![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SDW_HiSeq_4_GCCAAT_L006_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 22865788 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 45 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
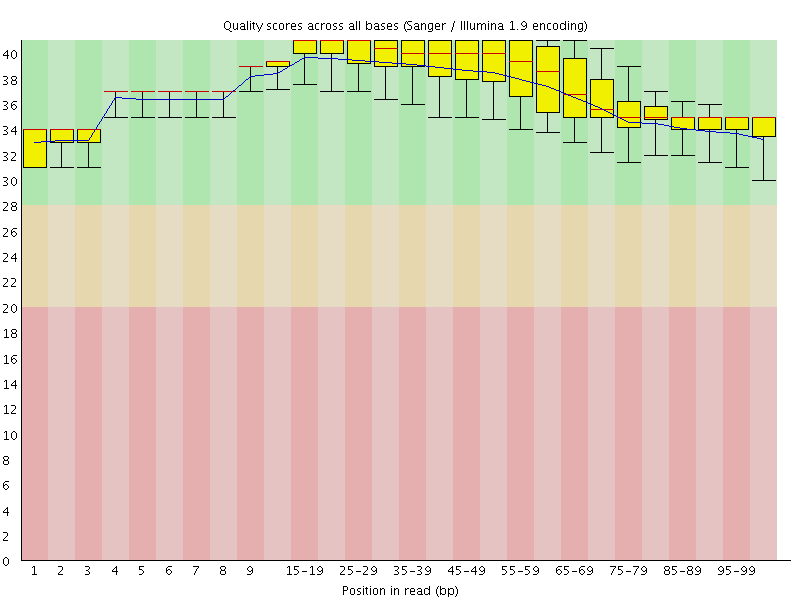
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
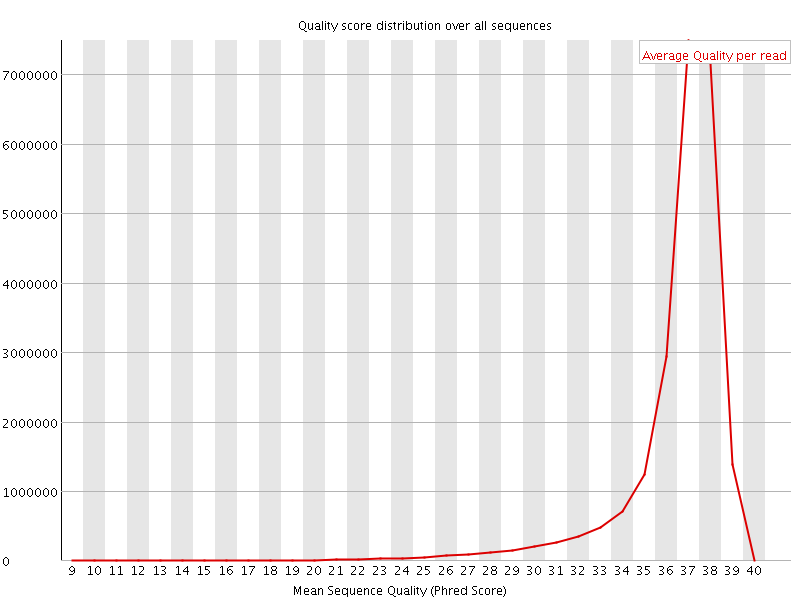
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content

![[FAIL]](Icons/error.png) Per base GC content
Per base GC content

![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
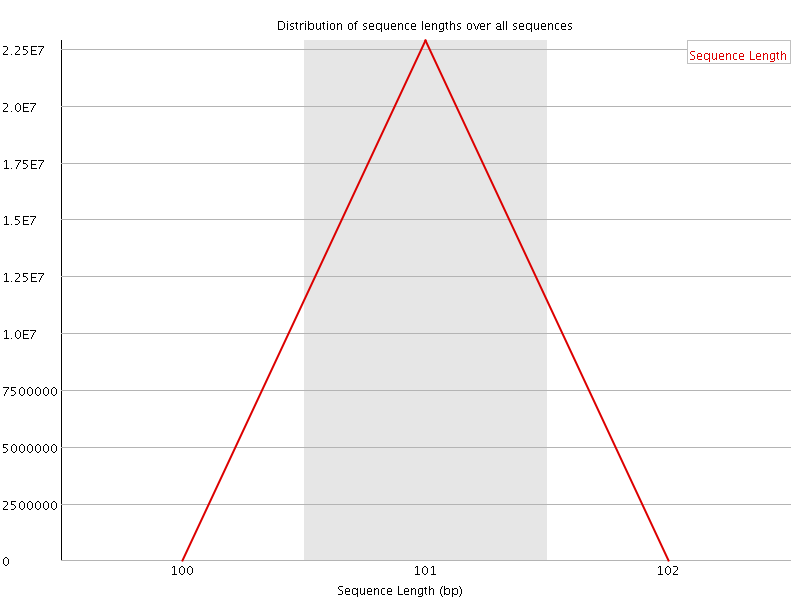
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
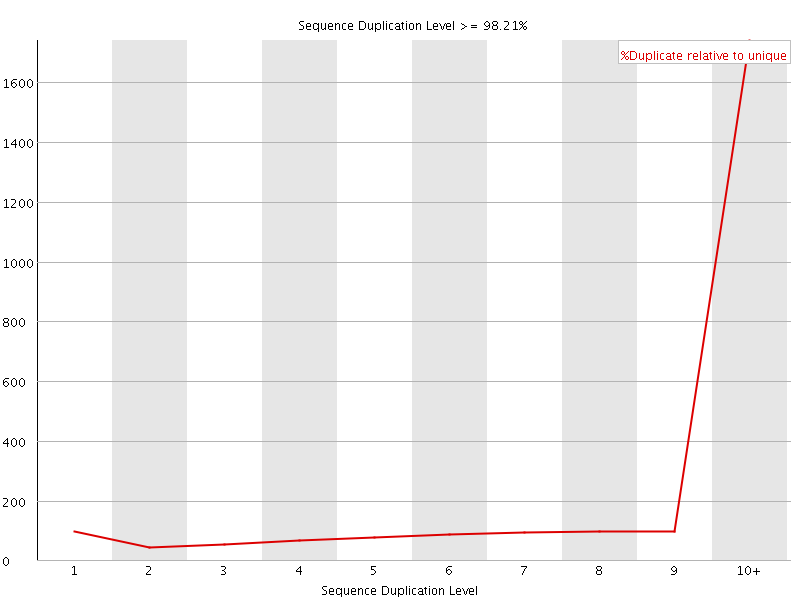
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GTAAATTCGTCGAAGAATTTAGAAGCCACCACGAAGACGAAGAACGAGGT | 44840 | 0.19610082976366264 | No Hit |
| GTTGAGTCCTTTTGGATGGAGTAGTCTTGGAGTGTGTTGCCATCTTCAAG | 38357 | 0.16774842835068707 | No Hit |
| GGAGTGTTGAGTCCTTTTGGATGGAGTAGTCTTGGAGTGTGTTGCCATCT | 37662 | 0.1647089529562681 | No Hit |
| GTCCTTTTGGATGGAGTAGTCTTGGAGTGTGTTGCCATCTTCAAGCTGCT | 36494 | 0.1596008849552878 | No Hit |
| CTTGATTTCATTACCTTCTTTCTTAACGAAGAAAAGAGCTGGAATTGAAG | 33014 | 0.14438164125373681 | No Hit |
| CGATAATCTCGCTCGTTAGCTGAATCAACGCTAGACAGGTCAACCCACGC | 32177 | 0.14072115074276031 | No Hit |
