![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-99_TAAGGCGA-TATCCTCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 3475656 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
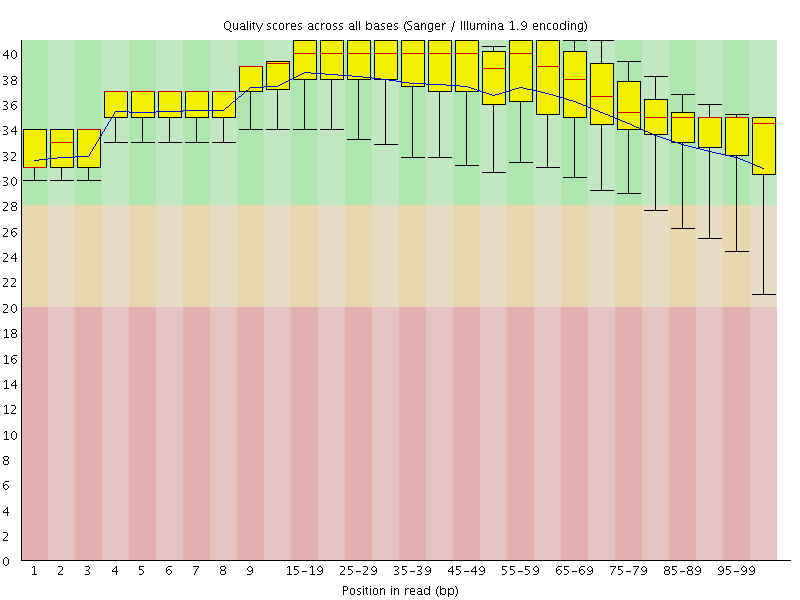
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
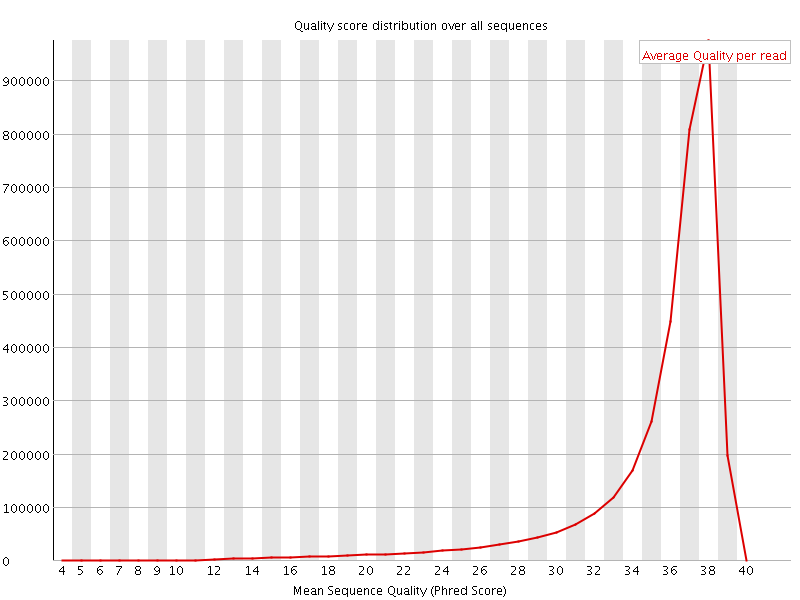
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
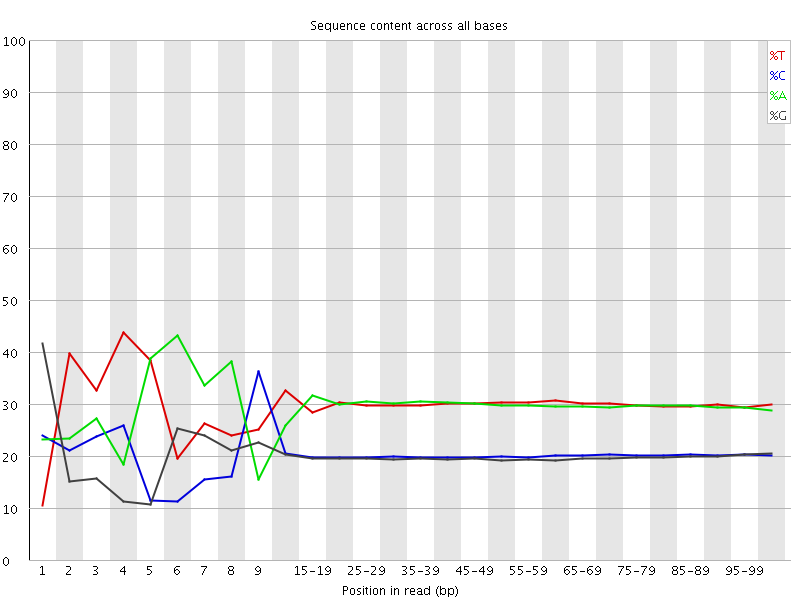
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
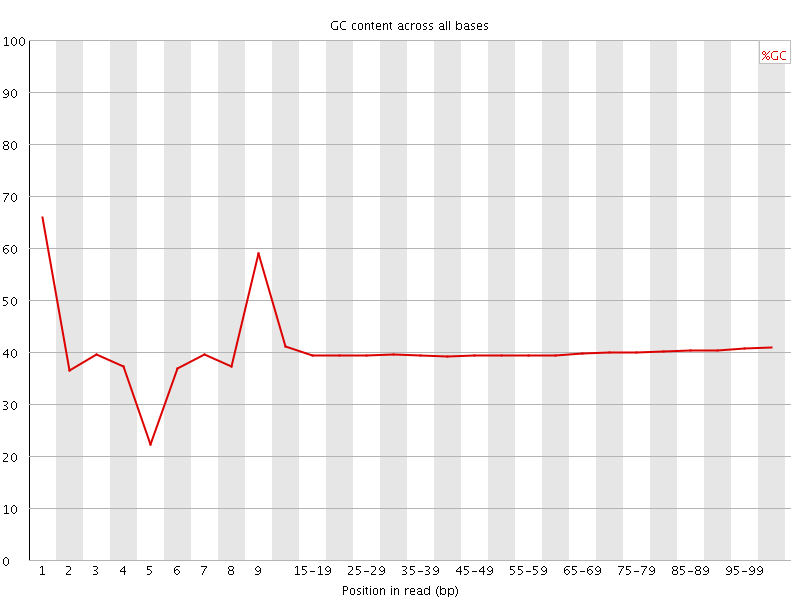
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
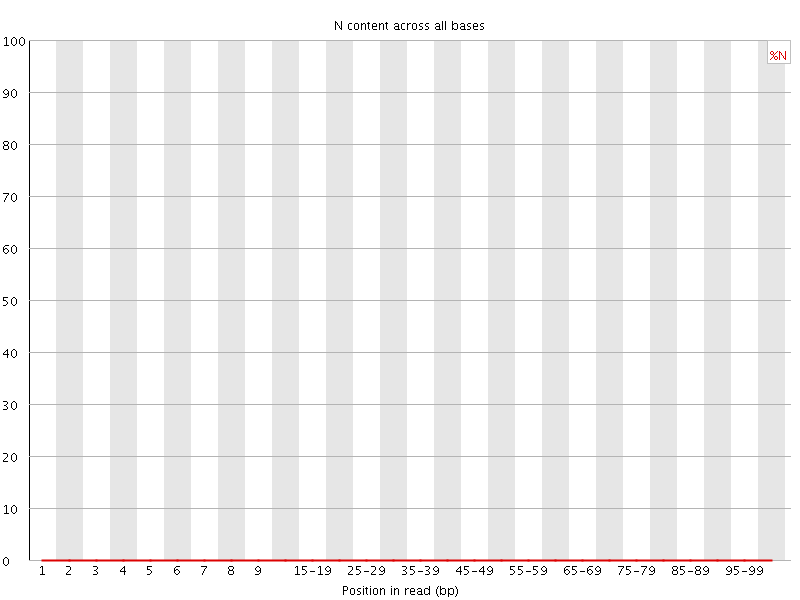
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
