![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL3_ACAGTG_L008_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 7195243 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
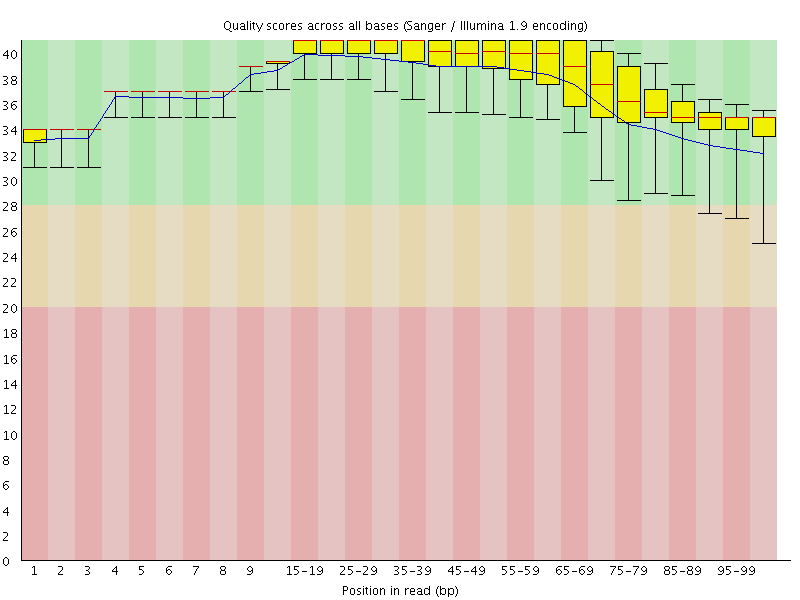
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
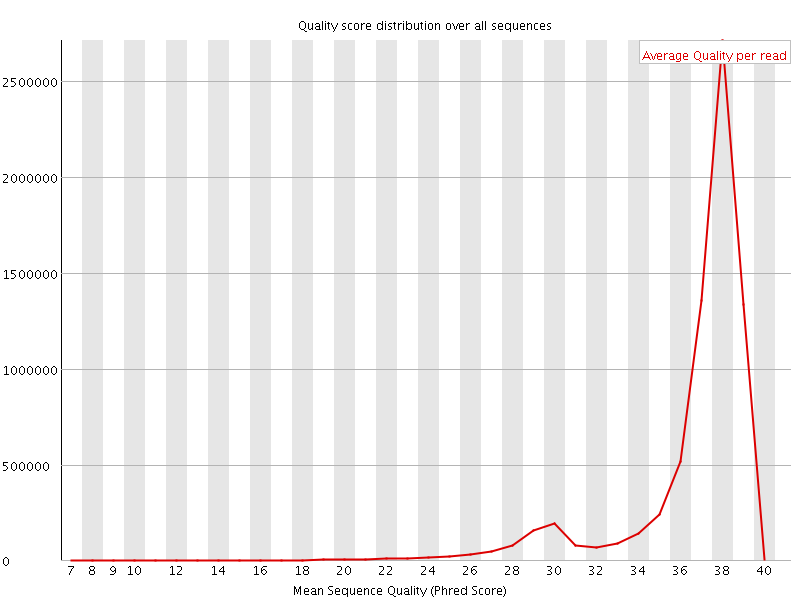
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
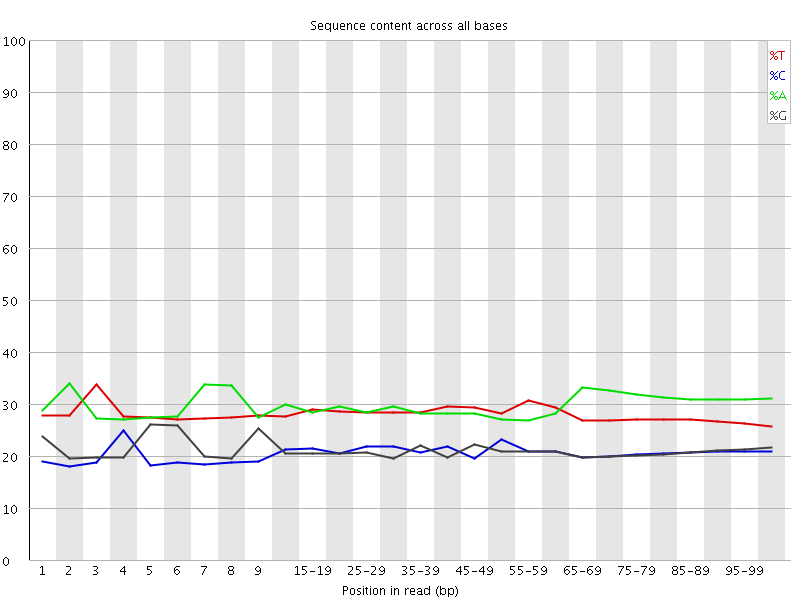
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
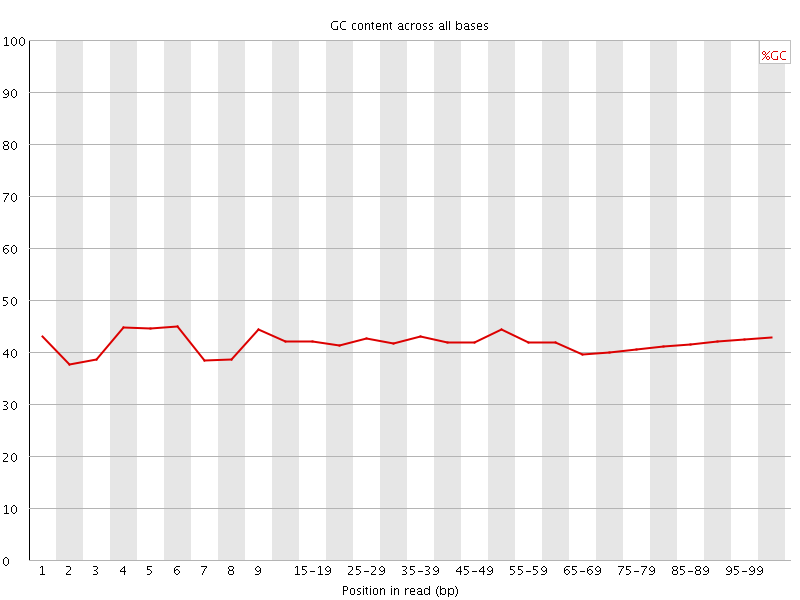
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
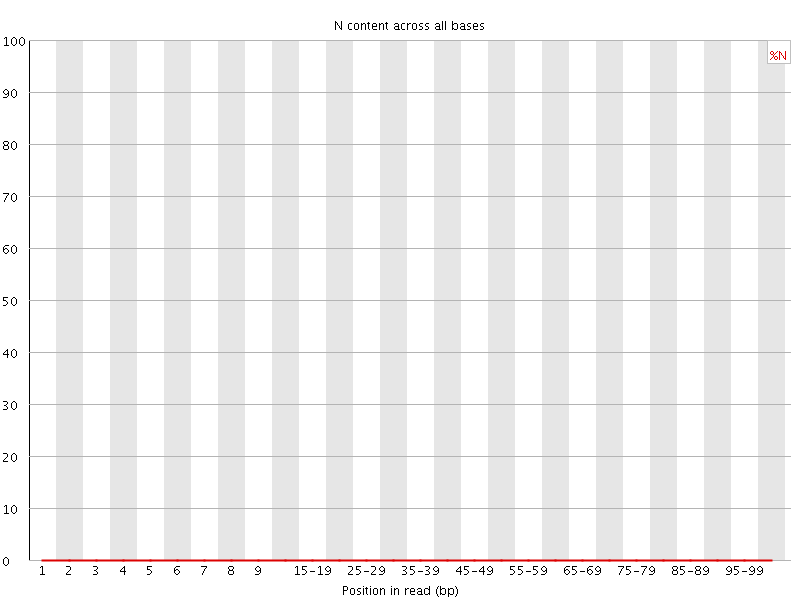
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
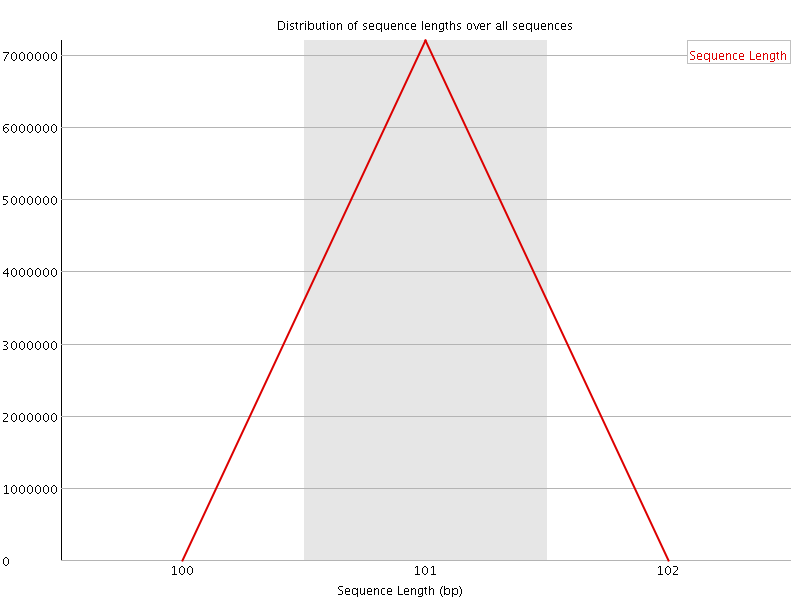
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACACAGTGATCTCGTATGC | 400945 | 5.5723621842931506 | TruSeq Adapter, Index 5 (100% over 50bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
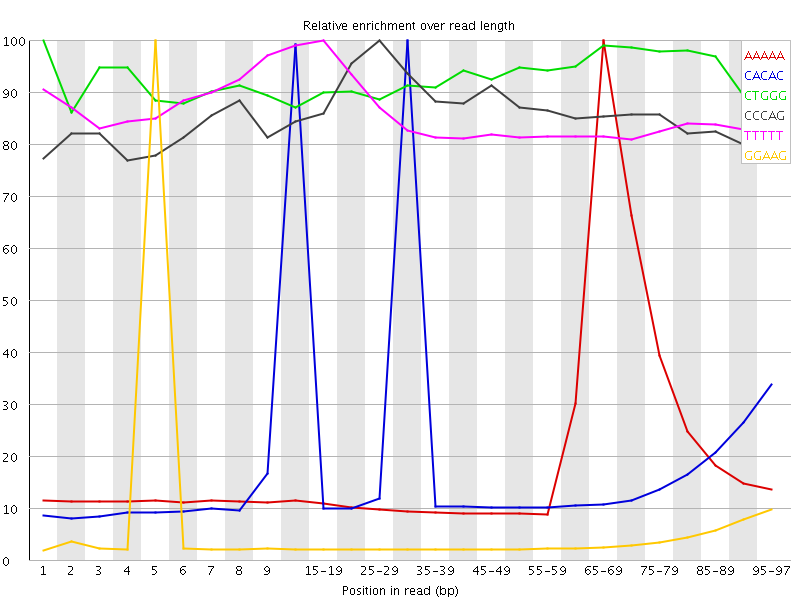
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 9221355 | 5.562278 | 25.595173 | 65-69 |
| CACAC | 2113175 | 3.6893888 | 16.536957 | 30-34 |
| CTGGG | 1311070 | 3.4488678 | 3.7018197 | 1 |
