| Sequence |
Count |
Percentage |
Possible Source |
| GCGAGTCGGGCTGTTTGGGAATGCAGCCCTAAGTGGGAGGTAAATTCCTT |
220296 |
1.605692022823836 |
No Hit |
| ACCCAATTGGTCAAGGAAGCTTCTCTGACGGTATGCCTTTAGGAATCTCT |
193179 |
1.4080418131835613 |
No Hit |
| GATACCGTCCTAGTCTCAACCATAAACGATGCCGACTAGGGATTGGCGGA |
134561 |
0.9807873237970648 |
No Hit |
| ACATGGGTCGTGAGTGGGAACTTAGCTACCGTTTAGGTATGCGTCCTTGG |
80264 |
0.5850277105346097 |
No Hit |
| GGTGTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTTCCTTAGTAAC |
75729 |
0.5519730326307617 |
No Hit |
| CTGGTACTTTCAACTTCATGATTGTGTTCCAAGCTGAACACAACATCCTT |
70698 |
0.5153031132185767 |
No Hit |
| CTGACGGTATGCCTTTAGGAATCTCTGGTACTTTCAACTTCATGATTGTG |
66271 |
0.4830356249979956 |
No Hit |
| ATGGTTCGGTGTTTTAATGATTCCTACTCTATTAACTGCAACTTCTGTAT |
63463 |
0.4625686932330551 |
No Hit |
| GCTACTGCTGTTTTCTTGATCTACCCAATTGGTCAAGGAAGCTTCTCTGA |
62975 |
0.4590117620716267 |
No Hit |
| GCTACTTTAGAAAGACGCGAAAGCGCAAGCCTATGGGGTCGCTTCTGCGA |
59883 |
0.43647481299142865 |
No Hit |
| GACTGCTACTTTAGAAAGACGCGAAAGCGCAAGCCTATGGGGTCGCTTCT |
56979 |
0.4153081570635842 |
No Hit |
| CATGGAGAGTTTGATCCTGGCTCAGGATGAACGCTGGCGGCATGCTTAAC |
53872 |
0.39266187608293246 |
No Hit |
| CAGACCTCTGATCAGGAAAGACTACCCGCTGAGTTTAAGCATATCACTAA |
47398 |
0.3454742278470974 |