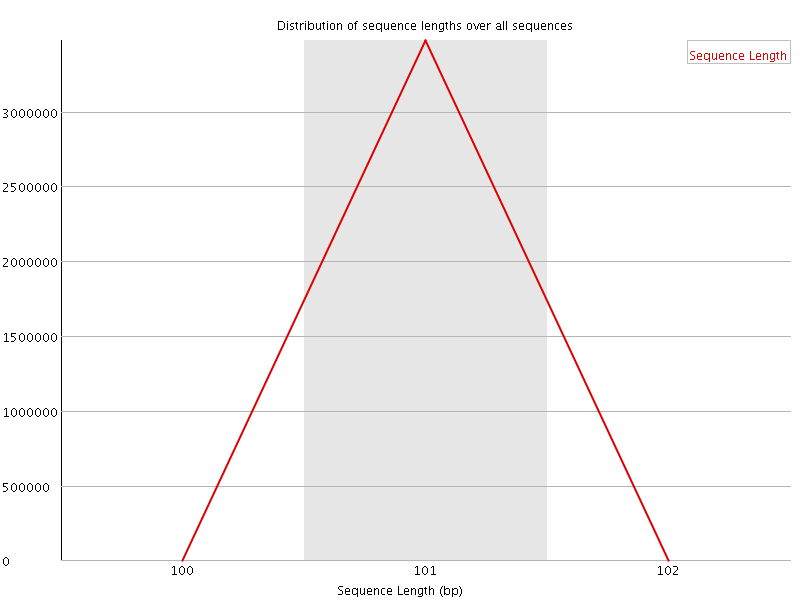
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
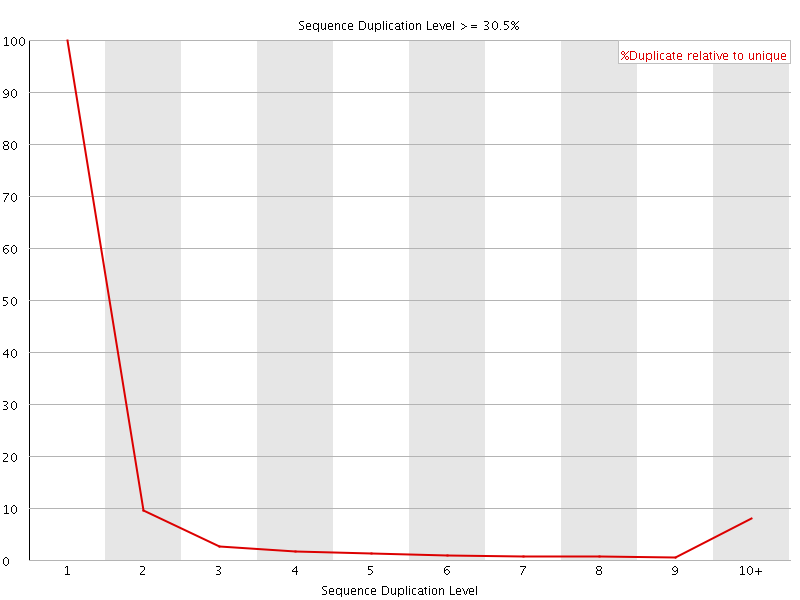
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
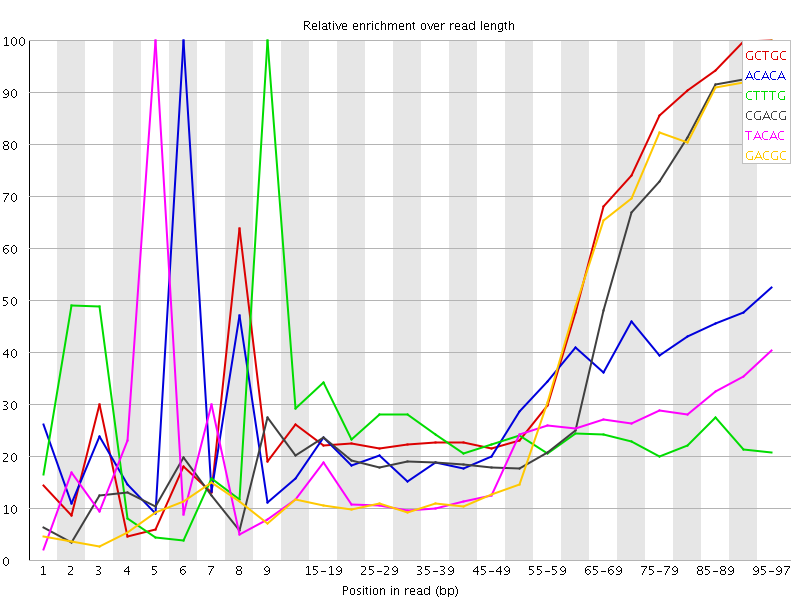
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 391390 | 2.385445 | 5.192059 | 95-97 |
| ACACA | 821525 | 2.252024 | 7.3327312 | 6 |
| CTTTG | 815950 | 2.2125309 | 8.889462 | 9 |
| CGACG | 327885 | 2.0158606 | 5.172827 | 95-97 |
| TACAC | 725880 | 1.9725949 | 9.207608 | 5 |
| GACGC | 320010 | 1.9674442 | 5.1797843 | 95-97 |
| CGCTG | 319610 | 1.9479601 | 5.112242 | 95-97 |
| CACAT | 694085 | 1.886191 | 5.0144997 | 7 |