| CCCAG | 1354650 | 3.369331 | 3.8836675 | 25-29 |
| TTTTT | 4147135 | 3.3222866 | 3.8904967 | 15-19 |
| GGAAG | 1840365 | 3.2096288 | 75.78967 | 5 |
| CCTCC | 1169910 | 3.080855 | 3.5426738 | 65-69 |
| CCTGG | 1159160 | 3.050353 | 3.2909825 | 65-69 |
| GGAGG | 1223000 | 3.0386095 | 3.3797126 | 4 |
| CTCCA | 1477950 | 2.7310202 | 17.521141 | 20-24 |
| CAGTG | 1474350 | 2.722411 | 17.481146 | 35-39 |
| TCCAG | 1450160 | 2.678706 | 17.203089 | 25-29 |
| CTGCT | 1337880 | 2.6156068 | 18.43888 | 55-59 |
| CACAG | 1470190 | 2.56588 | 16.826908 | 30-34 |
| TCTGC | 1283750 | 2.5097806 | 18.289454 | 55-59 |
| AGAGC | 1428885 | 2.4928958 | 74.88423 | 8 |
| GAGCA | 1376160 | 2.4009094 | 74.72939 | 9 |
| TTCTG | 1650410 | 2.3962917 | 13.908242 | 55-59 |
| GCACA | 1359360 | 2.3724518 | 15.973051 | 10-14 |
| AGCAC | 1353115 | 2.3615525 | 15.862683 | 10-14 |
| GAAGA | 1900155 | 2.3261743 | 53.175755 | 6 |
| CGTCT | 1178045 | 2.3031232 | 16.86976 | 50-54 |
| CTTCT | 1564840 | 2.2728658 | 14.08051 | 50-54 |
| AAGAG | 1854720 | 2.2705526 | 53.10785 | 7 |
| GTCTG | 1150295 | 2.2480633 | 17.741152 | 15-19 |
