![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL2_TGACCA_L008_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 11193634 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
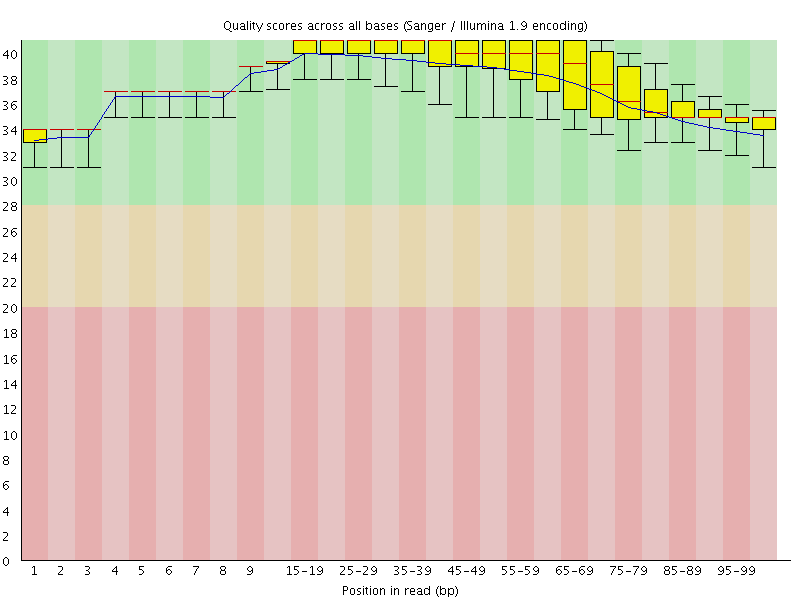
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
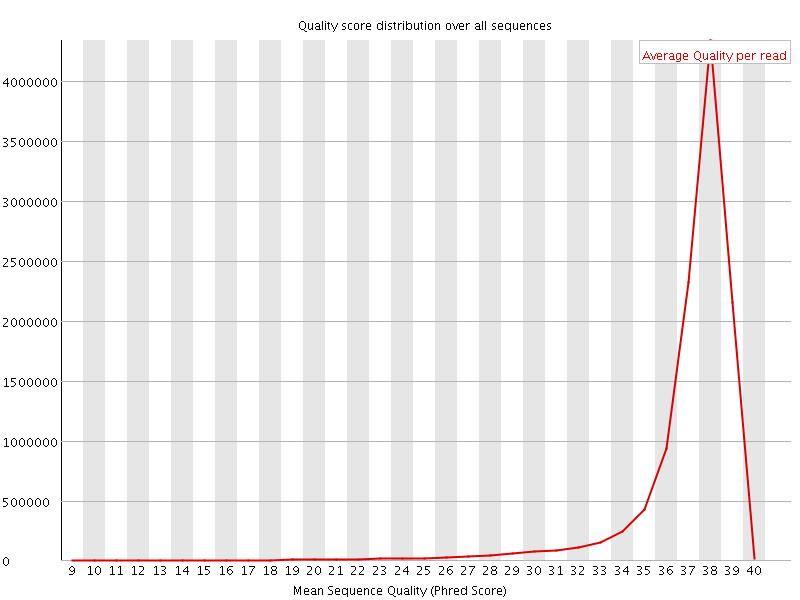
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
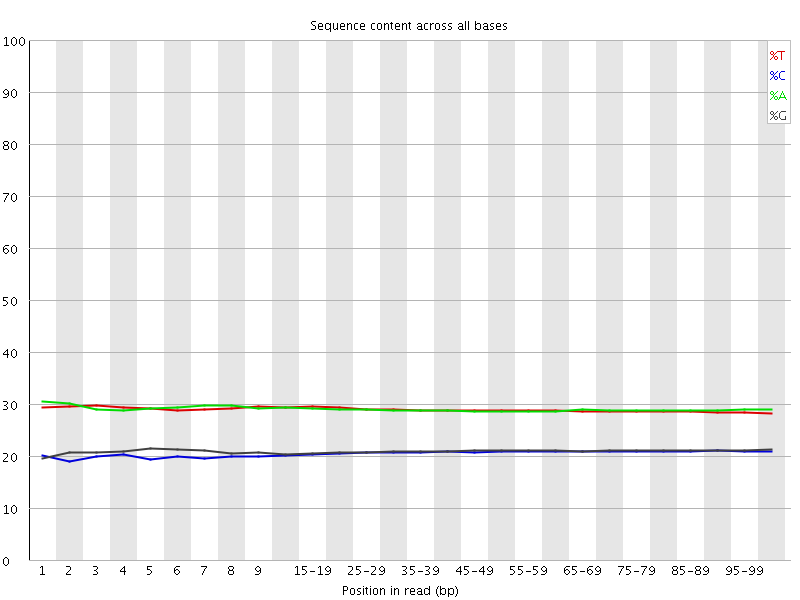
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
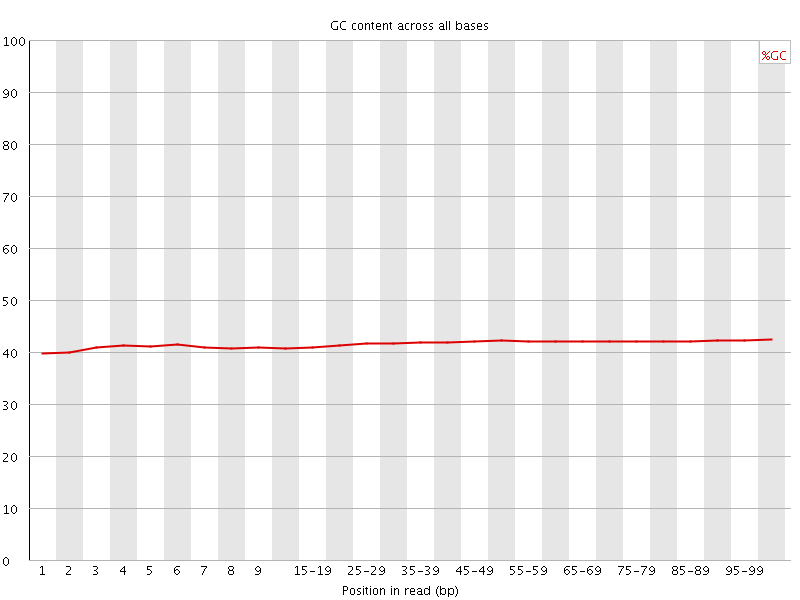