| GCTTT | 691315 | 1.8745706 | 5.6626143 | 1 |
| GAGAG | 405895 | 1.7178359 | 5.473825 | 7 |
| TGGCT | 399385 | 1.6360255 | 5.083442 | 6 |
| TTGAG | 550930 | 1.5300748 | 5.0048947 | 9 |
| GACTC | 364180 | 1.4821168 | 5.3959103 | 7 |
| GAGTC | 352350 | 1.455968 | 7.606883 | 9 |
| CCATG | 350090 | 1.4247743 | 5.4137125 | 9 |
| AAGAC | 506415 | 1.4095159 | 5.6110783 | 5 |
| GTCTT | 508155 | 1.3779136 | 6.7583065 | 1 |
| GTGTA | 485105 | 1.3472617 | 10.148507 | 1 |
| GACTT | 490435 | 1.3414869 | 6.794296 | 7 |
| TATAC | 723730 | 1.3218685 | 5.86335 | 5 |
| TGAGT | 462830 | 1.2853982 | 5.39826 | 8 |
| AACAC | 467615 | 1.2818604 | 5.1646256 | 5 |
| GTGTT | 463405 | 1.2758445 | 6.37584 | 1 |
| TCATA | 696695 | 1.27249 | 5.970518 | 2 |
| TATGA | 684460 | 1.2693195 | 5.82915 | 4 |
| GATTG | 444880 | 1.2355465 | 5.6715446 | 7 |
| GTTCT | 446660 | 1.2111639 | 5.2890525 | 1 |
| ACACC | 299415 | 1.2106203 | 5.127378 | 6 |
| ACATG | 410065 | 1.1314541 | 5.0687876 | 8 |
| TGGAC | 272615 | 1.12649 | 7.334109 | 5 |
| GTCCA | 274750 | 1.1181602 | 8.10199 | 1 |
| ACACT | 411215 | 1.1174856 | 5.5019774 | 6 |
| GGACT | 255730 | 1.0567183 | 7.165794 | 6 |
| AGACT | 373335 | 1.0301085 | 5.4967246 | 6 |
| GAGTA | 350095 | 0.98080194 | 5.1206408 | 1 |
| GTATA | 408955 | 0.7584002 | 7.148981 | 1 |