![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | ZL2_TGACCA_L007_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 11447782 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
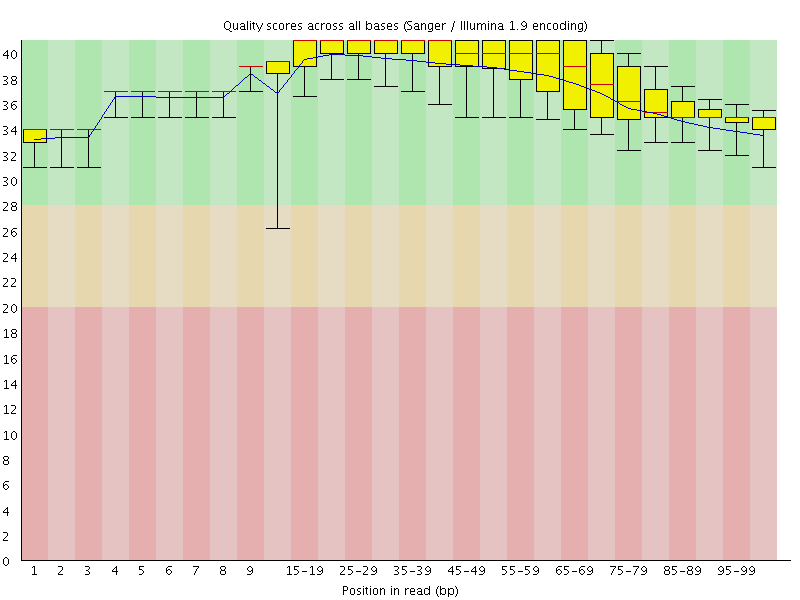
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
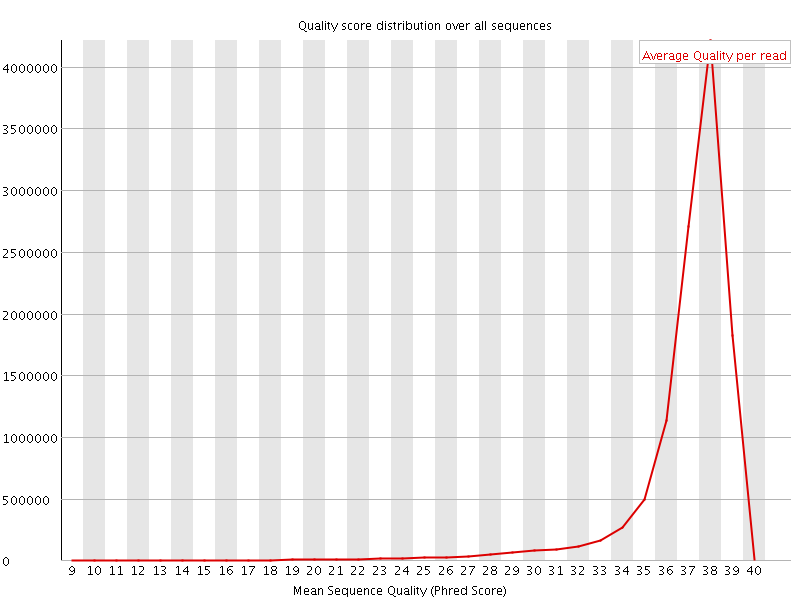
![[WARN]](Icons/warning.png) Per base sequence content
Per base sequence content
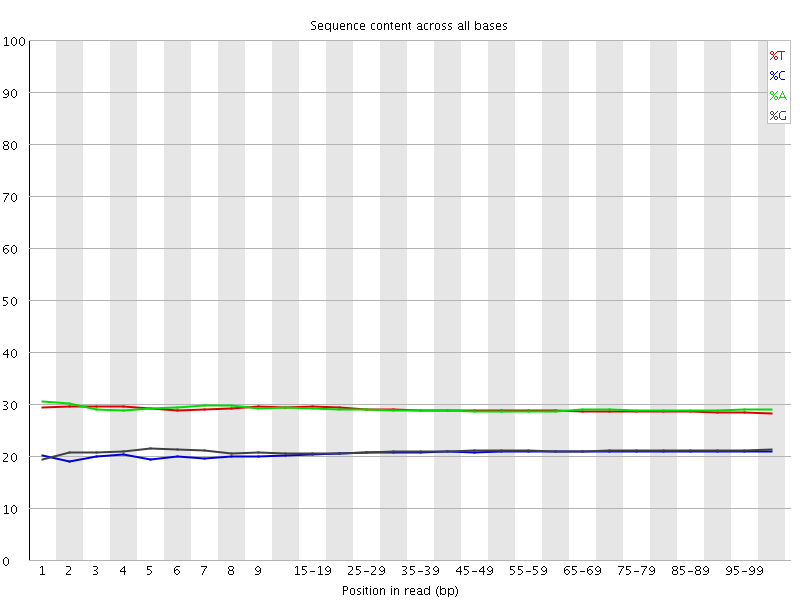
![[OK]](Icons/tick.png) Per base GC content
Per base GC content
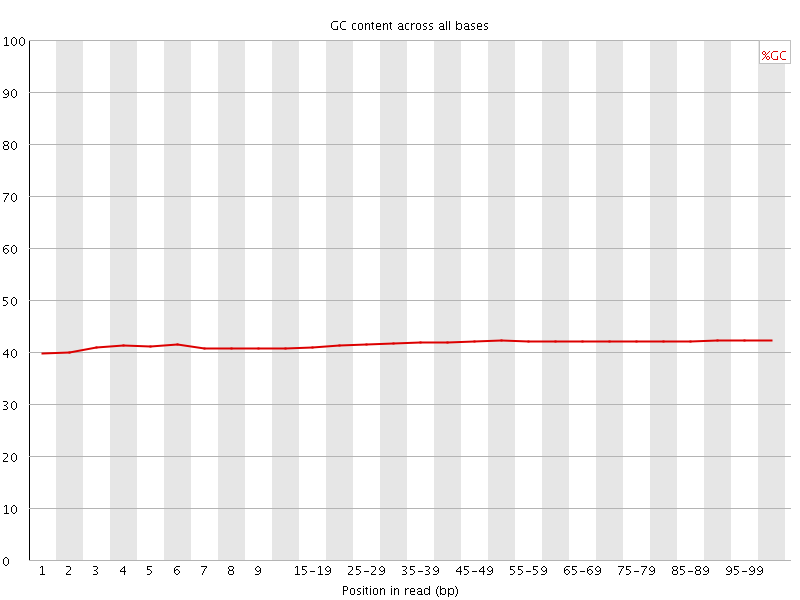
