![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL3_ACAGTG_L007_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 7569902 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
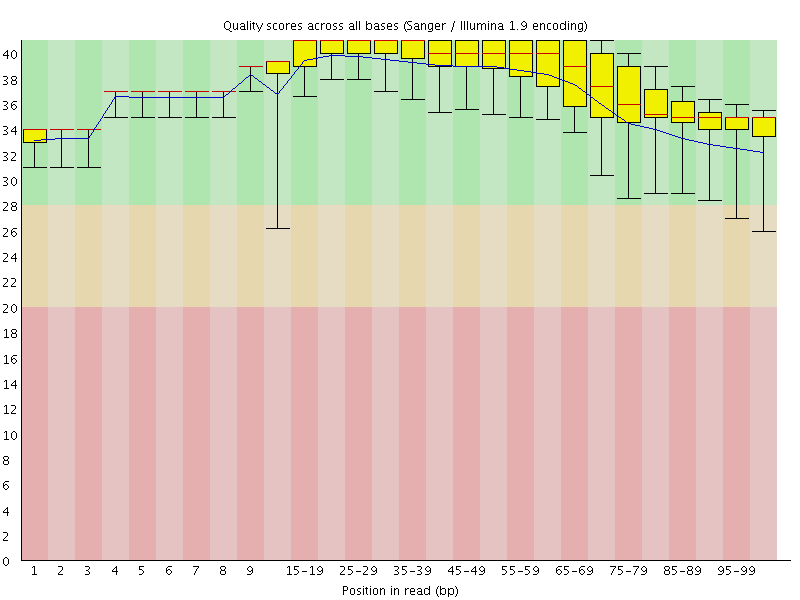
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
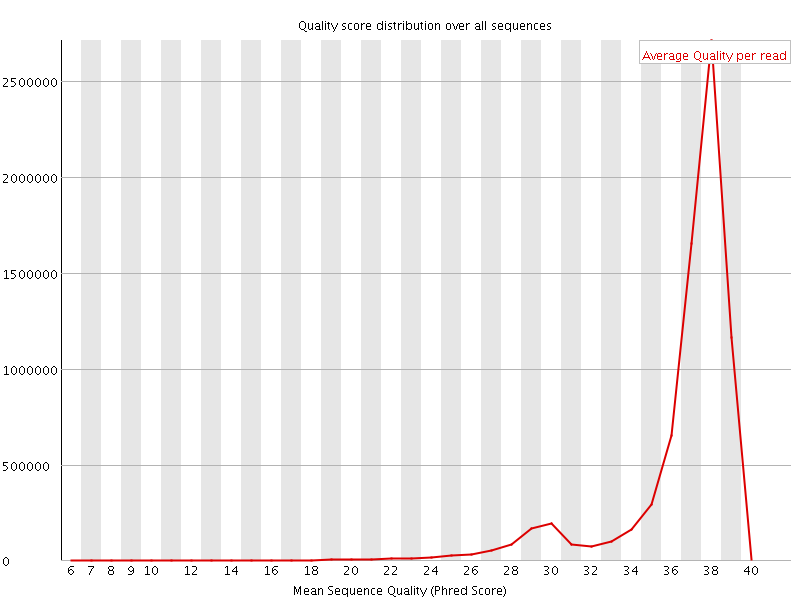
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
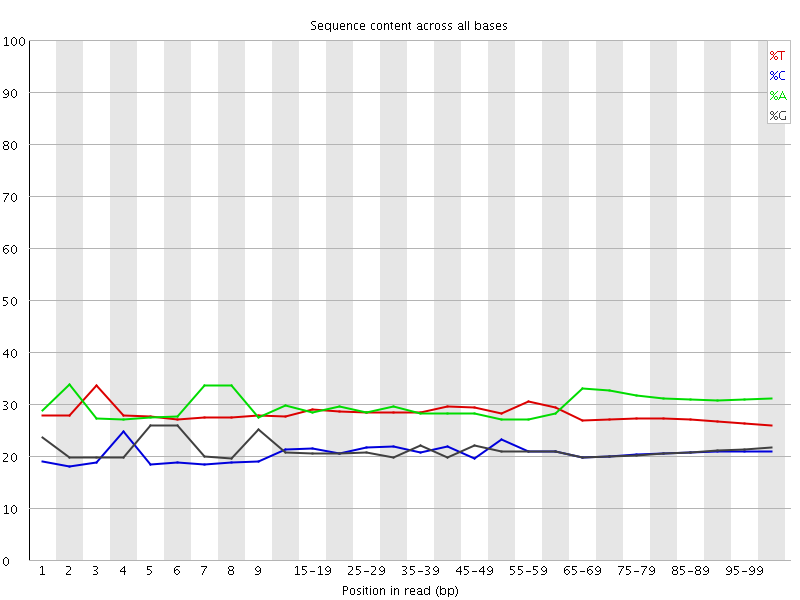
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
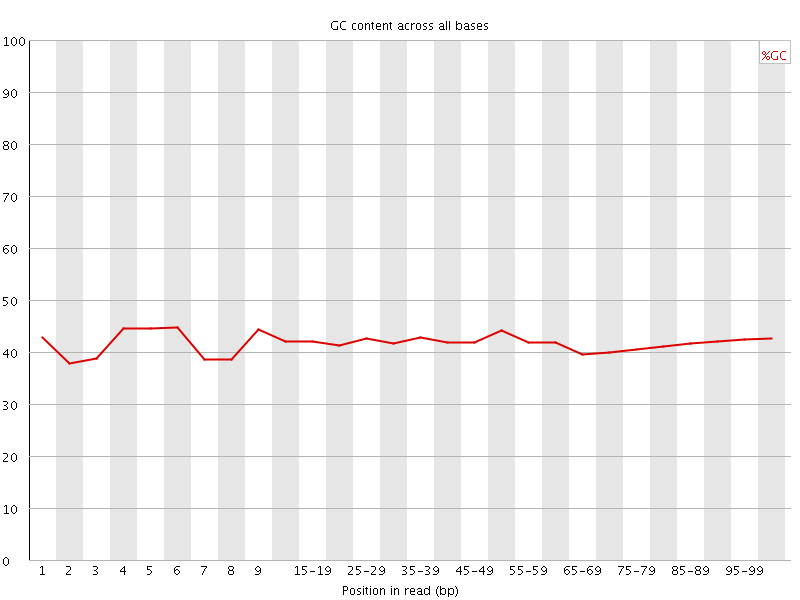
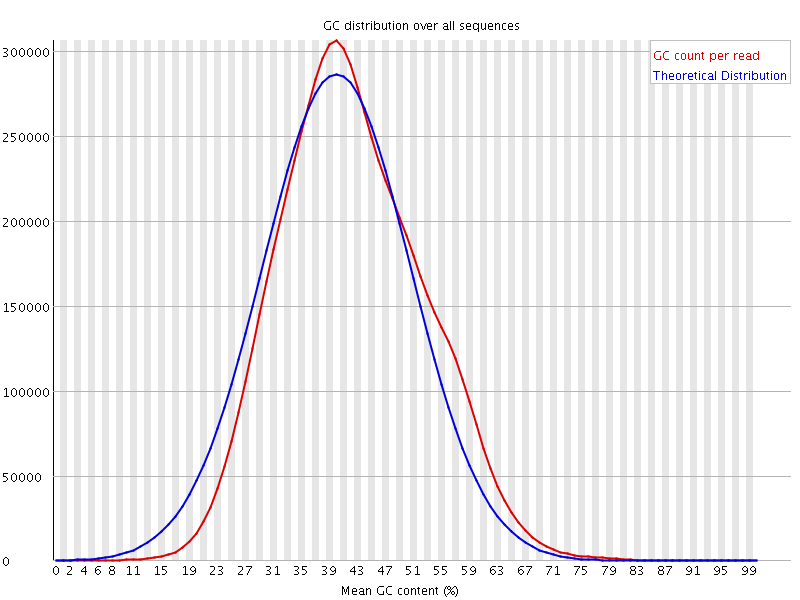
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
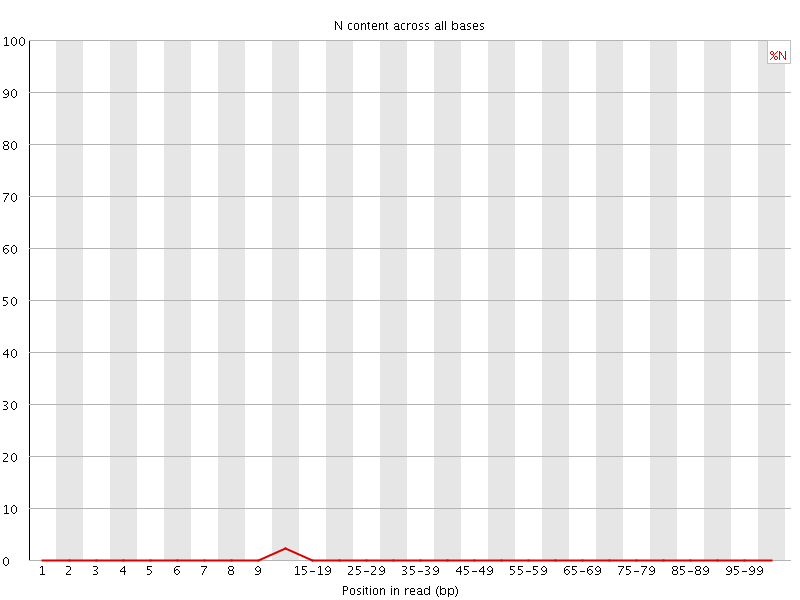
![[OK]](Icons/tick.png) Per base N content
Per base N content
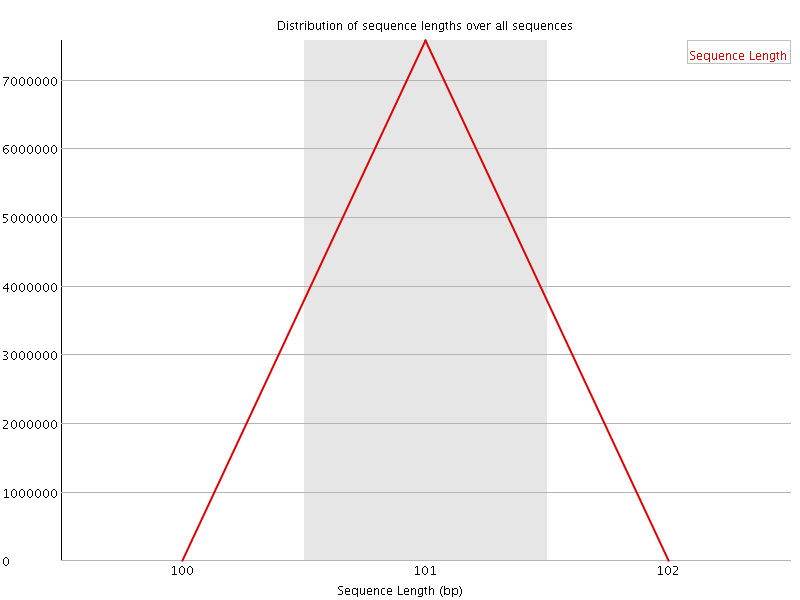
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
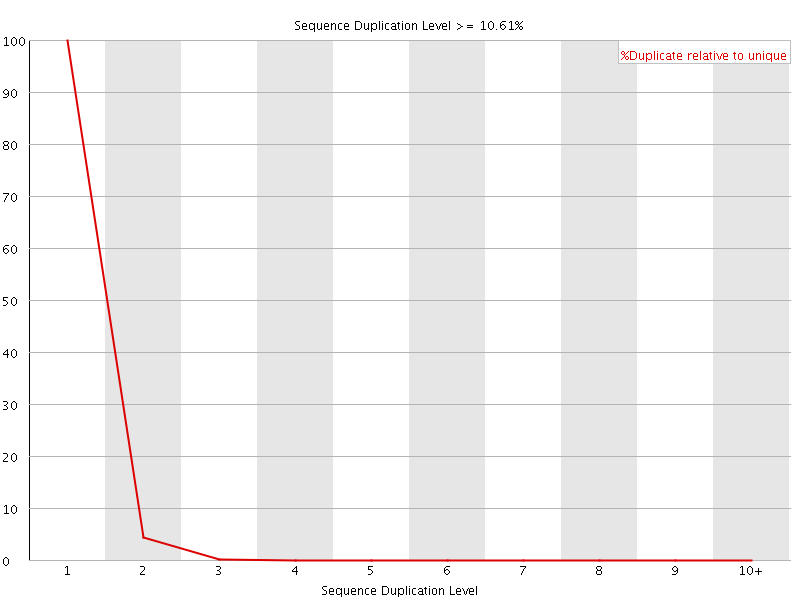