No Hit |
| GTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTTCCTTAGTAACTTC |
45512 |
0.3317275635633803 |
No Hit |
| GTAGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTTCCTT |
43551 |
0.31743423977739443 |
No Hit |
| GCTTGGCCTGTAGTAGGTATCTGGTTCACTGCATTAGGTATCAGCACTAT |
43134 |
0.3143948129447804 |
No Hit |
| CAACACGGGGAAACTTACCAGGTCCAGACATAGTAAGGATTGACAGATTG |
39658 |
0.2890589672129666 |
No Hit |
| CCCCGGTTGTTACCCGACTACCAATAGGGGTGCTTCGTGCACCCCCAGAA |
37242 |
0.27144924244655055 |
No Hit |
| TGCTACATGGGTCGTGAGTGGGAACTTAGCTACCGTTTAGGTATGCGTCC |
36919 |
0.2690949621901133 |
No Hit |
| TGGTGTAGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTT |
35736 |
0.2604723196409948 |
No Hit |
| CAAACGAATAAGGGCTTATGGTGGATACCTAGGCACCCAGAGACGAGGAA |
34668 |
0.25268788832868844 |
No Hit |
| CCTGTTGCGGCTGCTACTGCTGTTTTCTTGATCTACCCAATTGGTCAAGG |
34478 |
0.2513030175896077 |
No Hit |
| GGTACTTTCAACTTCATGATTGTGTTCCAAGCTGAACACAACATCCTTAT |
34346 |
0.2503408968656148 |
No Hit |
| TGCCAGTAGTCATATGCTTGTCTCAAAGATTAAGCCATGCATGTGTAAGT |
34270 |
0.2497869485699825 |
No Hit |
| AAGCCTATGGGGTCGCTTCTGCGACTGGGTTACCAGCACTGAAAACCGCC |
33714 |
0.24573437945983045 |
No Hit |
| TGGGTCGGGGGGAGCGGTCCGCCCCTTGTGGGTGTGCACTGGTCCACTCG |
33704 |
0.24566149152619463 |
