![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-99_TAAGGCGA-TATCCTCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 3475656 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
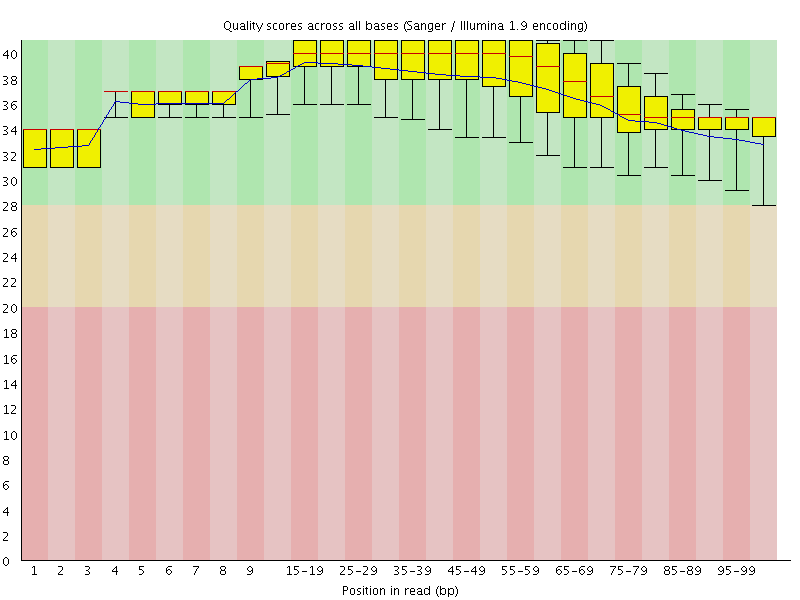
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
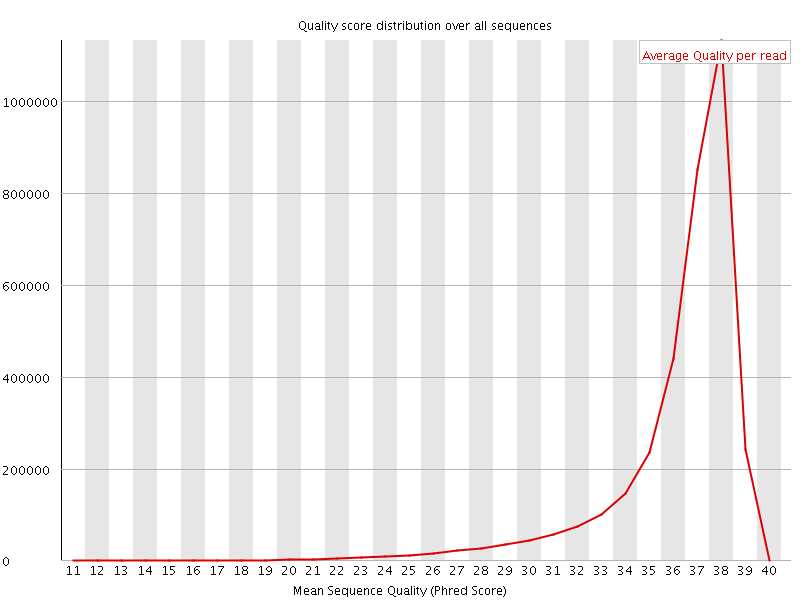
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
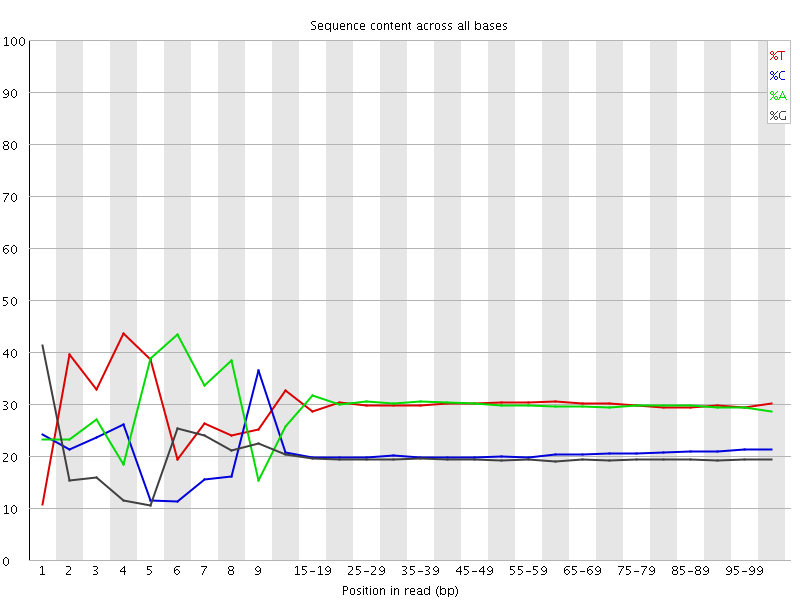
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
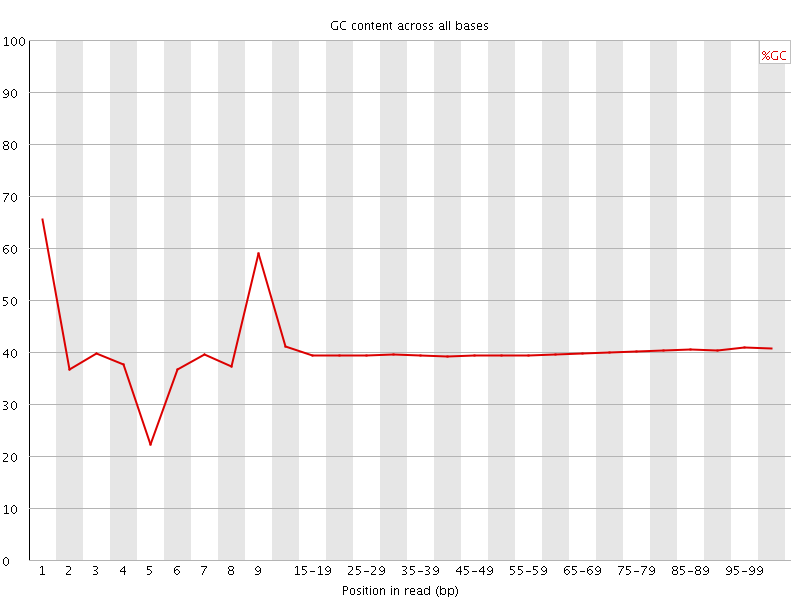
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
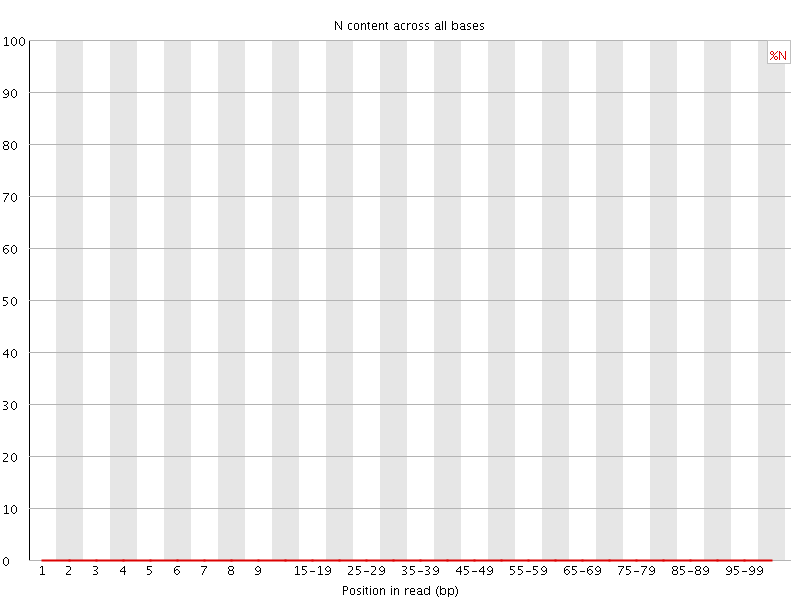
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
