![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | SDW_HiSeq_1_CGATGT_L006_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 19965718 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 45 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
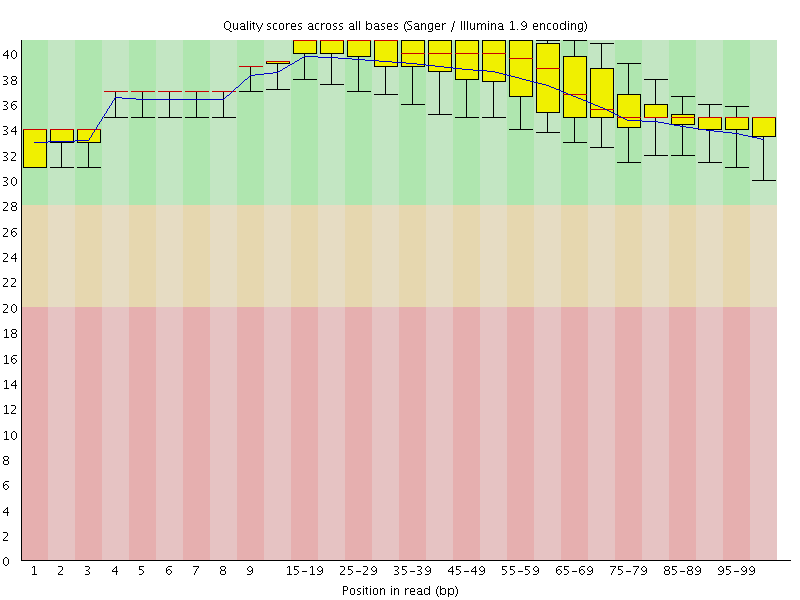
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
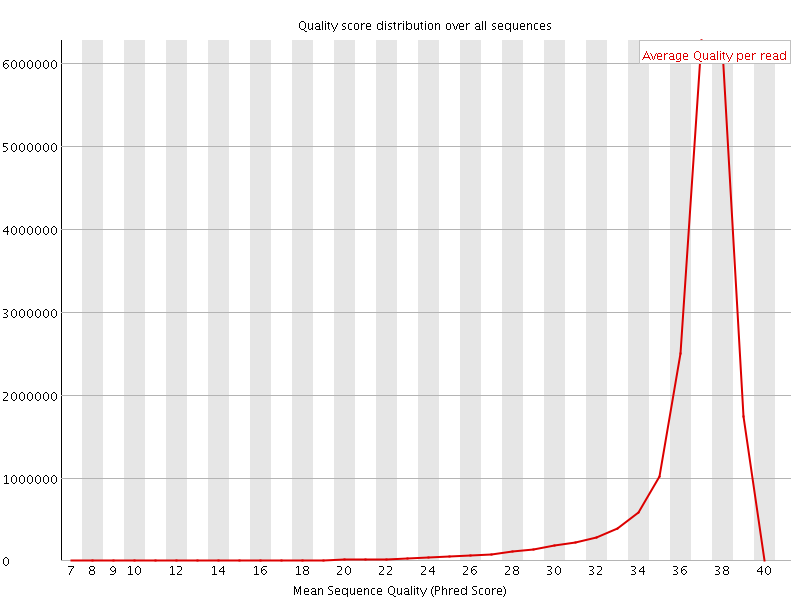
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content

![[FAIL]](Icons/error.png) Per base GC content
Per base GC content

![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content

![[OK]](Icons/tick.png) Per base N content
Per base N content

![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
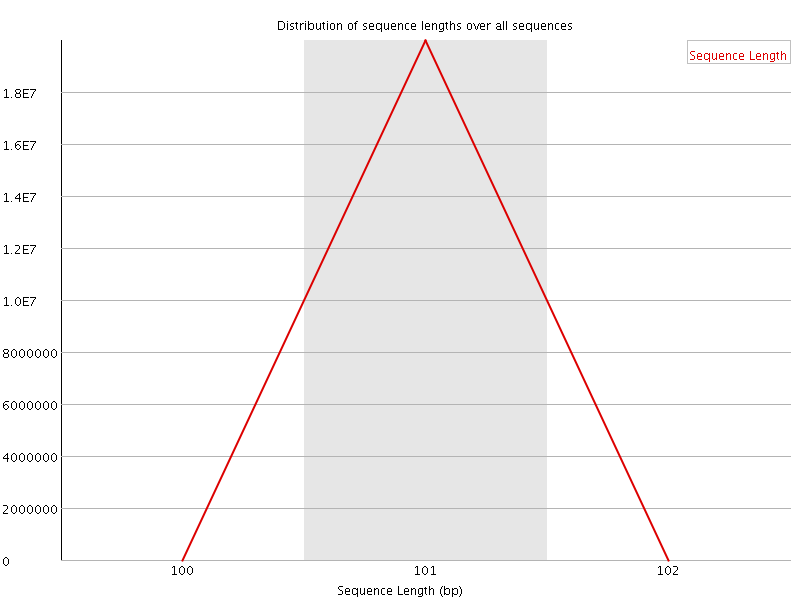
![[FAIL]](Icons/error.png) Sequence Duplication Levels
Sequence Duplication Levels
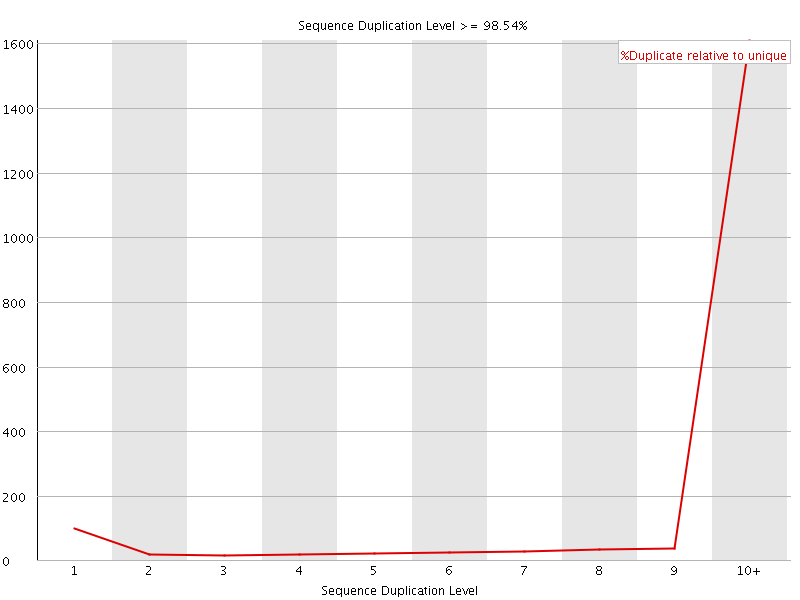
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| GTGAGTTTCTTCTTTCTTTGGCTTATCTTCTGTCTTCTTATCATCTTTCT | 127713 | 0.639661443680613 | No Hit |
| GTGCTGTATACCATCGGAGCCACTATACATTGATGATGTGAATGAGTAGT | 114093 | 0.5714445130398016 | No Hit |
| GCACAAAGTTTTTTTCTTTCCTTAGTGAGTTTCTTCTTTCTTTGGCTTAT | 110024 | 0.5510645797962287 | No Hit |
| GTTTAGTTTAAAGATACTTCTCAAGGCGGTGGAAGATATCAGCGCAGCGG | 51094 | 0.25590865302214527 | No Hit |
| CAGCACAAAGTTTTTTTCTTTCCTTAGTGAGTTTCTTCTTTCTTTGGCTT | 50869 | 0.2547817213485636 | No Hit |
| GTGAGTTTCTTCTTTCTTTGGCTTATCTTCTATCTTCTTATCATCTTTCT | 49271 | 0.24677800217352566 | No Hit |
