | 85-89 |
| CGGTG | 2340175 | 2.4032953 | 10.972718 | 9 |
| GGTGG | 2651520 | 2.3983998 | 9.37814 | 7 |
| GCAGC | 1733815 | 2.3981972 | 5.6658616 | 1 |
| AGAGC | 2109660 | 2.380852 | 8.727494 | 95-97 |
| ATGCA | 2659065 | 2.3432994 | 6.1660876 | 1 |
| CCGTG | 2005935 | 2.3388793 | 5.9405894 | 60-64 |
| TATGC | 3146465 | 2.3373868 | 6.953369 | 30-34 |
| ACCCA | 1584900 | 2.3056118 | 32.7107 | 1 |
| GAAGC | 2007065 | 2.2650688 | 6.798198 | 15-19 |
| ACATC | 2261685 | 2.2628884 | 5.729675 | 90-94 |
| TCTGC | 2478180 | 2.2563303 | 5.629725 | 85-89 |
| AAGCT | 2559440 | 2.2555048 | 5.256804 | 15-19 |
| AAGAG | 2448670 | 2.2546961 | 5.1330137 | 90-94 |
| TCAAC | 2238675 | 2.2398658 | 7.771921 | 8 |
| CCAAG | 1736535 | 2.225029 | 7.0602193 | 75-79 |
| TCGGA | 2335000 | 2.221343 | 5.7748685 | 85-89 |
| ATCGG | 2334570 | 2.2209342 | 5.732948 | 95-97 |
| CGTGA | 2329650 | 2.2162538 | 7.1816406 | 9 |
| CAGCA | 1719190 | 2.2028046 | 5.462557 | 2 |
| CATGG | 2299590 | 2.1876569 | 9.368575 | 2 |
| GAGCA | 1901065 | 2.1454427 | 5.9915676 | 90-94 |
| GCTGG | 2083950 | 2.1401594 | 7.9293923 | 4 |
| TCCTT | 2948570 | 2.0963326 | 5.718943 | 40-44 |
| ACTGC | 1934560 | 2.089504 | 6.9207234 | 4 |
| GATCG | 2195075 | 2.0882292 | 6.315501 | 95-97 |
| GCACA | 1611070 | 2.06427 | 6.531101 | 95-97 |
| CAAGG | 1814555 | 2.0478117 | 6.4879317 | 10-14 |
| CTCTG | 2229145 | 2.0295894 | 6.6691666 | 20-24 |
| CTCTC | 1914255 | 1.9787996 | 5.0004735 | 15-19 |
| GGCTC | 1678395 | 1.9569744 | 5.1177506 | 10-14 |
| CTTGG | 2437050 | 1.9543489 | 6.0222826 | 2 |
| CACAC | 1338930 | 1.9477904 | 7.435823 | 95-97 |
| CCTTA | 2300805 | 1.940527 | 6.917419 | 95-97 |
| AAGGA | 2100620 | 1.9342173 | 5.8336 | 10-14 |
| AGATC | 2166885 | 1.909566 | 5.919674 | 95-97 |
| GCTAC | 1760595 | 1.9016057 | 11.512498 | 1 |
| TGGCT | 2368530 | 1.8994007 | 7.2262936 | 9 |
| TTCCA | 2231670 | 1.8822176 | 5.2870584 | 75-79 |
| TAGCT | 2530805 | 1.880037 | 5.439993 | 6 |
| CATCC | 1532495 | 1.8792832 | 6.8338313 | 90-94 |
| TTTCA | 3213095 | 1.8638464 | 7.0335646 | 8 |
| TATTC | 3208340 | 1.8610879 | 5.8509574 | 2 |
| CCAGC | 1171840 | 1.8402746 | 6.3427873 | 1 |
| GTGGC | 1777625 | 1.8255721 | 8.593819 | 8 |
| CACAA | 1537440 | 1.8248212 | 6.3514013 | 85-89 |
| TGCTG | 2258390 | 1.8110759 | 5.3219457 | 6 |
| GCTTC | 1989000 | 1.8109423 | 7.0891733 | 15-19 |
| GCATG | 1901865 | 1.8092912 | 5.1167574 | 3 |
| TTCGG | 2251540 | 1.8055826 | 8.496718 | 7 |
| CTCTA | 2109760 | 1.7793972 | 5.3386908 | 95-97 |
| CACCC | 996665 | 1.777034 | 6.96975 | 50-54 |
| CCTGT | 1939480 | 1.7658554 | 7.7774277 | 1 |
| TGAAC | 1980385 | 1.745213 | 5.951833 | 80-84 |
| TCGGT | 2168260 | 1.7387975 | 9.353777 | 8 |
| TTGGT | 3148885 | 1.7367655 | 15.308171 | 7 |