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACACAGTGATCTCGTATGC | 350560 | 4.6309714445444605 | TruSeq Adapter, Index 5 (100% over 50bp) |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
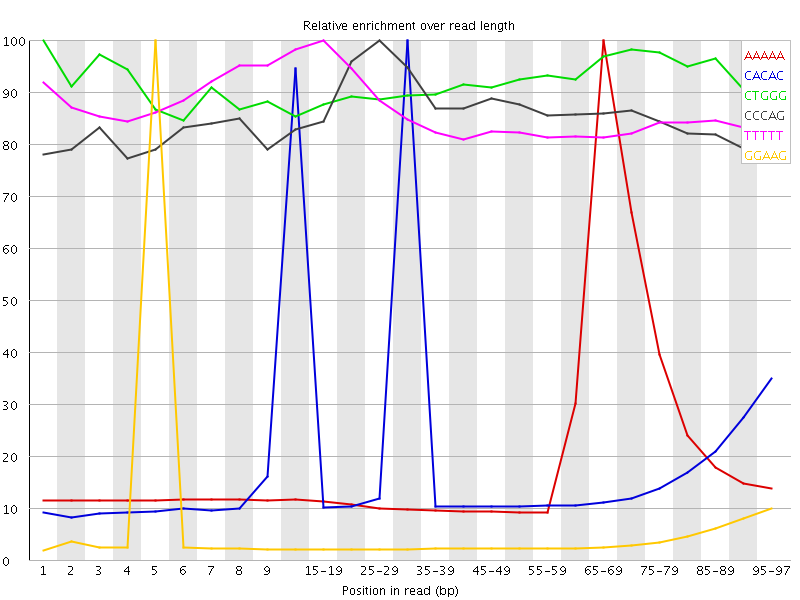
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 9595090 | 5.545954 | 25.236393 | 65-69 |
| CACAC | 2112150 | 3.535154 | 16.084497 | 30-34 |
| CTGGG | 1375710 | 3.4542356 | 3.754671 | 1 |
| CCCAG | 1422475 | 3.3878257 | 3.9245925 | 25-29 |
| TTTTT | 4314135 | 3.3023522 | 3.842875 | 15-19 |
| GGAAG | 1919805 | 3.1971486 | 73.42201 | 5 |
| CCTCC | 1221310 | 3.0819728 | 3.5783658 | 65-69 |
| CCTGG | 1213825 | 3.052861 | 3.3213785 | 70-74 |
| GGAGG | 1280280 | 3.0339155 | 3.5129707 | 4 |
| CCAGG | 1262740 | 3.0023715 | 3.4291997 | 25-29 |
| CTCCA | 1535160 | 2.7179265 | 17.09448 | 20-24 |
| CAGTG | 1529875 | 2.6995296 | 16.979504 | 35-39 |
| TCCAG | 1507250 | 2.6640565 | 16.739096 | 25-29 |
| CTGCT | 1384515 | 2.5885484 | 17.886635 | 55-59 |
| CACAG | 1527460 | 2.5522752 | 16.420498 | 30-34 |
