![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SDW_HiSeq_2_TGACCA_L006_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 26577734 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 45 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
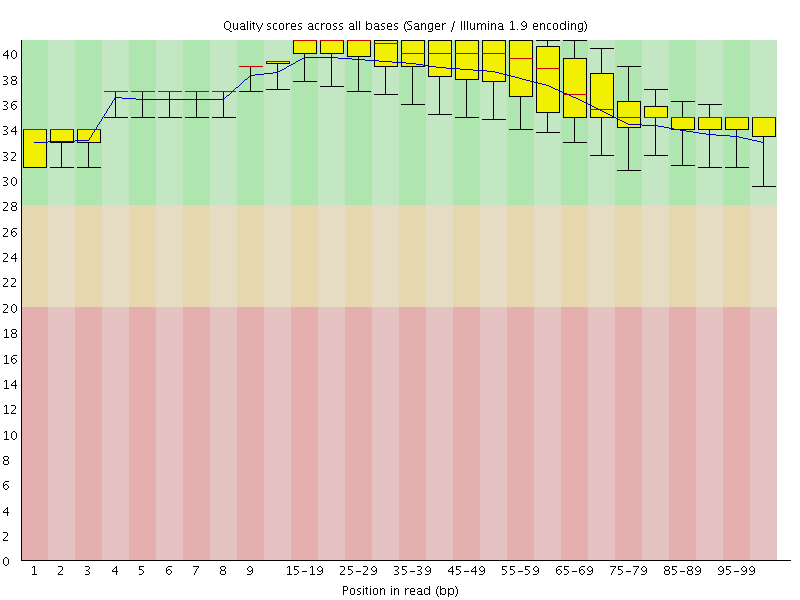
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
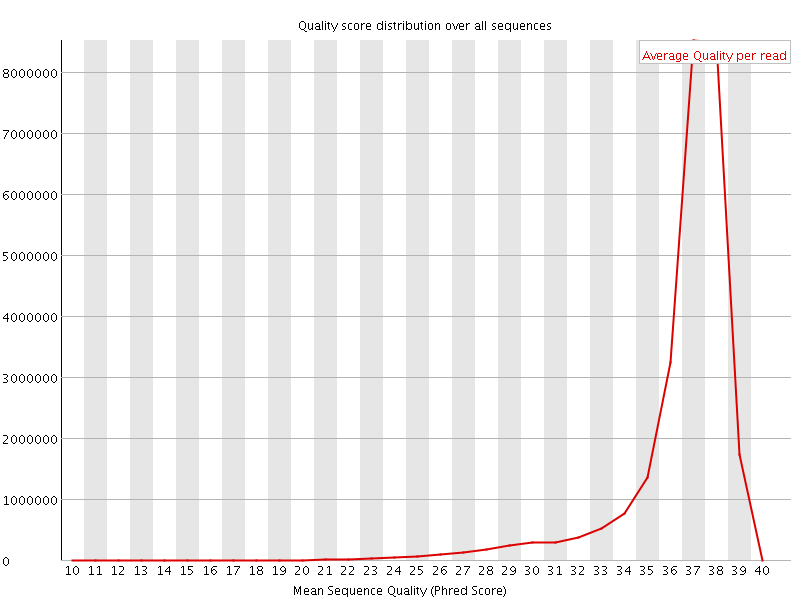
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content

![[FAIL]](Icons/error.png) Per base GC content
Per base GC content

![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
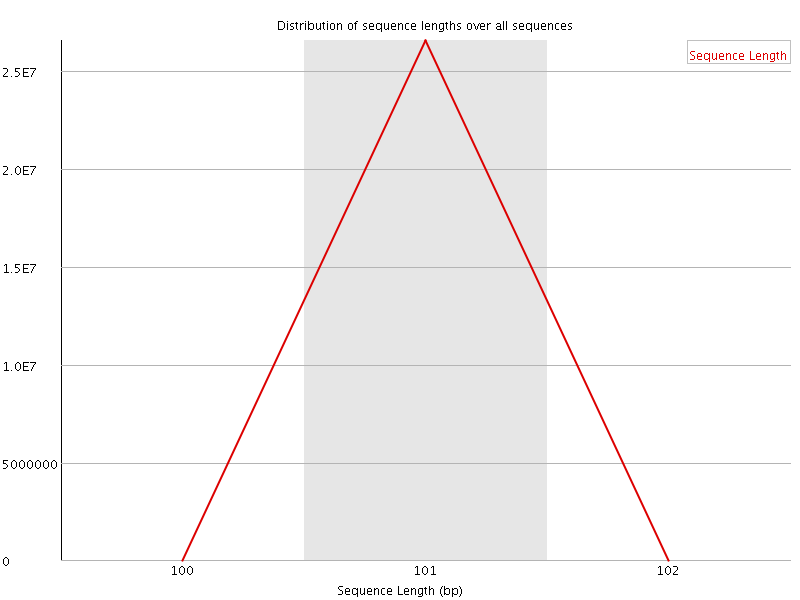
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
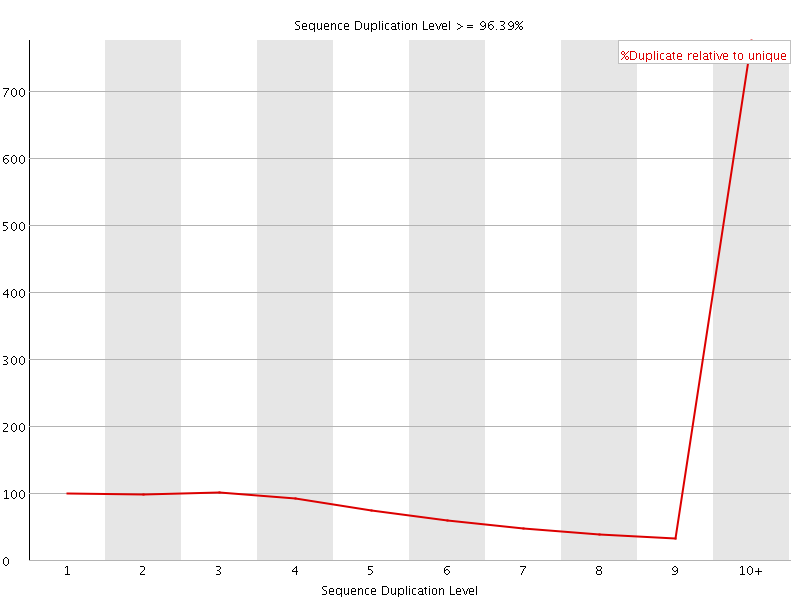
![[FAIL]](Icons/error.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACTGACCAATCTCGTATGC | 310526 | 1.1683689813435563 | TruSeq Adapter, Index 4 (100% over 50bp) |
| CGATAATCTCGCTCGTTAGCTGAATCAACGCTAGACAGGTCAACCCACGC | 45687 | 0.17189953063718677 | No Hit |
| GTTATTGCTACCCTCTTCCACCTGCTAAAATCGCAGGGAGTCCCAATTAG | 37732 | 0.141968461269121 | No Hit |
| GCCAGATCTTTTCCATATCATCCCAGTTGTTGACGATACCGTGTTCAATT | 37543 | 0.14125733969645418 | No Hit |
| GTAATCTTGAAGTGTGTTGCCGTCTTCAAGTTGCTTGCCGGCAAAGATCA | 37308 | 0.14037314091562508 | No Hit |
