![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SDW_HiSeq_6_GTGAAA_L006_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 22169699 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 46 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
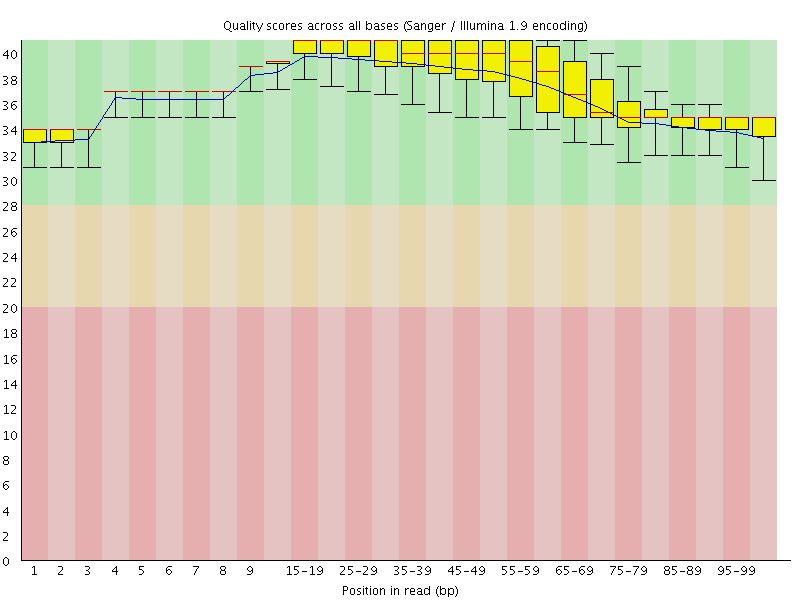
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
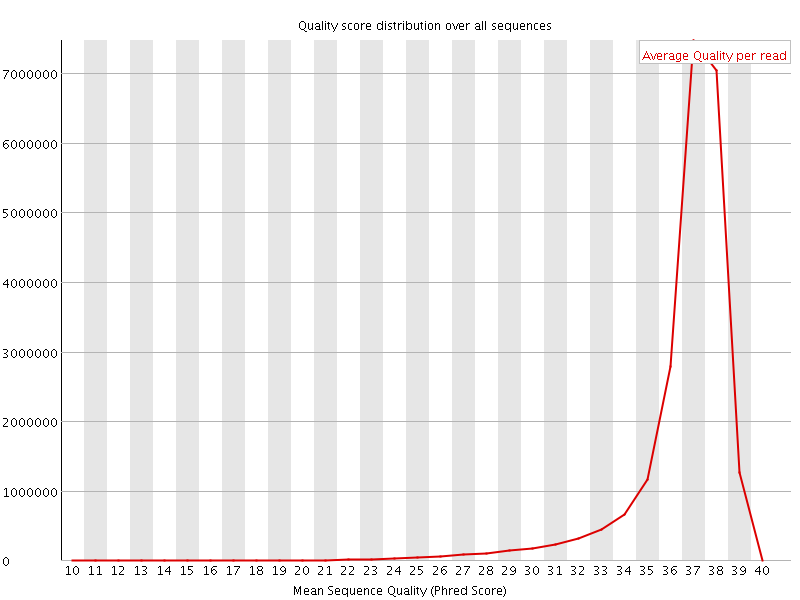
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content

![[FAIL]](Icons/error.png) Per base GC content
Per base GC content

![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
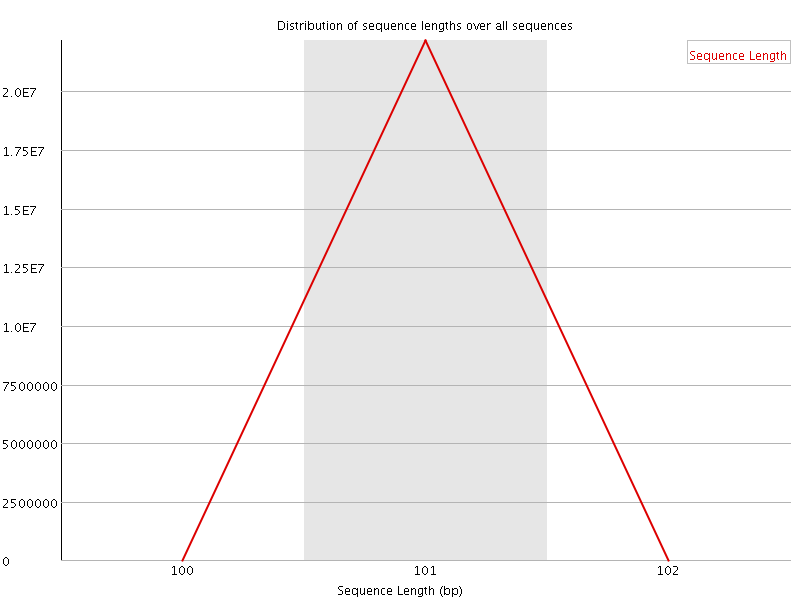
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
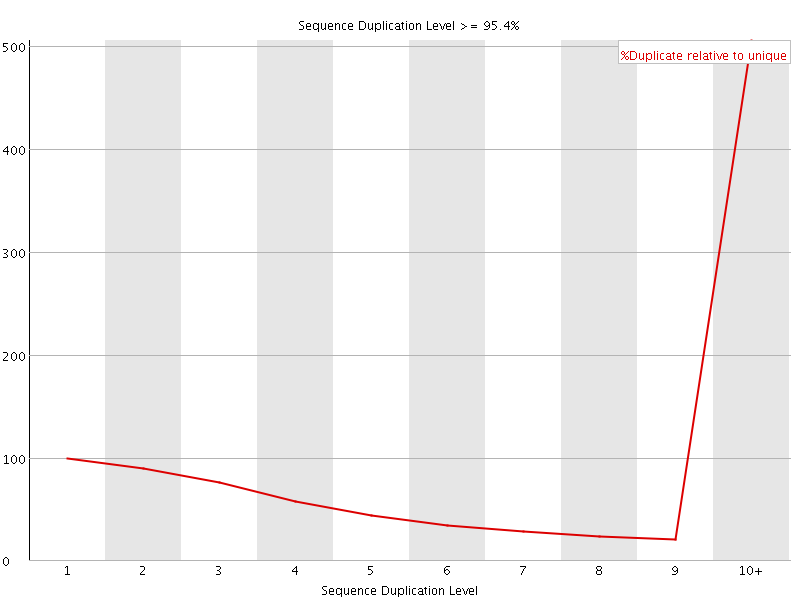
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CGATAATCTCGCTCGTTAGCTGAATCAACGCTAGACAGGTCAACCCACGC | 66060 | 0.2979742756092449 | No Hit |
| CCCGTGATAGCCTACCTAGGTAAAGACCCTAAGTACAAGCTGATAGGTCA | 29712 | 0.13402076410690106 | No Hit |
| AGCAAACTCGCAGAGCTGGAATTACCGCAGCTGCTGGCACCAGACTGGCC | 26866 | 0.12118342247226721 | No Hit |
| GCCAGATCTTTTCCATATCATCCCAGTTGTTGACGATACCGTGTTCAATT | 26482 | 0.11945132859043327 | No Hit |
| GTTATTGCTACCCTCTTCCACCTGCTAAAATCGCAGGGAGTCCCAATTAG | 25540 | 0.1152022857865594 | No Hit |
| CCCTGGTGTGCCCTTCCGTCAATTCCTTCAAGTTTCAGCCTTGCGGTCGT | 24740 | 0.11159375686607204 | No Hit |
| GTAATCTTGAAGTGTGTTGCCGTCTTCAAGTTGCTTGCCGGCAAAGATCA | 23799 | 0.10734922472334875 | No Hit |