| TCTGC | 1334150 | 2.494384 | 17.843924 | 55-59 |
| AGAGC | 1489515 | 2.4847147 | 72.6573 | 8 |
| TTCTG | 1719855 | 2.3863387 | 13.576778 | 55-59 |
| GAAGA | 1983605 | 2.3214965 | 51.57212 | 6 |
| GAGCA | 1369720 | 2.2848804 | 70.697914 | 9 |
| AAGAG | 1942405 | 2.2732785 | 51.558025 | 7 |
| CGTCT | 1211495 | 2.2650628 | 16.380444 | 15-19 |
| CTTCT | 1627500 | 2.2619722 | 13.719316 | 50-54 |
| GCACA | 1347450 | 2.2514913 | 14.7907715 | 10-14 |
| AGCAC | 1343045 | 2.244131 | 14.673268 | 10-14 |
| GTCTG | 1195555 | 2.2315273 | 17.233383 | 15-19 |
| ACTCC | 1246265 | 2.206452 | 16.740723 | 20-24 |
| ACACA | 1861845 | 2.186293 | 11.629201 | 30-34 |
| TCTTC | 1499120 | 2.0835443 | 13.663856 | 50-54 |
| CCAGT | 1169715 | 2.067465 | 16.338396 | 25-29 |
| GATCG | 1170695 | 2.065741 | 76.44713 | 1 |
| TGCTT | 1445980 | 2.006331 | 13.598981 | 55-59 |
| TCTGA | 1506360 | 1.9759165 | 12.698093 | 15-19 |
