![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SDW_HiSeq_5_CTTGTA_L006_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 25658979 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 46 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
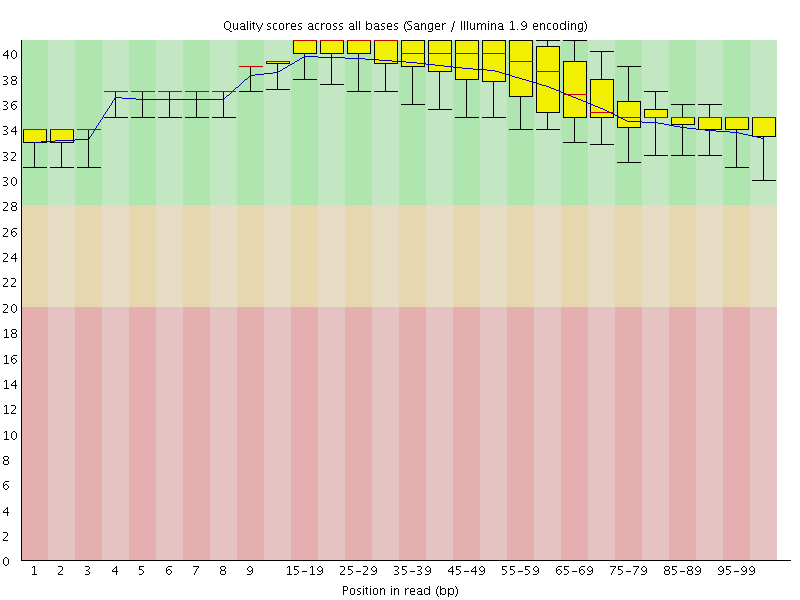
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
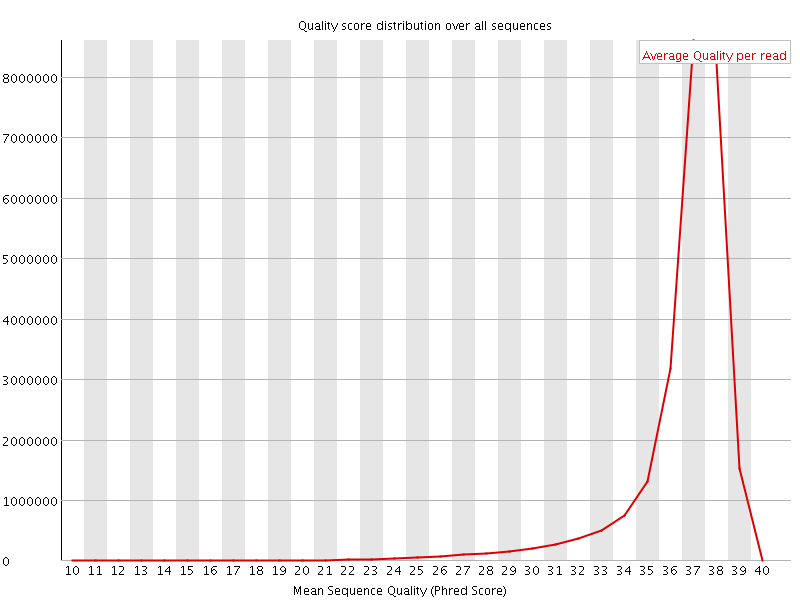
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content

![[FAIL]](Icons/error.png) Per base GC content
Per base GC content

![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
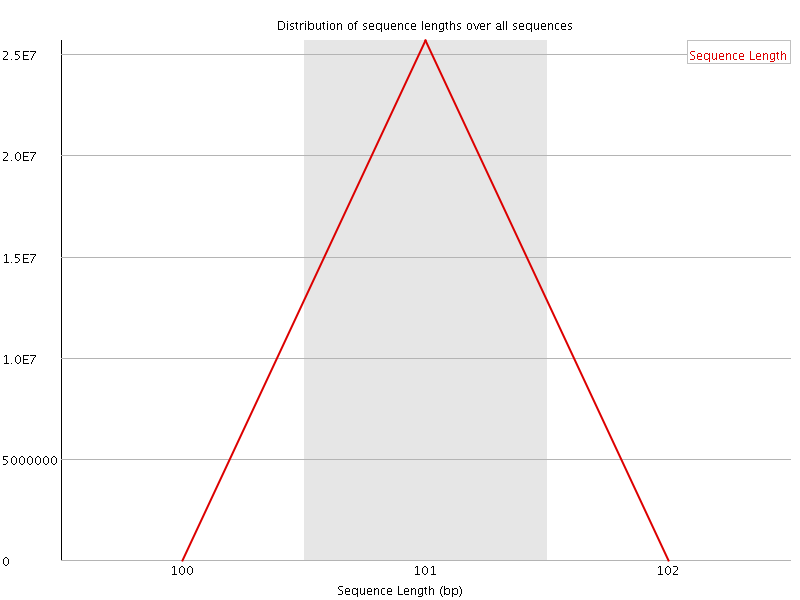
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
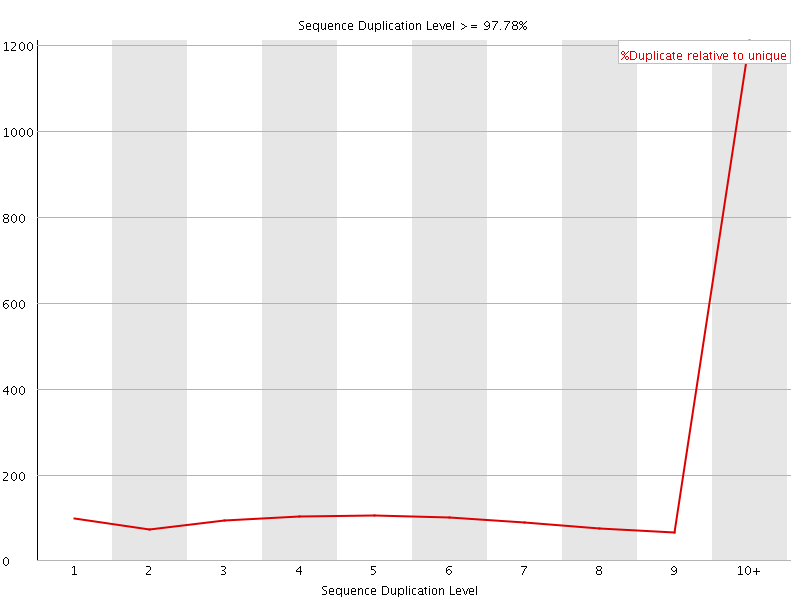
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CGATAATCTCGCTCGTTAGCTGAATCAACGCTAGACAGGTCAACCCACGC | 90950 | 0.3544568160720658 | No Hit |
| GTAATCTTGAAGTGTGTTGCCGTCTTCAAGTTGCTTGCCGGCAAAGATCA | 48999 | 0.190962391761574 | No Hit |
| GTAAACGTTTGTCTTGATGTCACCAAGATCGTGGATAGCGACAACTTGGA | 43179 | 0.1682802733499256 | No Hit |
| GCGATAATCTCGCTCGTTAGCTGAATCAACGCTAGACAGGTCAACCCACG | 39007 | 0.15202085788370612 | No Hit |
| CTGAAGAATAACTGGAGACTCTGTCTTGACGGAAGCTGTGATGATAGCCT | 36846 | 0.14359885481024012 | No Hit |
| GGCGATAATCTCGCTCGTTAGCTGAATCAACGCTAGACAGGTCAACCCAC | 31973 | 0.1246074522294905 | No Hit |
| GTTATTGCTACCCTCTTCCACCTGCTAAAATCGCAGGGAGTCCCAATTAG | 28718 | 0.11192183445802734 | No Hit |