| CCTGGTTGCAGATGTACTTCTCGCCACCATAGTATAAGCCAGAGAGAGCT | 23432 | 0.10569381208107517 | No Hit |
| CCCGTACGGTCCTCCATCAGAGTTTCCCCTGACTTCAACCTGTGCAGGCA | 23288 | 0.10504427687538744 | No Hit |
| GCGATAATCTCGCTCGTTAGCTGAATCAACGCTAGACAGGTCAACCCACG | 23124 | 0.10430452844668753 | No Hit |
| CTTCCGTCAATTCCTTCAAGTTTCAGCCTTGCGGTCGTAGTTCCCCCAGA | 22663 | 0.1022251136562567 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
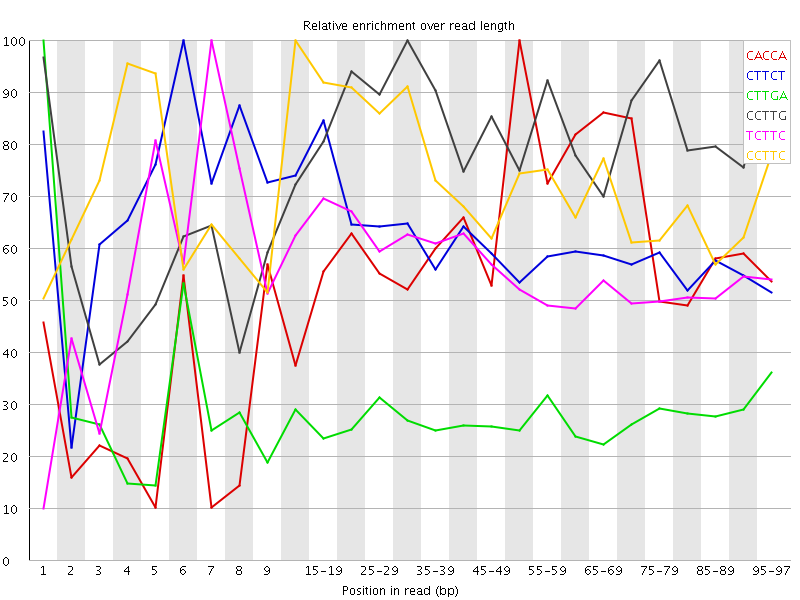
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACCA | 6048065 | 4.263828 | 7.083074 | 50-54 |
| CTTCT | 10269965 | 4.040078 | 6.5173516 | 6 |
| CTTGA | 8162080 | 3.4163375 | 12.247324 | 1 |
| CCTTG | 7132915 | 3.3843212 | 4.1528144 | 30-34 |
| TCTTC | 7935355 | 3.121671 | 5.540113 | 7 |
| CCTTC | 5894805 | 3.062251 | 4.138061 | 10-14 |
| TTCTT | 10095275 | 3.0073812 | 5.66741 | 6 |
| CCACC | 3760725 | 3.00538 | 5.0897045 | 20-24 |
| GAAGA | 5108120 | 2.6501231 | 5.0486217 | 3 |
| TCTTG | 6821535 | 2.4509592 | 5.0871983 | 7 |
| CTGGA | 4854525 | 2.4507012 | 17.807636 | 1 |
| TGGAA | 5401615 | 2.405596 | 9.764945 | 2 |
| CAGCC | 3177715 | 2.3194015 | 5.5617166 | 9 |
| CAACA | 3643780 | 2.2661607 | 5.8012896 | 9 |
| CTTTG | 6111075 | 2.1956928 | 5.498669 | 1 |
| TGGAG | 4677835 | 2.1568592 | 6.874332 | 2 |
| CTGTG | 4951560 | 2.1457534 | 6.048252 | 1 |
| GGATG | 4393365 | 2.0256956 | 5.4841666 | 4 |