| ACTCC | 1201120 | 2.2194817 | 17.15015 | 20-24 |
| ACACA | 1802940 | 2.2087498 | 11.962922 | 30-34 |
| TCTTC | 1443430 | 2.0965233 | 13.968934 | 50-54 |
| CCAGT | 1129480 | 2.086352 | 16.822737 | 25-29 |
| GATCG | 1122380 | 2.0724926 | 78.83819 | 1 |
| TGCTT | 1396940 | 2.0282693 | 13.984995 | 55-59 |
| TCTGA | 1447055 | 1.9851197 | 13.051907 | 15-19 |
| GCTTG | 1010485 | 1.9748274 | 17.929356 | 55-59 |
| ATGCC | 1062995 | 1.9635426 | 16.837372 | 45-49 |
| CTGAA | 1498825 | 1.9427027 | 12.321903 | 15-19 |
| ATCGG | 1040405 | 1.9211245 | 78.74357 | 2 |
| GAAAA | 2227475 | 1.9141166 | 8.75548 | 60-64 |
| TCGGA | 1008845 | 1.8628483 | 78.665665 | 3 |
| AGTGA | 1424420 | 1.8455995 | 12.147492 | 35-39 |
| CAGTC | 998600 | 1.8445935 | 16.63265 | 25-29 |
| TCACA | 1399345 | 1.8144137 | 12.065857 | 30-34 |
| CGGAA | 1002745 | 1.7494332 | 74.340614 | 4 |
| CTTGA | 1275195 | 1.7493563 | 12.672821 | 60-64 |
| ATCTC | 1274720 | 1.749333 | 12.421955 | 40-44 |
| GTCAC | 932010 | 1.7215899 | 16.743193 | 25-29 |
| GTCTT | 1175375 | 1.7065707 | 13.436787 | 50-54 |
| TGAAA | 1875810 | 1.7060454 | 8.940398 | 60-64 |