| CTTGATTTCATTACCTTCTTTCTTAACGAAGAAAAGAGCTGGAATTGAAG | 29538 | 0.111138142928212 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
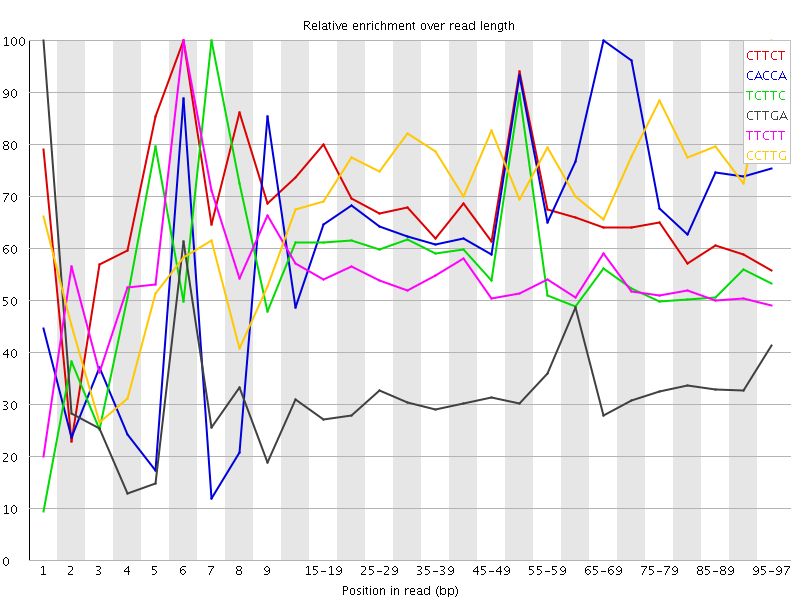
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CTTCT | 13170560 | 4.4492507 | 6.6037774 | 6 |
| CACCA | 6368980 | 3.8618546 | 5.689292 | 65-69 |
| TCTTC | 10582535 | 3.5749693 | 6.242661 | 7 |
| CTTGA | 10139055 | 3.5306704 | 10.774664 | 1 |
| TTCTT | 13417880 | 3.3127997 | 6.18093 | 6 |
| CCTTG | 7921860 | 3.2955074 | 4.4681687 | 95-97 |
| CCTTC | 6914355 | 3.1959925 | 4.50218 | 10-14 |
| GAAGA | 6980455 | 2.869832 | 16.820452 | 6 |
| TCTTG | 8524615 | 2.591782 | 5.6559753 | 7 |
| CAACA | 4834380 | 2.4537563 | 5.743269 | 9 |
| AAAAA | 6777710 | 2.4104767 | 11.028885 | 65-69 |
| CTTTG | 7702965 | 2.3419716 | 5.4504375 | 1 |
| CTGGA | 5428755 | 2.327943 | 11.210773 | 1 |
| CAGCC | 3570180 | 2.3275154 | 5.9240537 | 9 |
| AGAGC | 4738100 | 2.3270855 | 19.18173 | 8 |
| TTCTG | 7592865 | 2.3084974 | 5.072817 | 7 |
| GGAAG | 5186200 | 2.2924423 | 16.850922 | 5 |
| TGGAA | 6370915 | 2.2868547 | 6.9580865 | 2 |
| CCAGA | 3948765 | 2.154906 | 5.1241913 | 2 |
| GAGCA | 4364740 | 2.1437123 | 19.723614 | 9 |