No Hit |
| AGGCACCCAGAGACGAGGAAGGGCGTAGCAAGCGACGAAATGCTTCGGGG |
33287 |
0.24262206469358058 |
No Hit |
| AGCCTATGGGGTCGCTTCTGCGACTGGGTTACCAGCACTGAAAACCGCCT |
33020 |
0.24067595686550397 |
No Hit |
| GATCAGGAAAGACTACCCGCTGAGTTTAAGCATATCACTAAGCGGAGGAG |
32672 |
0.23813945677497714 |
No Hit |
| GCAACTTCTGTATTTATTATTGCTTTCATTGCAGCTCCTCCTGTAGATAT |
31727 |
0.23125154704639142 |
No Hit |
| GGTGTAGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTTC |
29668 |
0.21624392151077446 |
No Hit |
| ACCTCCCCGGTTGTTACCCGACTACCAATAGGGGTGCTTCGTGCACCCCC |
29081 |
0.21196539980635135 |
No Hit |
| CCTGTAGATATTGATGGTATCCGTGAGCCTGTTTCTGGTTCTCTTCTTTA |
29006 |
0.21141874030408264 |
No Hit |
| TGACTGCTACTTTAGAAAGACGCGAAAGCGCAAGCCTATGGGGTCGCTTC |
28976 |
0.21120007650317515 |
No Hit |
| ACTTTAGAAAGACGCGAAAGCGCAAGCCTATGGGGTCGCTTCTGCGACTG |
28520 |
0.2078763867293814 |
No Hit |
| AGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTTCCTTAG |
27599 |
0.2011634080415216 |
No Hit |
| GACGGTATGCCTTTAGGAATCTCTGGTACTTTCAACTTCATGATTGTGTT |
27298 |
0.19896948123908323 |
No Hit |
| CCACATGCTTGGTGTAGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTA |
24198 |
0.17637422181197654 |
No Hit |
| CAAGCCTATGGGGTCGCTTCTGCGACTGGGTTACCAGCACTGAAAACCGC |
23828 |
0.1736773682674509 |