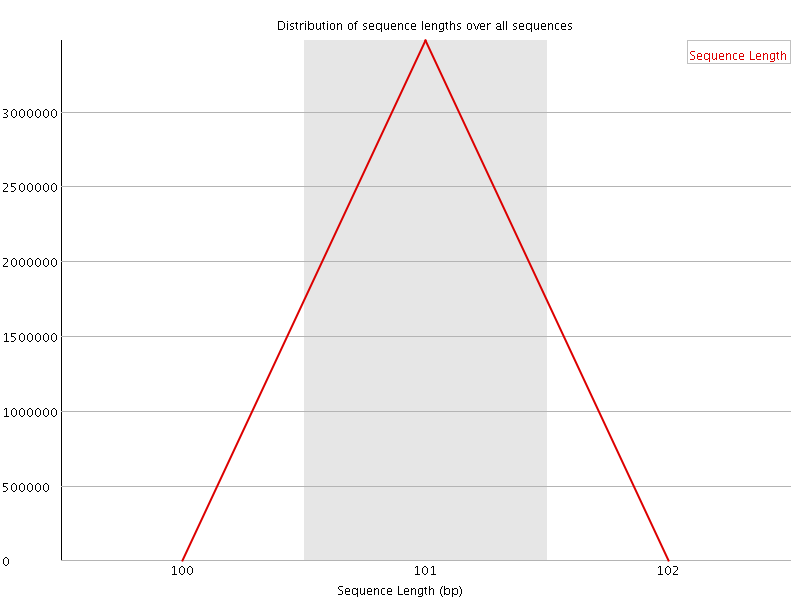
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
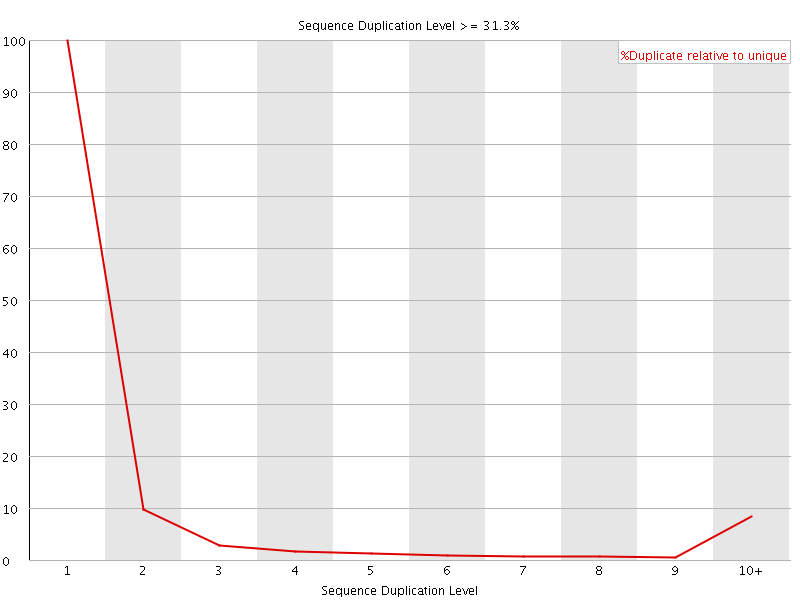
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
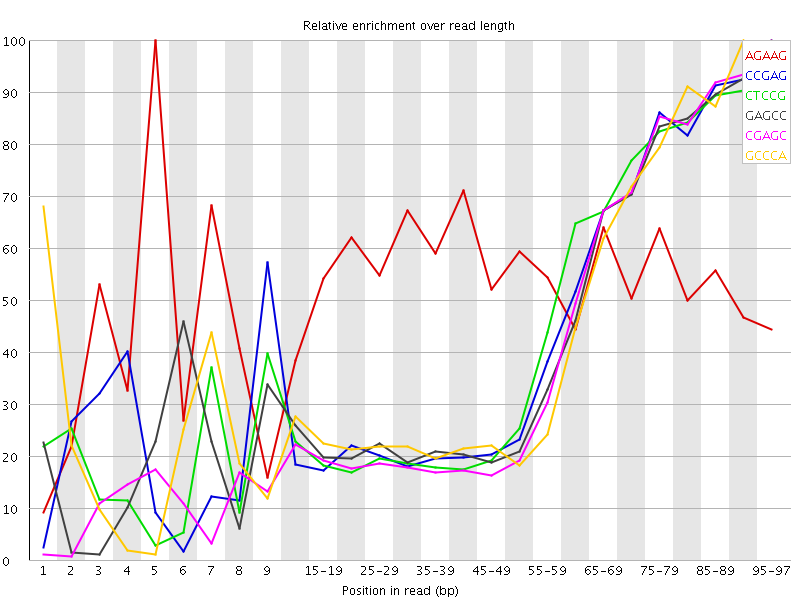
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AGAAG | 938100 | 2.722447 | 5.028881 | 5 |
| CCGAG | 413860 | 2.539744 | 5.7299876 | 95-97 |
| CTCCG | 416435 | 2.4321432 | 5.42309 | 95-97 |
| GAGCC | 395250 | 2.4255397 | 5.5316157 | 95-97 |
| CGAGC | 367565 | 2.2556443 | 5.3530803 | 95-97 |
| GCCCA | 374790 | 2.210343 | 5.0033174 | 90-94 |
| ACACA | 824035 | 2.208646 | 7.2487903 | 6 |
| CTTTG | 814230 | 2.2054815 | 8.789895 | 9 |
