![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SDW_HiSeq_3_ACAGTG_L006_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 41540808 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 45 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
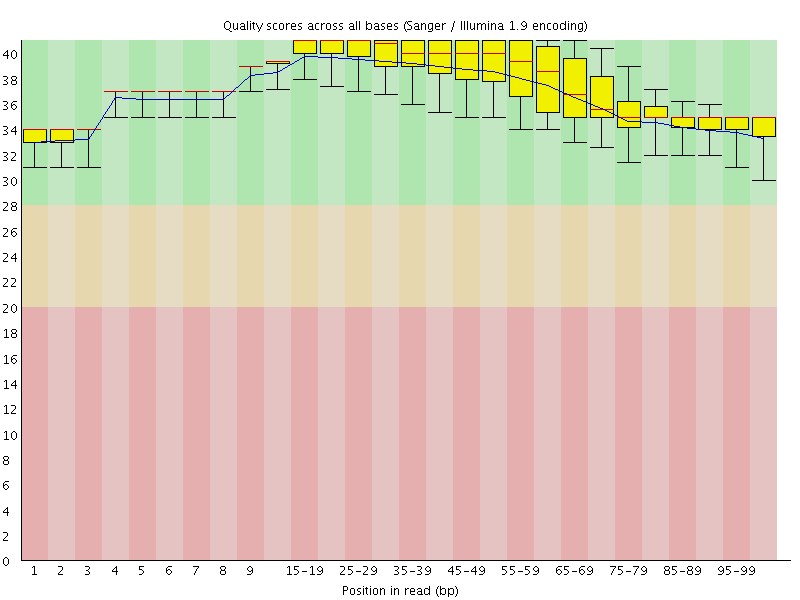
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
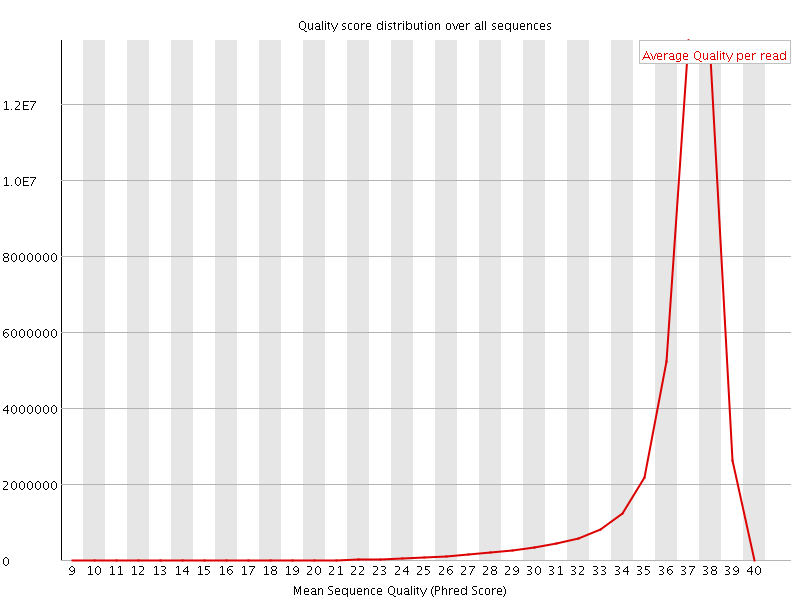
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content

![[FAIL]](Icons/error.png) Per base GC content
Per base GC content

![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
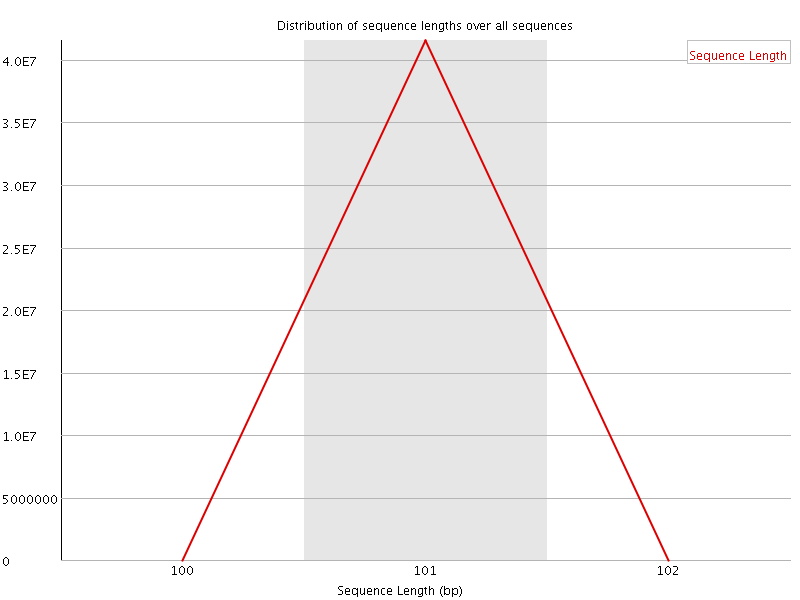
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
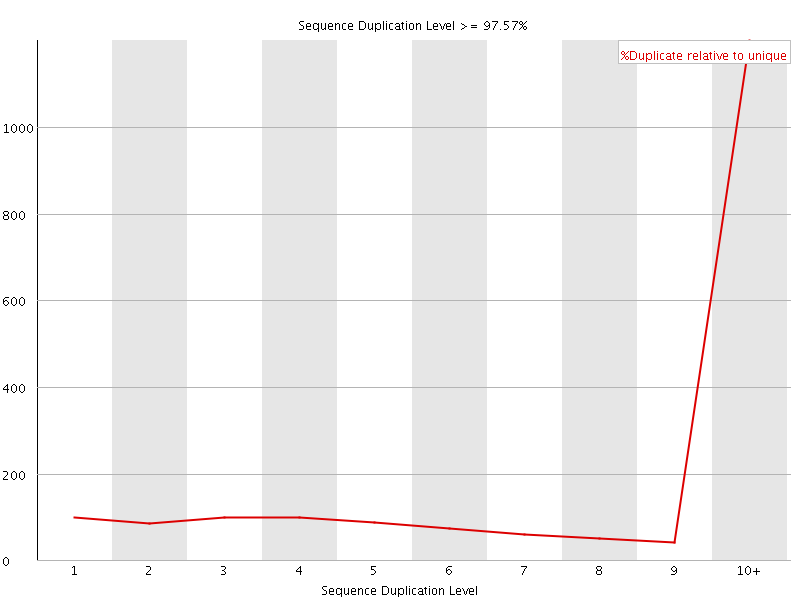
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CTTGATTTCATTACCTTCTTTCTTAACGAAGAAAAGAGCTGGAATTGAAG | 123703 | 0.2977866968788859 | No Hit |
| CTGGAATTGAAGAGACTCCGTAAGCATCTGCAGCATTGCCGTTCTTATCA | 57171 | 0.1376261145425963 | No Hit |
| GGCTGATCCTCGAGGAATTCCTTCTCATCATCCTCAGCAGCAGCGACCTT | 50952 | 0.12265529356097263 | No Hit |
| AACAATTTCACCGTTTTCCATTCTCTTGATTAATTCGAGAGCCTGAGCAA | 47290 | 0.11383986560877679 | No Hit |
| TTTGTCTTCTGGAGGACAACAGCCTGCTTGCCCTTCATGGCGAAGACCAT | 46662 | 0.11232809915493218 | No Hit |