| TCAAC | 3792310 | 2.024587 | 5.721136 | 8 |
| TGAAG | 4507305 | 2.0073173 | 5.1839747 | 8 |
| CCTGG | 3496425 | 2.0008407 | 9.306797 | 1 |
| CTGGG | 3634330 | 1.8995285 | 19.08719 | 1 |
| TGGGA | 4098460 | 1.8897204 | 10.813383 | 2 |
| CTTGT | 5119260 | 1.8393364 | 5.8467193 | 1 |
| TTGAA | 4912405 | 1.8138839 | 6.5173826 | 7 |
| CTGCT | 3735415 | 1.7723252 | 5.8588576 | 1 |
| GTTGA | 4588900 | 1.7542912 | 5.7410307 | 1 |
| TGGAT | 4580545 | 1.7510973 | 5.5374093 | 2 |
| CTGGC | 2914115 | 1.6676118 | 7.725192 | 1 |
| GATGT | 4309260 | 1.6473877 | 5.311964 | 5 |
| GCAGA | 2708150 | 1.5926572 | 5.132653 | 8 |
| GTGGA | 3391295 | 1.5636604 | 6.298959 | 1 |
| CTGGT | 3585705 | 1.5538615 | 11.53027 | 1 |
| CTCTG | 3125450 | 1.4829178 | 5.8908806 | 1 |
| GGAAT | 3140320 | 1.3985337 | 6.5285316 | 3 |
| CTGAA | 2840230 | 1.3849028 | 6.2670727 | 1 |
| GGGAA | 2540295 | 1.3644792 | 6.067837 | 3 |
| GTTGG | 3335350 | 1.3201168 | 5.3113723 | 1 |
| CTGAG | 2538015 | 1.2812617 | 6.2717824 | 1 |
| GAATT | 3411255 | 1.2595909 | 5.368513 | 4 |
| GTGAA | 2818525 | 1.2552232 | 5.6309934 | 1 |
| AATCT | 3032885 | 1.2261343 | 5.341657 | 5 |
| TGGTT | 3513605 | 1.1530288 | 5.458057 | 2 |
| TGGGT | 2714175 | 1.0742584 | 5.844362 | 2 |
| GGGAT | 2206280 | 1.017273 | 5.393606 | 3 |
| CTCGC | 1586875 | 0.99425495 | 5.897894 | 8 |
| CGGGC | 522385 | 0.36054766 | 5.0403976 | 1 |