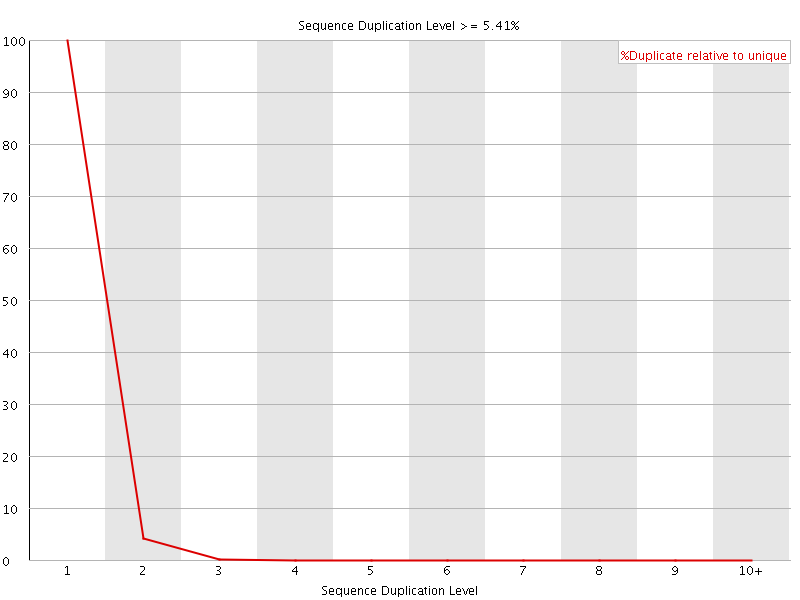
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
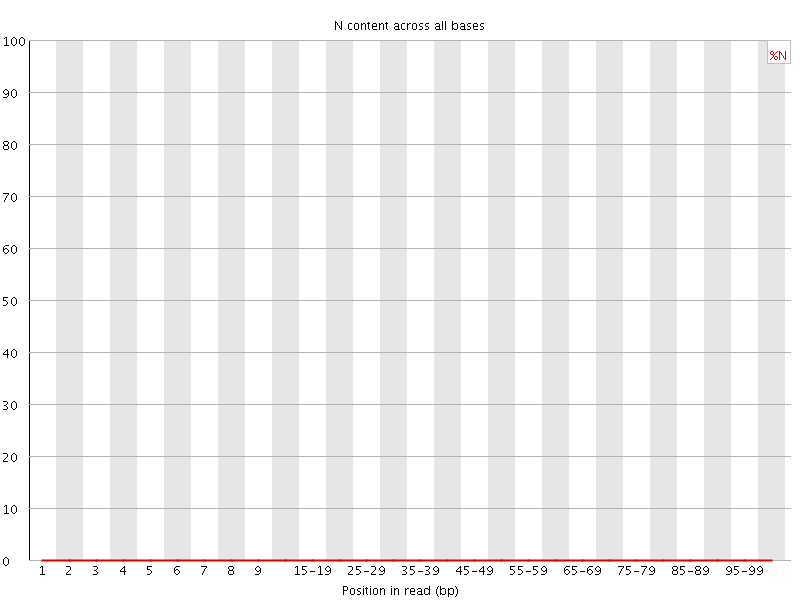
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
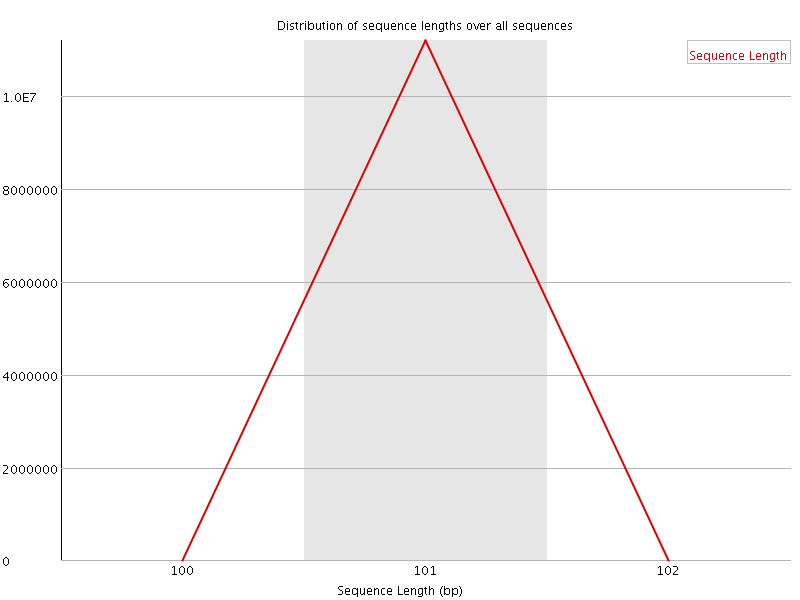
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTGACCAATCTCGTATGC | 32852 | 0.2934882451936521 | TruSeq Adapter, Index 4 (100% over 50bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
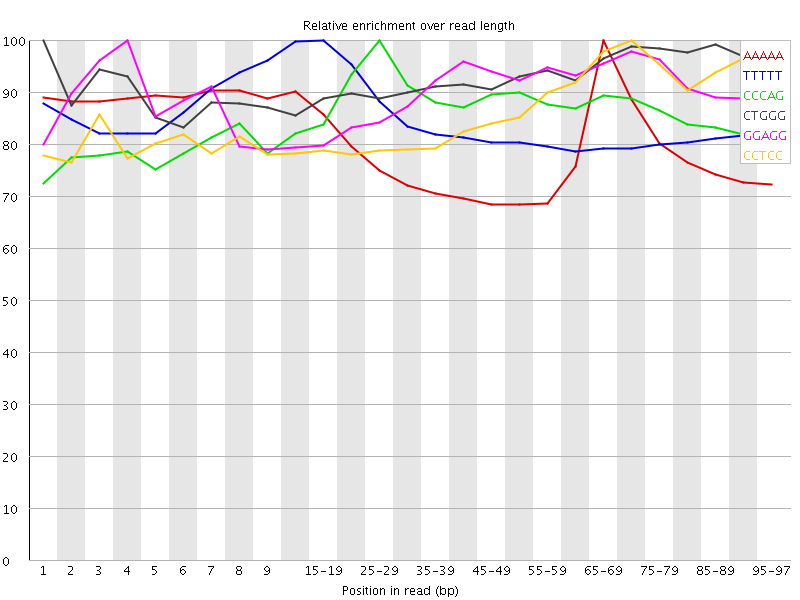
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 8209875 | 3.6412446 | 4.6408796 | 65-69 |
| TTTTT | 7719200 | 3.470884 | 4.1076446 | 15-19 |
| CCCAG | 2081615 | 3.4463112 | 3.965414 | 25-29 |
| CTGGG | 2003830 | 3.2613993 | 3.5065513 | 1 |
| GGAGG | 2023650 | 3.2522707 | 3.6093037 | 4 |
| CCTCC | 1925065 | 3.2276862 | 3.722181 | 70-74 |
| CCAGG | 1868475 | 3.0629532 | 3.4654157 | 25-29 |
| GGGGG | 1356830 | 3.0085773 | 3.507695 | 70-74 |
| GGAAG | 1799145 | 2.0957184 | 6.24654 | 5 |
| GAGCA | 1185110 | 1.3942043 | 5.4666204 | 9 |
| AGAGC | 1176970 | 1.384628 | 5.4569216 | 8 |