| TACAC | 728030 | 1.9324139 | 9.327723 | 5 |
| GCTTT | 690460 | 1.8702292 | 5.4390273 | 1 |
| GAGAG | 400135 | 1.7570934 | 5.6144214 | 7 |
| TTGAG | 550435 | 1.566596 | 5.1968217 | 9 |
| GAGTC | 355010 | 1.4836597 | 7.3635154 | 9 |
| GACTC | 368415 | 1.4796743 | 5.314213 | 7 |
| AGACT | 528985 | 1.4610298 | 5.6417484 | 6 |
| CCATG | 354130 | 1.4223009 | 5.608256 | 9 |
| GTCTT | 524670 | 1.4211588 | 6.336119 | 1 |
| AAGAC | 508430 | 1.418001 | 5.8389473 | 5 |
| GACTT | 492370 | 1.346721 | 6.439784 | 7 |
| TGAGT | 469395 | 1.3359476 | 5.467288 | 8 |
| GTGTT | 468495 | 1.320463 | 6.434538 | 1 |
| TATAC | 728395 | 1.3166642 | 5.8098197 | 5 |
| TATGA | 690085 | 1.2980026 | 5.9916286 | 4 |
| TCATA | 699025 | 1.2635742 | 5.905195 | 2 |
| GATTG | 442520 | 1.2594585 | 5.9626837 | 7 |
| AACAC | 463085 | 1.2411985 | 5.176037 | 5 |
| GTGTA | 430260 | 1.2245653 | 10.637813 | 1 |
| GTTCT | 447755 | 1.2128212 | 5.104096 | 1 |
| ACACC | 300180 | 1.1699717 | 5.232667 | 6 |
| TGGAC | 271250 | 1.1336095 | 7.1629133 | 5 |
| ACATG | 410165 | 1.1328549 | 5.025749 | 8 |
| GTATG | 393725 | 1.1205828 | 5.23546 | 3 |
| GTCCA | 277095 | 1.1129035 | 7.7415094 | 1 |
| ACACT | 412475 | 1.0948343 | 5.6432195 | 6 |
| GGACT | 256290 | 1.0710887 | 6.911654 | 6 |
| GAGTA | 352195 | 1.0121942 | 5.117325 | 1 |
| GTATA | 411760 | 0.7744923 | 7.115131 | 1 |