| GTGAT | 1163795 | 1.5959601 | 12.532661 | 35-39 |
| AACTC | 1222730 | 1.5854117 | 11.910497 | 20-24 |
| TGAAC | 1218815 | 1.5797677 | 11.764915 | 20-24 |
| AGATC | 1212990 | 1.5722177 | 6.64527 | 95-97 |
| ACAGT | 1203705 | 1.5601829 | 12.023053 | 30-34 |
| TTGAA | 1594385 | 1.5347625 | 9.265241 | 60-64 |
| GAACT | 1175650 | 1.5238194 | 11.831293 | 20-24 |
| CCGTC | 574140 | 1.511404 | 22.242311 | 50-54 |
| AGTCA | 1144335 | 1.4832305 | 12.067745 | 25-29 |
| TGCCG | 557170 | 1.4662043 | 23.05722 | 45-49 |
| CACGT | 786720 | 1.453213 | 16.162943 | 10-14 |
| TGATC | 1045025 | 1.4336011 | 12.388545 | 35-39 |
| GCCGT | 534410 | 1.4063107 | 22.73435 | 45-49 |
| GATCT | 1024785 | 1.4058352 | 12.313768 | 35-39 |
| ACACG | 795180 | 1.3878046 | 15.298661 | 10-14 |
| TCTCG | 645105 | 1.2612052 | 16.682608 | 40-44 |
| ACGTC | 682410 | 1.2605338 | 15.743968 | 15-19 |
| TATGC | 893045 | 1.2251097 | 12.258097 | 45-49 |
| GTATG | 889175 | 1.2193625 | 12.076926 | 45-49 |
| CTCGT | 560720 | 1.0962292 | 16.57642 | 40-44 |
| CGTAT | 543080 | 0.7450157 | 11.864116 | 40-44 |
| TCGTA | 518535 | 0.7113441 | 11.950204 | 40-44 |