| GCACC | 1097940 | 1.7242209 | 6.3421535 | 50-54 |
| GCAGG | 1411865 | 1.7200582 | 5.1493964 | 90-94 |
| TGTAG | 2617030 | 1.7123172 | 6.358765 | 3 |
| GTGTA | 2603080 | 1.7031896 | 8.869972 | 2 |
| CATGC | 1549110 | 1.6731822 | 6.1068287 | 2 |
| CTTTC | 2339840 | 1.6635463 | 8.233443 | 7 |
| CCACA | 1132230 | 1.6470964 | 7.011929 | 1 |
| CCCAA | 1111265 | 1.6165977 | 32.187775 | 2 |
| TGGTA | 2465940 | 1.6134592 | 9.446455 | 2 |
| GTAGC | 1683895 | 1.601931 | 6.8401804 | 5 |
| GTCTG | 1996070 | 1.600713 | 6.063332 | 90-94 |
| GCTTG | 1994935 | 1.5998029 | 5.6303515 | 1 |
| GAACA | 1523690 | 1.5928929 | 5.8974915 | 80-84 |
| CGCAG | 1119535 | 1.5485308 | 5.9577727 | 90-94 |
| ACACA | 1298305 | 1.5409865 | 6.0974092 | 85-89 |
| GCTGA | 1617990 | 1.5392339 | 5.217301 | 80-84 |
| TCCAA | 1538045 | 1.5388633 | 5.4234357 | 75-79 |
| TACTT | 2651205 | 1.5379062 | 7.360319 | 5 |
| CCAAT | 1524165 | 1.524976 | 22.490192 | 3 |
| CCGCC | 773230 | 1.4882817 | 5.200771 | 45-49 |
| ATGCC | 1376420 | 1.4866612 | 7.115652 | 9 |
| ACACG | 1156560 | 1.4819047 | 6.329976 | 95-97 |
| AACAC | 1246545 | 1.4795516 | 6.1451917 | 85-89 |
| GCAAC | 1152950 | 1.4772793 | 5.708476 | 1 |
| ATCTC | 1736265 | 1.464387 | 5.1944013 | 45-49 |
| GAGCC | 1037160 | 1.4345902 | 5.1415405 | 95-97 |
| ACGGT | 1496215 | 1.4233865 | 6.8798532 | 4 |
| GTATT | 2757390 | 1.4088104 | 5.7569256 | 4 |
| ACATG | 1585115 | 1.3968816 | 8.992122 | 1 |
| GCTGT | 1704765 | 1.367106 | 5.4258437 | 7 |
| ACTTT | 2343865 | 1.3596249 | 7.3051877 | 6 |
| GGTAC | 1429030 | 1.3594718 | 11.084618 | 3 |
| GGTCA | 1404880 | 1.3364972 | 21.283825 | 9 |
| GACGG | 1089700 | 1.3275684 | 8.532249 | 3 |
| TGGTC | 1596875 | 1.2805856 | 18.197754 | 8 |
| TGTAT | 2463485 | 1.2586482 | 5.534696 | 3 |
| AACGC | 972095 | 1.2455491 | 5.1319203 | 95-97 |
| GCGGC | 827485 | 1.2355839 | 5.589014 | 7 |
| ATGGG | 1468240 | 1.2302507 | 7.523967 | 3 |
| ATTCG | 1655190 | 1.2295765 | 6.990006 | 6 |
| CAATT | 1717420 | 1.1818278 | 15.586326 | 4 |
| GGTCG | 1133290 | 1.1638577 | 7.0848126 | 6 |
| CTGAC | 1076225 | 1.1624225 | 7.8768873 | 1 |
| TGGGT | 1629310 | 1.1508249 | 5.7399726 | 4 |
| GTACT | 1546440 | 1.1487904 | 8.623589 | 4 |
| TGACG | 1203670 | 1.1450812 | 6.63767 | 2 |
| ATTGG | 1669840 | 1.0925727 | 14.860702 | 6 |
| ACTTG | 1451540 | 1.0782928 | 6.4076214 | 8 |
| GGGTC | 1015435 | 1.0428239 | 7.022565 | 5 |
| TGGCC | 878105 | 1.0238527 | 8.327368 | 1 |
| TCGTG | 1224520 | 0.9819822 | 5.5868354 | 8 |
| CGGTA | 987070 | 0.9390241 | 6.3982534 | 5 |
| AATTG | 1534425 | 0.9300176 | 12.773544 | 5 |
| CTTGT | 1350490 | 0.8456835 | 5.26991 | 9 |
| GTCGT | 829825 | 0.6654634 | 5.3523483 | 7 |
| CCCCG | 326070 | 0.6276064 | 5.1788697 | 1 |
| TGACC | 564840 | 0.6100795 | 5.9033027 | 1 |