| GTTTCTTCTTTCTTTGGCTTATCTTCTGTCTTCTTATCATCTTTCTTCTC | 47996 | 0.2403920560232294 | No Hit |
| GATCGGAAGAGCACACGTCTGAACTCCAGTCACCGATGTATCTCGTATGC | 47575 | 0.23828344164732768 | TruSeq Adapter, Index 2 (100% over 50bp) |
| ATACCATCGGAGCCACTATACATTGATGATGTGAATGAGTAGTATTGTGG | 33988 | 0.17023179431864158 | No Hit |
| GGAAGATATCAGCGCAGCGGCATGAGTACATCCACTCGTTATCGTACCAG | 30644 | 0.15348308535660976 | No Hit |
| CTGTATACCATCGGAGCCACTATACATTGATGATGTGAATGAGTAGTATT | 29269 | 0.14659628068472166 | No Hit |
| GCTGTATACCATCGGAGCCACTATACATTGATGATGTGAATGAGTAGTAT | 27531 | 0.13789135957945514 | No Hit |
| GTGGATTTTCTTCATCATCGACTTCGTCTTCGCTTTGGGCAGCTTCCTTT | 27422 | 0.13734542379092002 | No Hit |
| CCTTAGTGAGTTTCTTCTTTCTTTGGCTTATCTTCTGTCTTCTTATCATC | 24210 | 0.12125784807738946 | No Hit |
| GTTCGTAGTCAGCTGATGGTTCGAAGAGCTGAGCCTTGAGCCAGTCCTTG | 23493 | 0.11766669247757582 | No Hit |
| CACAAAGTTTTTTTCTTTCCTTAGTGAGTTTCTTCTTTCTTTGGCTTATC | 23165 | 0.11602387652675451 | No Hit |
| GTTTACAAACATATTAATATTAAAAGAAGATTAGAAATTCAGCACAAAGT | 23111 | 0.11575341292509492 | No Hit |
| GTAAACGTTTGTCTTGATGTCACCAAGATCGTGGATAGCGACAACTTGGA | 22633 | 0.11335930919188582 | No Hit |
| GTTTAGTTTAAAGGTACTTCTCAAGGCGGTGGAAGATATCAGCGCAGCGG | 22345 | 0.11191683664970124 | No Hit |
| GTACTGTCACACCCATGCTTCTCGACCGAGATCGAGTTTAGCGCAATTTG | 21859 | 0.10948266423476481 | No Hit |
| GGGAGCTTGATGGAGCCGACGAAACGCTTATCCTTAGAAACGTTGTAGCC | 21596 | 0.10816540632297822 | No Hit |
| CTAGCATACATTAGTCATTGTTTTTCTTCCTTCTTTTCTTCTTTCTTCAC | 20868 | 0.10451915628578946 | No Hit |