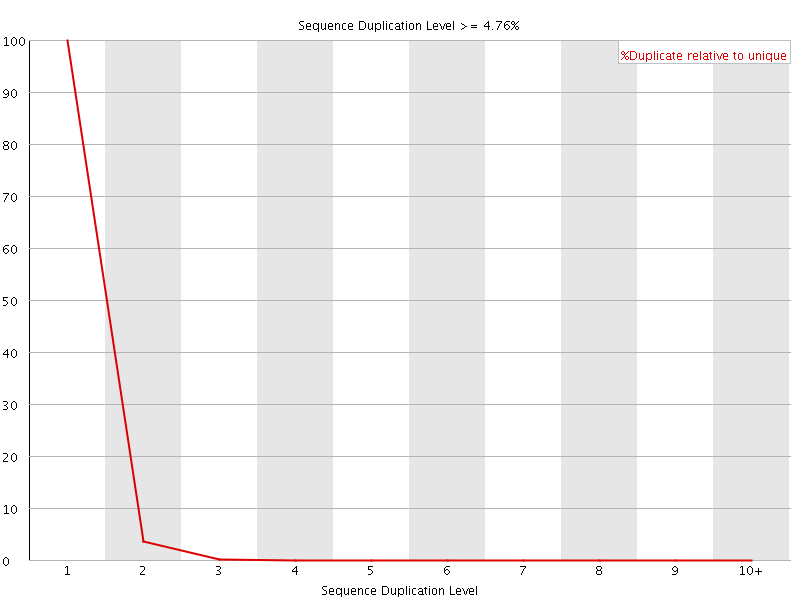
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
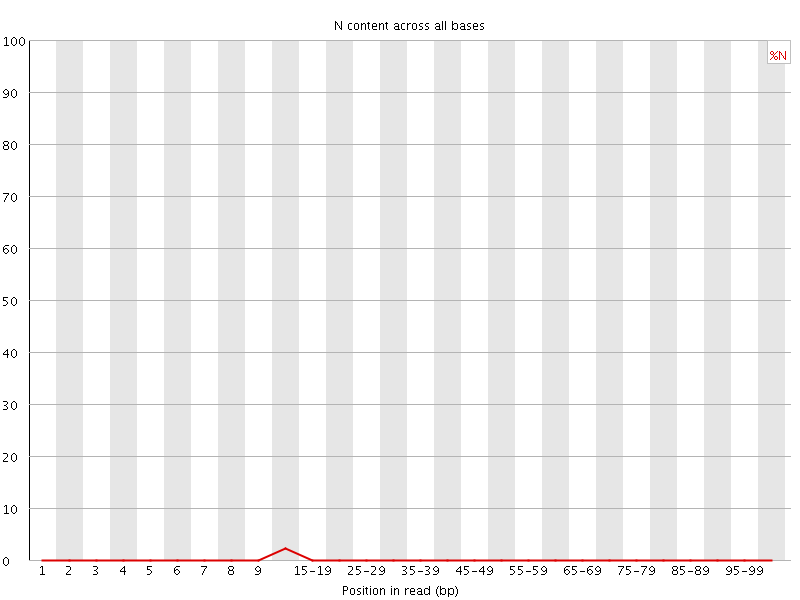
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
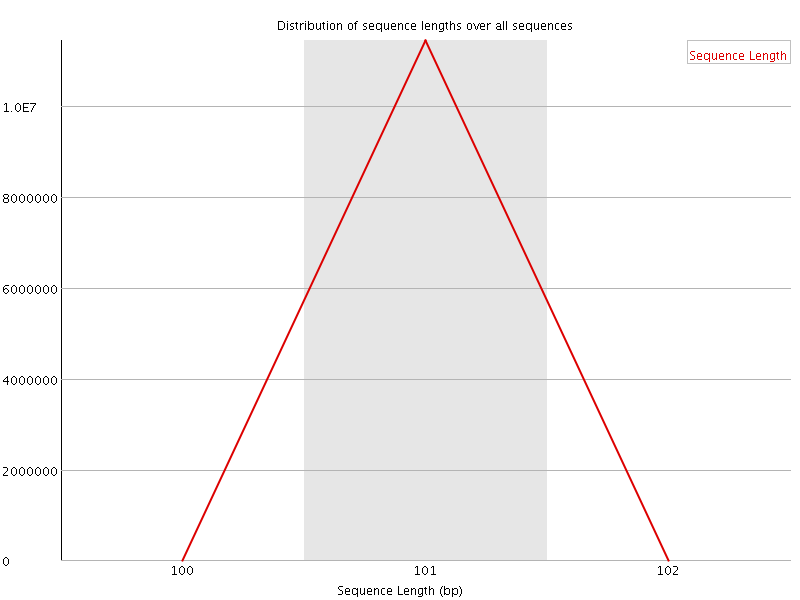
![[OK]](Icons/tick.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTGACCAATCTCGTATGC | 28239 | 0.246676605127526 | TruSeq Adapter, Index 4 (100% over 50bp) |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
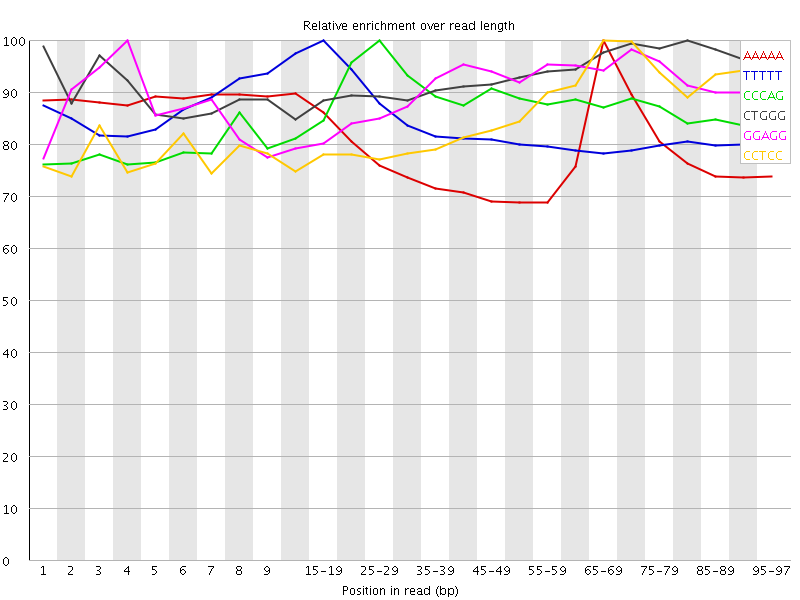
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AAAAA | 8310655 | 3.6216133 | 4.5918503 | 65-69 |
| TTTTT | 7863115 | 3.4696789 | 4.135772 | 15-19 |
| CCCAG | 2114275 | 3.451042 | 3.9443674 | 25-29 |
| CTGGG | 2034810 | 3.26339 | 3.503137 | 80-84 |
| GGAGG | 2046050 | 3.2404935 | 3.587519 | 4 |
| CCTCC | 1948070 | 3.2199082 | 3.7525291 | 65-69 |
| CCAGG | 1891995 | 3.0573428 | 3.4027522 | 25-29 |
| GGGGG | 1378510 | 3.014476 | 3.5231748 | 70-74 |
| GGAAG | 1829395 | 2.098434 | 6.130299 | 5 |
| GAGCA | 1203830 | 1.3948183 | 5.2769976 | 9 |
| AGAGC | 1201910 | 1.3925937 | 5.357962 | 8 |