| GCTTG | 1048210 | 1.956505 | 17.391693 | 55-59 |
| ATGCC | 1099335 | 1.9430686 | 16.389793 | 45-49 |
| CTGAA | 1565805 | 1.9416771 | 12.0049305 | 15-19 |
| ATCGG | 1084290 | 1.913276 | 76.329956 | 2 |
| GAAAA | 2324490 | 1.9118232 | 8.564081 | 60-64 |
| TCGGA | 1050685 | 1.8539784 | 76.306145 | 3 |
| CAGTC | 1037750 | 1.8342175 | 16.180977 | 25-29 |
| AGTGA | 1475570 | 1.8267251 | 11.828703 | 35-39 |
| TCACA | 1451725 | 1.803224 | 11.760478 | 30-34 |
| CGGAA | 1045545 | 1.7441121 | 72.11143 | 4 |
| CTTGA | 1328520 | 1.742641 | 12.382197 | 60-64 |
| ATCTC | 1326300 | 1.7426394 | 12.112216 | 40-44 |
| TGAAA | 1957585 | 1.703104 | 8.730381 | 60-64 |
| GTCAC | 959270 | 1.6955045 | 16.231236 | 25-29 |
| GTCTT | 1220950 | 1.6940967 | 13.044549 | 50-54 |
| AGATC | 1275995 | 1.5822979 | 6.673619 | 95-97 |
| GTGAT | 1203440 | 1.575935 | 12.162173 | 35-39 |
| AACTC | 1268665 | 1.5758404 | 11.640543 | 20-24 |
| TGAAC | 1264915 | 1.5685582 | 11.446237 | 20-24 |