| CTTTGGAACTGTAAGCGTAAGAACACCATCAGCGAAAGCAGCATTGATTG | 20116 | 0.10075270020341869 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
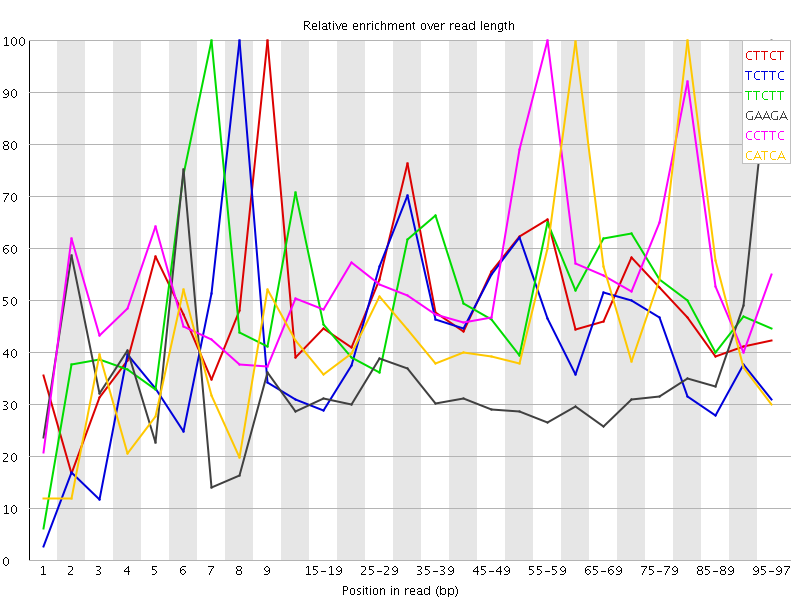
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CTTCT | 16256580 | 6.2795296 | 12.591199 | 9 |
| TCTTC | 14466075 | 5.5879 | 12.8602085 | 8 |
| TTCTT | 20133060 | 5.5777984 | 10.843492 | 7 |
| GAAGA | 5908175 | 3.8396363 | 11.085944 | 95-97 |
| CCTTC | 6642545 | 3.5774808 | 6.26725 | 55-59 |
| CATCA | 5865140 | 3.5175128 | 7.2224016 | 80-84 |
| GCAGC | 4269630 | 3.4742203 | 5.3320556 | 40-44 |
| CTTGA | 7684940 | 3.449395 | 6.8966603 | 1 |
| CACCA | 4070970 | 3.4040828 | 4.7259936 | 9 |
| TCATC | 6644540 | 3.198104 | 5.9168115 | 80-84 |
| TCCTT | 8241495 | 3.1834934 | 5.0292106 | 55-59 |
| CAGCA | 3874655 | 3.0214128 | 6.748053 | 1 |
| CCTTG | 5981335 | 3.0041094 | 3.8755448 | 85-89 |
| TGGAA | 5406770 | 2.819973 | 7.1422415 | 2 |
| CTTTG | 7771690 | 2.7995503 | 5.9653487 | 1 |
| GGAAG | 4126855 | 2.7986329 | 8.001951 | 1 |
| CAGCG | 3366015 | 2.7389438 | 5.154315 | 40-44 |
| TCTGG | 5563955 | 2.6060102 | 5.2033024 | 85-89 |
| CTGGA | 4437735 | 2.5899036 | 5.232021 | 1 |
| GCCTT | 5154965 | 2.5890677 | 7.874641 | 1 |
| TTTCT | 9204915 | 2.5501916 | 8.488426 | 6 |
| GAGAA | 3902145 | 2.5359468 | 5.750287 | 2 |
| CTTCC | 4595555 | 2.4750316 | 5.6615343 | 50-54 |
| ATCAT | 5603995 | 2.410517 | 5.4735727 | 60-64 |
| TCAGC | 3721580 | 2.329028 | 5.7425833 | 9 |