No Hit |
| CTGATCAGGAAAGACTACCCGCTGAGTTTAAGCATATCACTAAGCGGAGG |
22666 |
0.16520779037896768 |
No Hit |
| GATGGTTCGGTGTTTTAATGATTCCTACTCTATTAACTGCAACTTCTGTA |
22203 |
0.16183307905162886 |
No Hit |
| ATGCATGGTTCCTTAGTAACTTCAAGTTTAATCCGTGAAACTACTGAGAA |
21162 |
0.15424544516013916 |
No Hit |
| GAACACAACATCCTTATGCACCCATTCCACATGCTTGGTGTAGCTGGTGT |
21123 |
0.15396118221895944 |
No Hit |
| CTATTCAGTGCTATGCATGGTTCCTTAGTAACTTCAAGTTTAATCCGTGA |
21052 |
0.15344367789014507 |
No Hit |
| ACATGCTTGGTGTAGCTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATG |
20905 |
0.15237222526569838 |
No Hit |
| GGTTCGGTGTTTTAATGATTCCTACTCTATTAACTGCAACTTCTGTATTT |
20573 |
0.1499523458689889 |
No Hit |
| AAGTGGGAGGTAAATTCCTTCTAAGGCTAAATATTGGCAAGAGACCGATA |
20378 |
0.14853103116309027 |
No Hit |
| TGGCCTGTAGTAGGTATCTGGTTCACTGCATTAGGTATCAGCACTATGGC |
19883 |
0.14492307844811678 |
No Hit |
| GAAACATCTTAGTAGCCAGAGGAAAAGAAAGCAAAAGCGATTCCCATAGT |
19881 |
0.1449085008613896 |
No Hit |
| GTATTTATTATTGCTTTCATTGCAGCTCCTCCTGTAGATATTGATGGTAT |
19551 |
0.14250319905140726 |
No Hit |
| CCACGAGGCGCGATCTGCGAGTCGGGCTGTTTGGGAATGCAGCCCTAAGT |
19312 |
0.140761177437511 |
No Hit |
| GCAAGCCTATGGGGTCGCTTCTGCGACTGGGTTACCAGCACTGAAAACCG |
18987 |
0.13839231959434659 |
No Hit |