| GCCAGATCTTTTCCATATCATCCCAGTTGTTGACGATACCGTGTTCAATT | 30049 | 0.1314146706861797 | No Hit |
| CTTGTCGATACCATCGTATTCCATACCATCGAAGTATGGCTTGAAGAGCA | 23193 | 0.10143101125576778 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
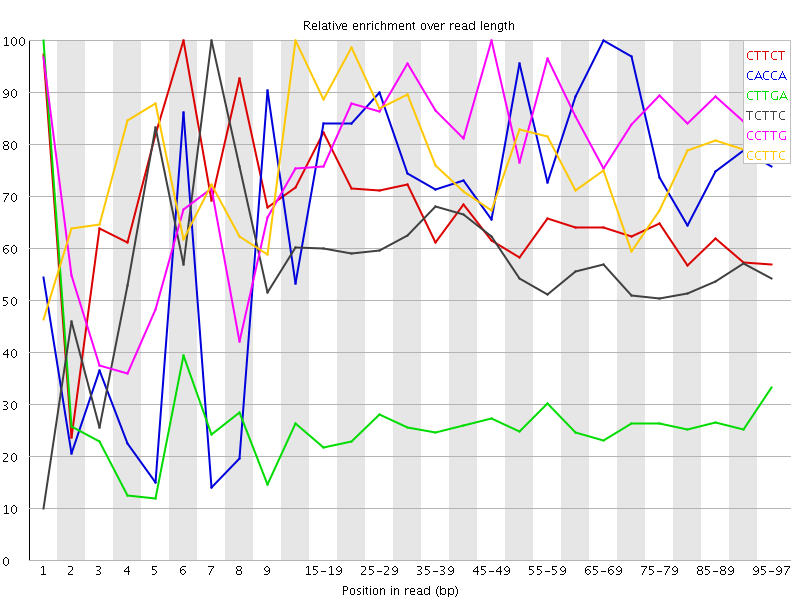
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CTTCT | 11621100 | 4.503107 | 6.8078003 | 6 |
| CACCA | 5582455 | 4.0489993 | 5.3767085 | 65-69 |
| CTTGA | 8747050 | 3.5307727 | 13.372888 | 1 |
| TCTTC | 9096410 | 3.5248046 | 6.136094 | 7 |
| CCTTG | 7353865 | 3.4561617 | 4.1420093 | 45-49 |
| CCTTC | 6380365 | 3.372173 | 4.284629 | 10-14 |
| TTCTT | 11730775 | 3.332671 | 6.3564568 | 6 |
| GAAGA | 6564060 | 3.2333515 | 7.12907 | 2 |
| GCAGC | 4796425 | 3.2028637 | 4.726852 | 8 |
| TCCTT | 8072875 | 3.128191 | 4.3866377 | 2 |
| CAACA | 4254825 | 2.6505375 | 6.5881815 | 9 |
| TCTTG | 7612935 | 2.623199 | 5.237989 | 7 |
| CAGCC | 3311935 | 2.4870732 | 6.833236 | 9 |
| TGGAG | 5671280 | 2.468975 | 6.6163263 | 2 |
| AAGAA | 5162455 | 2.4561355 | 6.0091515 | 3 |
| CTGGA | 4944070 | 2.4205058 | 15.836832 | 1 |
| TGGAA | 5711080 | 2.401428 | 8.556718 | 2 |
| CTTTG | 6896285 | 2.376262 | 6.731572 | 1 |
| TGTTG | 7658130 | 2.3464746 | 5.0476046 | 5 |
| TCAGC | 4048615 | 2.2290199 | 5.3508472 | 8 |