| TGGAG | 5344125 | 2.0624802 | 5.627446 | 2 |
| CCTGG | 4002745 | 2.0505266 | 7.271102 | 1 |
| CTGGG | 4290975 | 1.9783564 | 12.034439 | 1 |
| TTGAA | 6671095 | 1.9445605 | 6.000648 | 7 |
| TCAAC | 4262955 | 1.8891456 | 5.365194 | 8 |
| GCCAG | 3126190 | 1.834251 | 5.4488115 | 1 |
| CTTGT | 6009875 | 1.8272128 | 5.316985 | 1 |
| CTGGC | 3418290 | 1.751122 | 6.0279107 | 1 |
| TGGGA | 4521000 | 1.7448082 | 7.947338 | 2 |
| AGCAC | 3186760 | 1.7390676 | 5.6671796 | 10-14 |
| GTTGA | 5421440 | 1.6990862 | 7.721516 | 1 |
| GATCT | 4856140 | 1.6910284 | 5.404532 | 5 |
| AAGAG | 4075420 | 1.6755025 | 15.6769285 | 7 |
| GATGT | 5042355 | 1.5802807 | 5.246161 | 5 |
| ATTTC | 5471195 | 1.5471395 | 5.3942885 | 5 |
| CTGGT | 4120590 | 1.5427499 | 7.3534117 | 1 |
| GTGGA | 3800315 | 1.4666712 | 6.67747 | 1 |
| GTTGG | 3861870 | 1.3012922 | 7.109043 | 1 |
| GTGAA | 3455140 | 1.2402306 | 6.014553 | 1 |
| CGGAA | 2444385 | 1.2005429 | 17.376606 | 4 |
| TTTCA | 4008605 | 1.1335497 | 5.1839204 | 6 |
| GTAAA | 3364195 | 1.1231596 | 5.034474 | 1 |
| GTGGG | 2581700 | 1.0712612 | 5.098356 | 1 |
| GATCG | 2031850 | 0.87129205 | 14.720607 | 1 |
| TCGGA | 1744740 | 0.74817437 | 15.587435 | 3 |
| ATCGG | 1466445 | 0.6288367 | 14.655577 | 2 |