| TAACAATTTCACCGTTTTCCATTCTCTTGATTAATTCGAGAGCCTGAGCA | 43443 | 0.10457909244326687 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
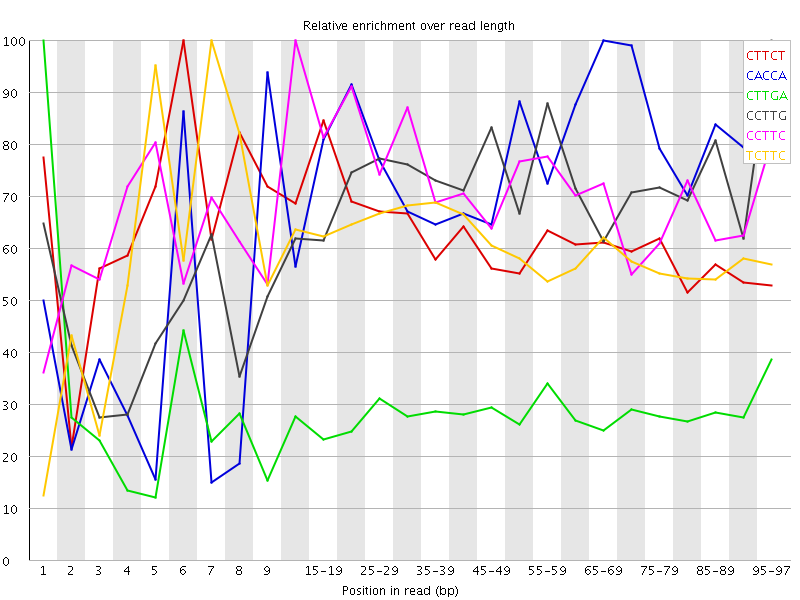
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CTTCT | 21177935 | 4.604292 | 7.3714085 | 6 |
| CACCA | 10478965 | 4.1048126 | 5.464513 | 65-69 |
| CTTGA | 16245065 | 3.63636 | 12.726742 | 1 |
| CCTTG | 13774925 | 3.5750408 | 5.087554 | 95-97 |
| CCTTC | 11917280 | 3.4795187 | 4.804915 | 10-14 |
| TCTTC | 15942060 | 3.465961 | 5.7432866 | 7 |
| TTCTT | 21336540 | 3.4541442 | 6.291849 | 6 |
| GCAGC | 9262765 | 3.3240027 | 4.7946415 | 35-39 |
| AGCCT | 10243835 | 3.0794332 | 3.859466 | 9 |
| TCCTT | 14040085 | 3.052453 | 3.885038 | 40-44 |
| GAAGA | 11310695 | 3.0193768 | 5.4228144 | 3 |
| CAACA | 7855225 | 2.6539164 | 6.5187426 | 9 |
| TCTTG | 13291770 | 2.5686905 | 5.0846443 | 7 |
| CTGGA | 9246680 | 2.470835 | 14.371094 | 1 |
| CTTTG | 12614060 | 2.43772 | 6.066263 | 1 |
| CAGCC | 6017085 | 2.4291656 | 6.9236636 | 9 |
| TGGAA | 10440810 | 2.406278 | 8.201559 | 2 |
| CCTGG | 7571875 | 2.34589 | 7.818574 | 1 |
| GATGA | 10057080 | 2.3178399 | 5.1066933 | 6 |
| CTGTG | 9621125 | 2.2195623 | 5.3122296 | 1 |
| TTCTG | 11377630 | 2.1987748 | 5.35719 | 7 |
| CTGGG | 7797735 | 2.1474469 | 14.49173 | 1 |