| GACACTGAGAGACGAAAGCTAGGGGAGCGAATGGGATTAGATACCCCAGT |
18941 |
0.13805703509962178 |
No Hit |
| ATTAACTGCAACTTCTGTATTTATTATTGCTTTCATTGCAGCTCCTCCTG |
18153 |
0.13231346592911852 |
No Hit |
| GGCTCTCTATTCAGTGCTATGCATGGTTCCTTAGTAACTTCAAGTTTAAT |
17859 |
0.1301705606802252 |
No Hit |
| TGGCAAGAGACCGATAGCGAACAAGTACCGCGAGGGAAAGATGAAAAGGA |
17858 |
0.13016327188686158 |
No Hit |
| CAACGCGAAGAACCTTACCAGGGCTTGACATGCCGCGAATCCTTTTGAAA |
17728 |
0.12921572874959583 |
No Hit |
| TTACATCGGATGGTTCGGTGTTTTAATGATTCCTACTCTATTAACTGCAA |
17436 |
0.12708740108742966 |
No Hit |
| ACCAATAGGGGTGCTTCGTGCACCCCCAGAAGCATGTGTCGAGAGCTCCC |
17277 |
0.12592848294262 |
No Hit |
| CCTCCTGTAGATATTGATGGTATCCGTGAGCCTGTTTCTGGTTCTCTTCT |
17212 |
0.12545471137398712 |
No Hit |
| ACATCGGTAGGGGAGCGTTCCGCCTTAGAGGGAAGCATCAGCGCGAGCAG |
17142 |
0.12494449583853633 |
No Hit |
| AACACGGACCAAGGAGTCTAACATGTATGCAAGCAGGTGGGCGGCAAACC |
17052 |
0.12428850443581387 |
No Hit |
| TACCATGACTGCTACTTTAGAAAGACGCGAAAGCGCAAGCCTATGGGGTC |
17037 |
0.12417917253536014 |
No Hit |
| GCTCACGGTTACTTTGGTAGATTAATTTTCCAATACGCTAGCTTTAACAA |
16856 |
0.12285990093655164 |
No Hit |
| CTGGTGTATTCGGTGGCTCTCTATTCAGTGCTATGCATGGTTCCTTAGTA |
16803 |
0.12247359488828174 |
No Hit |
| TTCATGCGTTGGAGAGACCGTTTCTTATTCTGTGCAGAAGCTATTTACAA |
16772 |