| CTCGTTAGCTGAATCAACGCTAGACAGGTCAACCCACGCACCACCAACGG | 28076 | 0.10941978634457745 | No Hit |
| CCTGGTTGCAGATGTACTTCTCGCCACCATAGTATAAGCCAGAGAGAGCT | 27471 | 0.10706193726570336 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
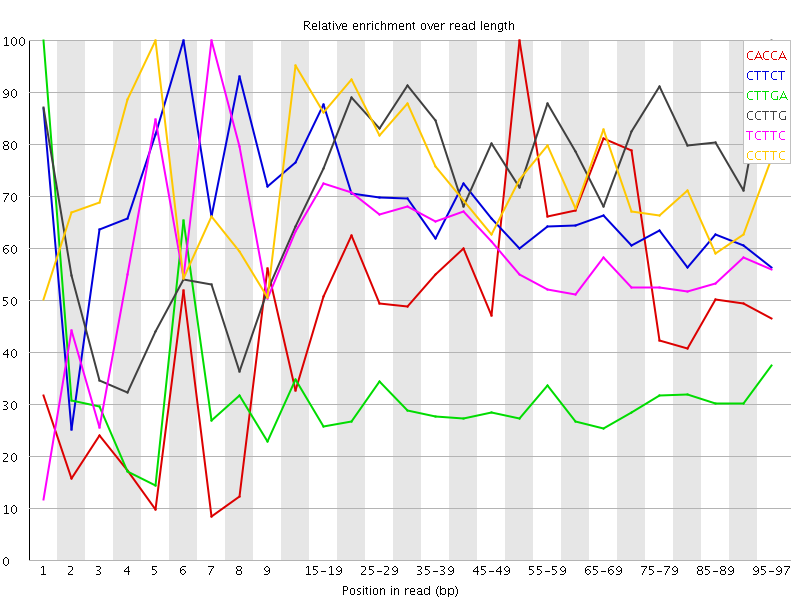
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CACCA | 7213270 | 4.4866686 | 8.231708 | 50-54 |
| CTTCT | 12080380 | 4.183248 | 6.245006 | 6 |
| CTTGA | 9892165 | 3.583408 | 11.748986 | 1 |
| CCTTG | 8496315 | 3.5183504 | 4.554038 | 95-97 |
| TCTTC | 9474030 | 3.2807095 | 5.50335 | 7 |
| CCTTC | 7003615 | 3.223126 | 4.3112516 | 5 |
| TTCTT | 12099675 | 3.1527154 | 5.914797 | 6 |
| CCACC | 4333925 | 3.0816033 | 5.2433457 | 20-24 |
| GCAGC | 5265715 | 3.0315154 | 4.2084274 | 8 |
| AGCCT | 6280460 | 3.0235586 | 4.036589 | 9 |
| GAAGA | 6094240 | 2.6848178 | 5.7956977 | 3 |
| TCTTG | 8402735 | 2.6182265 | 5.3192067 | 7 |
| CTGGA | 5598950 | 2.4254203 | 14.04073 | 1 |
| TGGAA | 6353410 | 2.4075968 | 8.481573 | 2 |
| CAGCC | 3746220 | 2.3968573 | 5.7927885 | 9 |
| CTTTG | 7484740 | 2.3321865 | 5.2091665 | 1 |
| CAACA | 4171610 | 2.2698228 | 5.2461925 | 9 |
| TGGAG | 5730200 | 2.2335913 | 7.2791834 | 2 |
| AAGAA | 5133770 | 2.1987436 | 5.281314 | 4 |
| CTGTG | 5853305 | 2.181038 | 6.5193834 | 1 |
| CCTGG | 4259105 | 2.1091251 | 9.361251 | 1 |
| TGAAG | 5522535 | 2.0927403 | 5.82835 | 8 |
| GGATG | 5278875 | 2.0576682 | 5.413212 | 4 |
| TCAAC | 4310230 | 2.017299 | 5.1998844 | 8 |