| TGGAG | 8918000 | 2.1182399 | 6.1714196 | 2 |
| GGATG | 8523310 | 2.0244913 | 5.212232 | 4 |
| TGGGA | 8289695 | 1.9690021 | 9.902538 | 2 |
| TCAAC | 6673055 | 1.9464211 | 5.1029615 | 8 |
| GAGAA | 7160750 | 1.9115539 | 5.113665 | 2 |
| TTGAT | 11340005 | 1.8901482 | 5.321249 | 2 |
| TTGAA | 9668570 | 1.8666433 | 5.9874506 | 7 |
| CTTGT | 9582435 | 1.8518459 | 5.0068626 | 1 |
| TTCAT | 8951035 | 1.6784413 | 5.911653 | 7 |
| GATGT | 8402025 | 1.6717799 | 5.8132095 | 5 |
| GTTGA | 8328300 | 1.6571107 | 5.3911586 | 1 |
| CTGGC | 5306425 | 1.6440169 | 6.8870964 | 1 |
| CTGGT | 6817425 | 1.5727576 | 8.369472 | 1 |
| ATTTC | 8323385 | 1.5607485 | 7.4099 | 5 |
| GGAAA | 5824415 | 1.5548208 | 5.149654 | 3 |
| GAAAG | 5684545 | 1.5174826 | 5.5446362 | 4 |
| GGAAT | 6259960 | 1.4427236 | 6.1107087 | 3 |
| GTGGA | 6055995 | 1.4384446 | 6.1087193 | 1 |
| GGGAA | 5086940 | 1.399525 | 6.156544 | 3 |
| GTGAA | 6024045 | 1.3883528 | 5.6472764 | 1 |
| GAATT | 6985335 | 1.3486098 | 5.362268 | 4 |
| GATTT | 7667655 | 1.2780421 | 6.0096235 | 4 |
| CTGAA | 4728545 | 1.2259969 | 5.0742135 | 1 |
| TTTCA | 6054545 | 1.1353097 | 7.0888915 | 6 |
| GTAAA | 4989055 | 1.1156652 | 5.1571503 | 1 |
| GTGGG | 4513865 | 1.1049746 | 5.46394 | 1 |