0.1222476422940107 |
No Hit |
| CCACGTAGTTGGTTAGCTACTTCTCACTTTGTATTAGGATTTTTCTTCTT |
16464 |
0.12000269393802718 |
No Hit |
| GCGACTGGGTTACCAGCACTGAAAACCGCCTTTACATCGGATGGTTCGGT |
16070 |
0.11713090935277556 |
No Hit |
| GCAACACGGGGAAACTTACCAGGTCCAGACATAGTAAGGATTGACAGATT |
15938 |
0.11616878862878263 |
No Hit |
| GTAGGTGGCTTTTTAAGTCTGCTGTCAAATCCCAGGGCTCAACCCTGGAC |
15821 |
0.11531599980524344 |
No Hit |
| ATCACTAAGCGGAGGAGAAGAAACTAACCAGGATTTCCCTAGTAACGGCG |
15691 |
0.11436845666797768 |
No Hit |
| GCAGCTATCGGTTTGCACTTCTACCCAATTTGGGAAGCTGCTTCCGTTGA |
15504 |
0.11300545230898769 |
No Hit |
| GCTAAAAACTATGGTAGAGCTGTATACGAATGTCTTCGTGGTGGACTTGA |
15130 |
0.11027944359100773 |
No Hit |
| GTGAGCCTGTTTCTGGTTCTCTTCTTTACGGAAACAACATCATCTCTGCT |
14756 |
0.10755343487302776 |
No Hit |
| ACCTGGTTGATCCTGCCAGTAGTCATATGCTTGTCTCAAAGATTAAGCCA |
14617 |
0.10654029259548976 |
No Hit |
| AAGCGGAGGAGAAGAAACTAACCAGGATTTCCCTAGTAACGGCGAGCGAA |
14568 |
0.10618314172067418 |
No Hit |
| GTTGCGGCTGCTACTGCTGTTTTCTTGATCTACCCAATTGGTCAAGGAAG |
14524 |
0.10586243481267654 |
No Hit |
| GGCCTGTAGTAGGTATCTGGTTCACTGCATTAGGTATCAGCACTATGGCT |
14472 |
0.10548341755777024 |
No Hit |
| TAGTAGCCAGAGGAAAAGAAAGCAAAAGCGATTCCCATAGTAGCGGCGAG |
14472 |
0.10548341755777024 |
No Hit |