| GAGCA | 3123995 | 2.2717588 | 7.096061 | 9 |
| CAAGG | 3067100 | 2.230385 | 5.0883145 | 9 |
| AGAGC | 2965665 | 2.1566217 | 7.060442 | 8 |
| TCAAC | 3507365 | 2.1034796 | 5.0666556 | 8 |
| TGGAG | 3797040 | 2.0665326 | 5.121501 | 2 |
| GGAGA | 3025330 | 2.0516322 | 9.95889 | 1 |
| CTGGG | 3303465 | 2.011787 | 7.3343477 | 1 |
| AAGAG | 3063735 | 1.9910762 | 6.3506904 | 95-97 |
| AGAGA | 3055335 | 1.9856174 | 5.5507755 | 3 |
| AGAAA | 3092810 | 1.9261937 | 5.579161 | 3 |
| GGCGG | 2399060 | 1.8996481 | 6.497377 | 1 |
| TGGGA | 3402880 | 1.8520116 | 9.056853 | 2 |
| GTGGA | 3297820 | 1.7948328 | 12.359215 | 1 |
| GAGAG | 2645925 | 1.7943382 | 6.4960117 | 2 |
| GCTTG | 3812685 | 1.7857615 | 5.6014695 | 1 |
| GCTGG | 2882195 | 1.7552366 | 5.9679 | 1 |
| GGATG | 3168690 | 1.724554 | 5.8862715 | 1 |
| GTTCT | 4760370 | 1.7148002 | 6.1456823 | 1 |
| CAAAG | 2445035 | 1.7039112 | 9.065539 | 4 |
| GCCAC | 1938475 | 1.691424 | 5.0097623 | 15-19 |
| GTTGA | 4003780 | 1.6758978 | 8.072531 | 1 |
| TTTGG | 4927150 | 1.6551732 | 5.265521 | 2 |
| GGCGA | 2134270 | 1.6195387 | 6.0017986 | 1 |
| GAAAG | 2476320 | 1.6093241 | 6.709026 | 4 |
| GCCAG | 1956820 | 1.5922747 | 6.631831 | 1 |
| GAGTT | 3482180 | 1.4575671 | 8.894029 | 3 |
| ACAAA | 2039725 | 1.362208 | 9.245393 | 3 |
| TGAGT | 3218590 | 1.3472339 | 8.519921 | 2 |
| GGGAA | 1941225 | 1.3164446 | 7.319673 | 1 |
| GGAGT | 2373770 | 1.2919201 | 5.080787 | 1 |
| GTGAA | 2466120 | 1.2862378 | 5.7320952 | 1 |
| AAGTT | 3134125 | 1.2571979 | 7.1458187 | 6 |