| CTGGG | 4487340 | 1.9995234 | 17.281458 | 1 |
| GCCTG | 3976330 | 1.9690939 | 5.3164825 | 1 |
| TGGGA | 5036120 | 1.963044 | 11.035883 | 2 |
| CTTGT | 6042680 | 1.8828518 | 6.2139025 | 1 |
| TTGAA | 5858270 | 1.8563966 | 7.086212 | 7 |
| TGGAT | 5551610 | 1.8095785 | 5.103028 | 2 |
| GTTGA | 5241595 | 1.7085274 | 5.779912 | 1 |
| CTGCT | 4092940 | 1.6948992 | 5.03572 | 1 |
| GATGT | 5084855 | 1.6574371 | 5.533782 | 5 |
| CTGGC | 3326440 | 1.6472657 | 6.9543943 | 1 |
| GTGGA | 4180990 | 1.6297204 | 7.919337 | 1 |
| CTGGT | 4289120 | 1.5981967 | 10.781751 | 1 |
| ATTTC | 5157555 | 1.5623323 | 5.1350837 | 5 |
| CTGAA | 3615255 | 1.5225173 | 5.695645 | 1 |
| GGCGA | 2896810 | 1.500638 | 5.1070433 | 1 |
| GATAA | 3864955 | 1.42385 | 5.253694 | 2 |
| GGAAT | 3675435 | 1.3927901 | 5.77567 | 3 |
| GGGAA | 2994625 | 1.3570467 | 6.526839 | 3 |
| GTTGG | 3886805 | 1.3031917 | 5.9386873 | 1 |
| AATCT | 3691330 | 1.2999623 | 6.087386 | 5 |
| CTGAG | 2967690 | 1.2855796 | 5.2979283 | 1 |
| GAATT | 4029540 | 1.2768999 | 5.091948 | 4 |
| GTGAA | 3306585 | 1.2530158 | 6.1844807 | 1 |
| TCTCG | 3015115 | 1.2485685 | 5.14383 | 7 |
| TGGTT | 4063680 | 1.1393564 | 5.1582294 | 2 |
| GTGGG | 2737530 | 1.0976149 | 6.2295904 | 1 |
| GGGAT | 2691400 | 1.0490887 | 5.3290915 | 3 |
| TGGGT | 2974090 | 0.9971711 | 5.136582 | 2 |
| ATCTC | 2470110 | 0.99441516 | 5.1458383 | 6 |
| CTCGC | 1774055 | 0.9763321 | 6.8884172 | 8 |