| ACAGT | 1249525 | 1.5494739 | 11.720994 | 30-34 |
| TTGAA | 1654610 | 1.5227079 | 9.03508 | 60-64 |
| GAACT | 1220725 | 1.5137602 | 11.506356 | 20-24 |
| CCGTC | 589360 | 1.4847646 | 21.631945 | 50-54 |
| AGTCA | 1190570 | 1.4763665 | 11.735077 | 25-29 |
| TGCCG | 571255 | 1.4367491 | 22.383108 | 45-49 |
| CACGT | 812025 | 1.4352498 | 17.292908 | 10-14 |
| TGATC | 1084400 | 1.4224249 | 12.052422 | 35-39 |
| GATCT | 1059545 | 1.389822 | 11.94141 | 35-39 |
| GCCGT | 547810 | 1.3777832 | 22.0328 | 45-49 |
| ACACG | 757815 | 1.2662541 | 14.089312 | 10-14 |
| TCTCG | 665050 | 1.2434058 | 16.208353 | 40-44 |
| ACGTC | 702620 | 1.2418771 | 15.294006 | 15-19 |
| TATGC | 922065 | 1.2094874 | 11.914961 | 45-49 |
| GTATG | 918515 | 1.2028185 | 11.706606 | 45-49 |
| CTCGT | 576480 | 1.0778116 | 16.107975 | 40-44 |
| CGTAT | 557330 | 0.73105866 | 11.524421 | 40-44 |
| TCGTA | 532930 | 0.6990528 | 11.607236 | 40-44 |