| GTTTC | 3473460 | 1.2512242 | 9.382183 | 5 |
| GCTGT | 2499685 | 1.170787 | 7.102331 | 3 |
| GGAGG | 1602595 | 1.1340718 | 5.1455126 | 1 |
| TACCA | 1881360 | 1.128312 | 8.249456 | 9 |
| GTGAG | 2071495 | 1.1274077 | 14.119494 | 1 |
| CTGTA | 2498445 | 1.1214303 | 6.3687677 | 4 |
| GTGGG | 1949390 | 1.1070975 | 9.76992 | 1 |
| TGCTG | 2331520 | 1.0920227 | 7.5738926 | 2 |
| AGTTT | 3345505 | 1.0770088 | 6.9685345 | 4 |
| ATACC | 1769310 | 1.0611122 | 8.255275 | 8 |
| GTTGG | 2389795 | 1.0438259 | 6.466296 | 1 |
| GATTT | 3207540 | 1.0325942 | 5.3303394 | 4 |
| AAAGT | 2050120 | 1.024699 | 8.803334 | 5 |
| GTAGA | 1863295 | 0.9718264 | 5.1888704 | 1 |
| GTGCT | 2068330 | 0.96875143 | 10.408317 | 1 |
| GTTCA | 2125465 | 0.9540177 | 5.6404247 | 1 |
| GTCGA | 1600755 | 0.93421555 | 6.495323 | 1 |
| GTTTG | 2770755 | 0.9307773 | 6.38887 | 1 |
| GCCGT | 1394370 | 0.91057444 | 5.762475 | 1 |
| GTAAA | 1809905 | 0.90463376 | 6.0868073 | 1 |
| GTCAA | 1596250 | 0.89275664 | 5.6762958 | 1 |
| GGGAG | 1240480 | 0.8778223 | 7.4232826 | 1 |
| TAAAG | 1724545 | 0.8619688 | 5.406735 | 9 |
| CACAA | 1138360 | 0.8506809 | 9.052551 | 2 |
| TGTAT | 2276270 | 0.7327931 | 5.402312 | 5 |
| TTAAA | 1876455 | 0.7213323 | 5.277428 | 8 |
| GCACA | 895730 | 0.69848037 | 9.790016 | 1 |
| TTTAA | 2235070 | 0.6895387 | 5.109148 | 7 |
| GTTTA | 1819275 | 0.58567405 | 10.698091 | 1 |
| TATAC | 900260 | 0.38724017 | 5.424607 | 7 |
| GTATA | 901885 | 0.36177492 | 5.3582454 | 6 |