| TACCAGCACTGAAAACCGCCTTTACATCGGATGGTTCGGTGTTTTAATGA |
14385 |
0.10484929253513854 |
No Hit |
| TAGCTGCTTGGCCTGTAGTAGGTATCTGGTTCACTGCATTAGGTATCAGC |
14124 |
0.10294691746724344 |
No Hit |
| GCCCTAGGTAACTTCTGTATTATAGTTGCTCCAGTTTTAAACTGAAAAAA |
13789 |
0.10050517169044319 |
No Hit |
![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality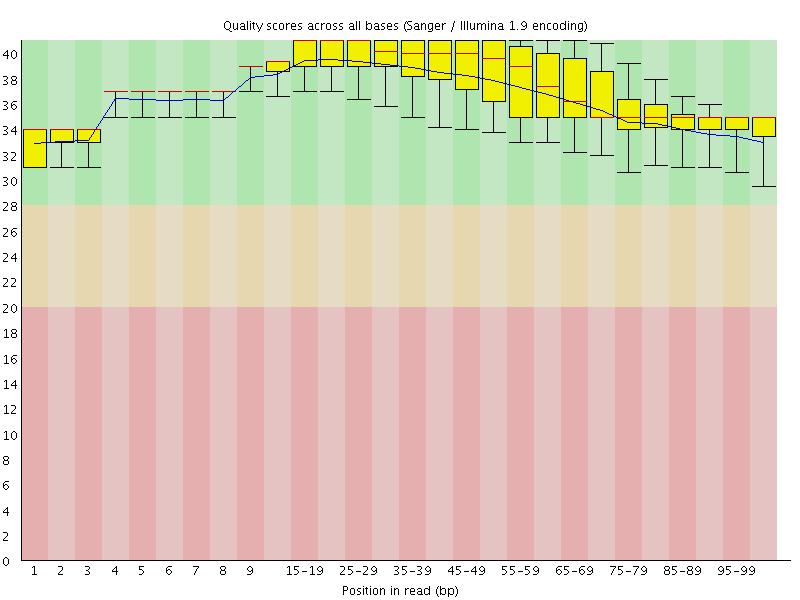
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores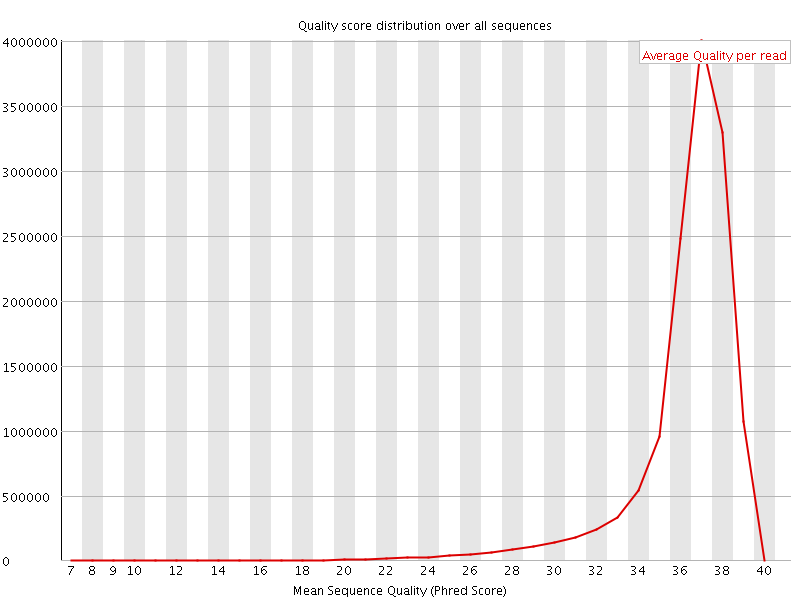
![[OK]](Icons/tick.png) Per base sequence content
Per base sequence content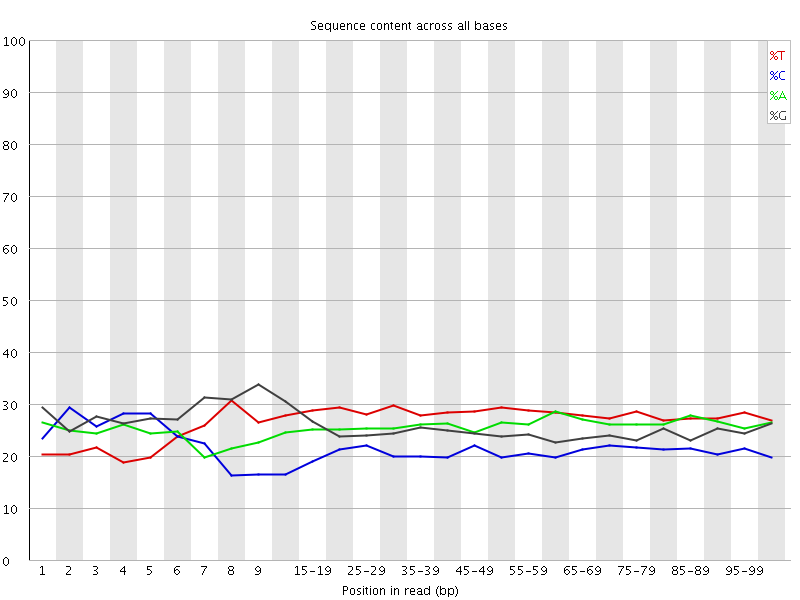
![[WARN]](Icons/warning.png) Per base GC content
Per base GC content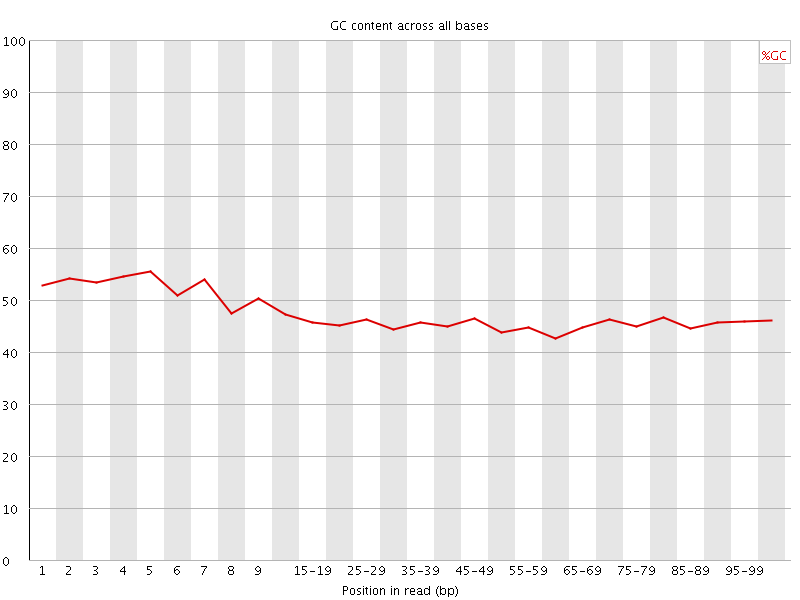
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content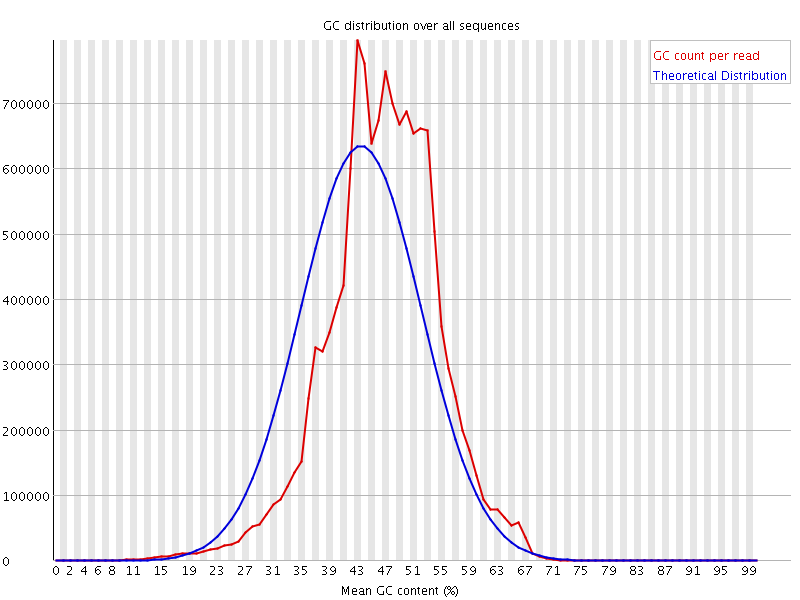
![[OK]](Icons/tick.png) Per base N content
Per base N content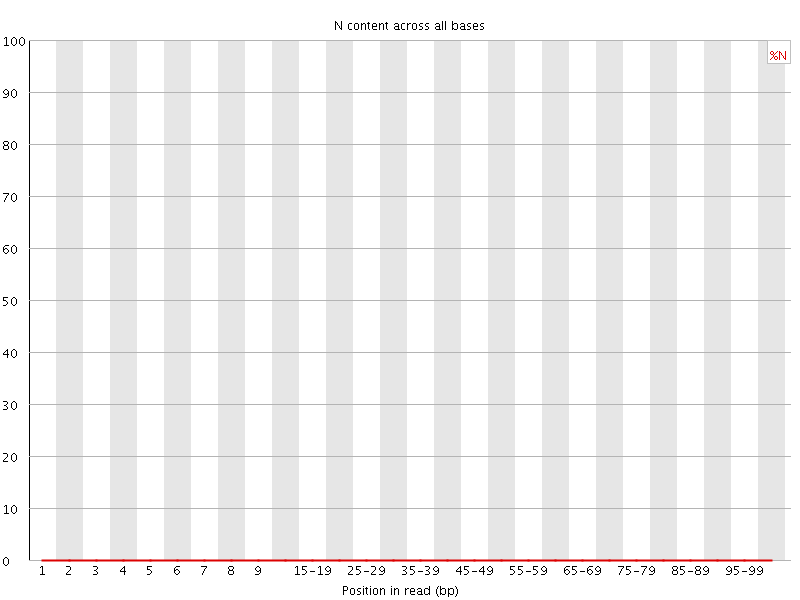
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution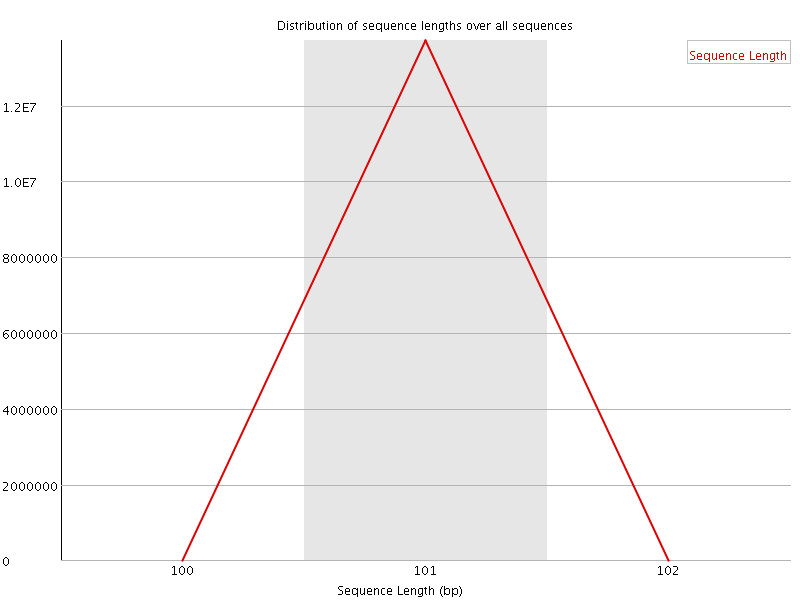
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels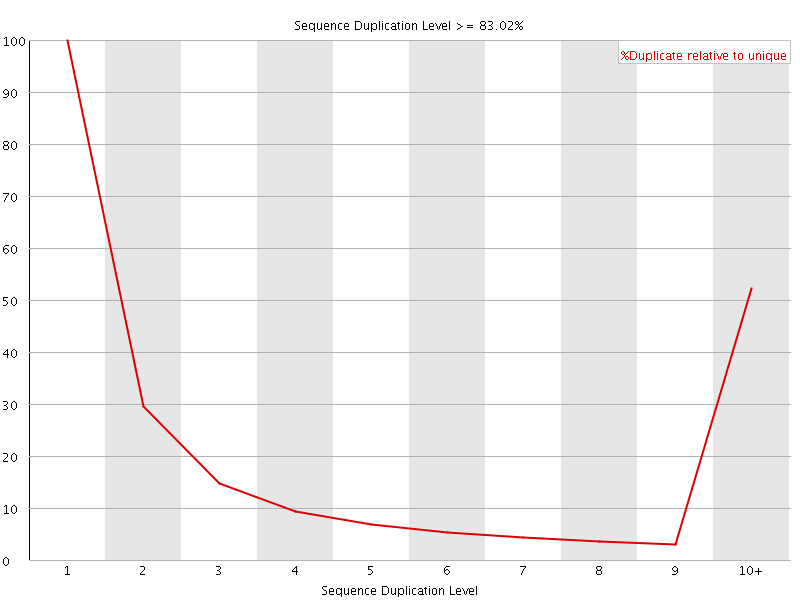
![[FAIL]](Icons/error.png) Overrepresented sequences
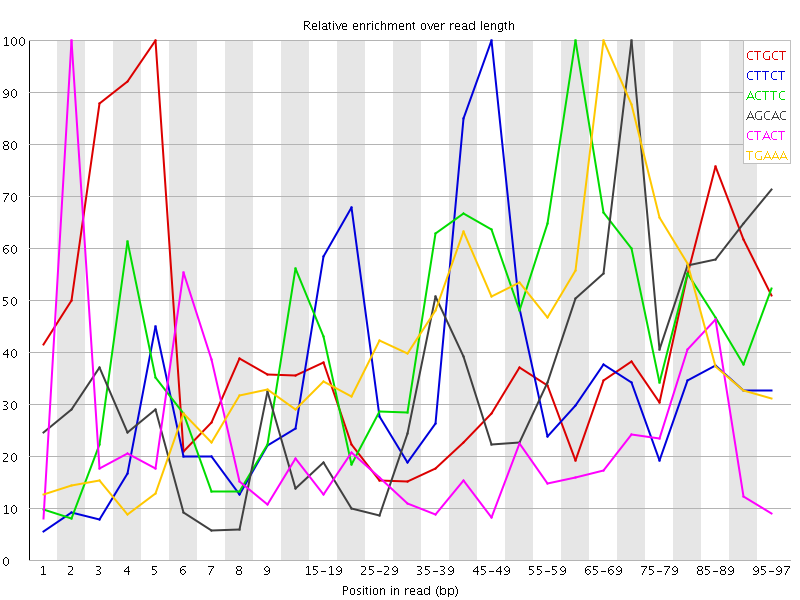
Overrepresented sequences![[WARN]](Icons/warning.png) Kmer Content
Kmer Content