| CCTGG | 3888550 | 2.2165596 | 8.331518 | 1 |
| CTGTG | 5095560 | 2.129533 | 6.2329535 | 1 |
| CTGGG | 4085230 | 2.0707245 | 16.55751 | 1 |
| TCAAC | 3852330 | 2.0485504 | 5.9230423 | 8 |
| TGAAG | 4826640 | 2.0295334 | 5.0602927 | 2 |
| GTTGA | 5331280 | 1.9136112 | 8.062142 | 1 |
| TTGAT | 6371155 | 1.8855021 | 5.3185 | 2 |
| CTTGG | 4497180 | 1.8794583 | 5.0795817 | 1 |
| TGGGA | 4311535 | 1.877014 | 9.823781 | 2 |
| TTGAG | 5229165 | 1.876958 | 5.8998675 | 2 |
| CTTGT | 5404885 | 1.8623681 | 6.6020794 | 1 |
| TTGAA | 5330665 | 1.8480738 | 5.436855 | 2 |
| TGGAT | 4999260 | 1.794436 | 5.33796 | 2 |
| GATCT | 4397905 | 1.7752275 | 5.213404 | 5 |
| CTGGC | 2996110 | 1.707849 | 7.910555 | 1 |
| GATGT | 4551290 | 1.6336415 | 5.1584826 | 5 |
| GTGGA | 3653495 | 1.5905383 | 6.371544 | 1 |
| CTGGT | 3753655 | 1.5687249 | 9.696314 | 1 |
| ATTTC | 4574240 | 1.5223473 | 5.671121 | 5 |
| CTCTG | 3130685 | 1.471356 | 5.2119126 | 1 |
| GGGAA | 2542800 | 1.2968116 | 5.669757 | 3 |
| GTGAA | 3070665 | 1.2911711 | 5.5082784 | 1 |
| GATTT | 4331720 | 1.2819448 | 5.123303 | 4 |
| GTTGG | 3392890 | 1.2608873 | 5.5446935 | 1 |
| CTGAA | 2576215 | 1.218202 | 5.535273 | 1 |
| GTAAA | 2881795 | 1.1703887 | 6.074899 | 1 |
| TTTCA | 3423885 | 1.1394991 | 5.775224 | 6 |