![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-98_TAAGGCGA-CTCTCTAT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1517524 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
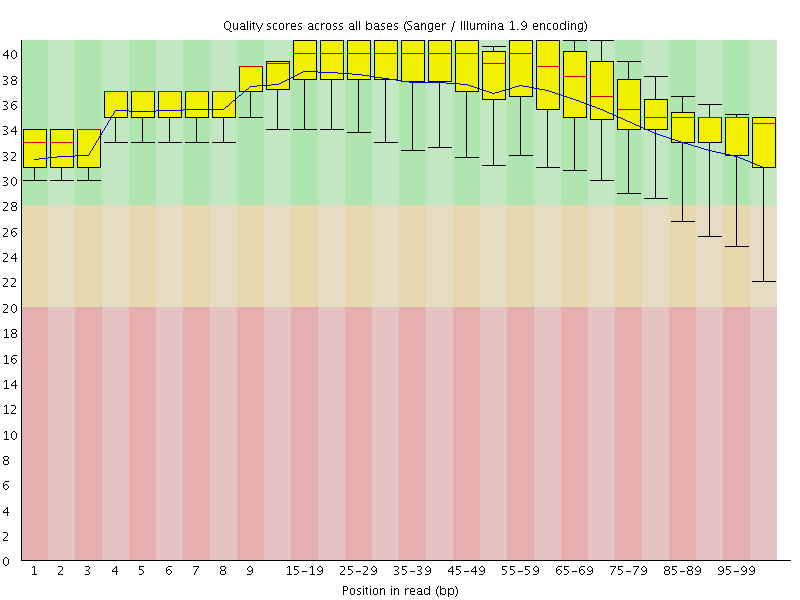
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
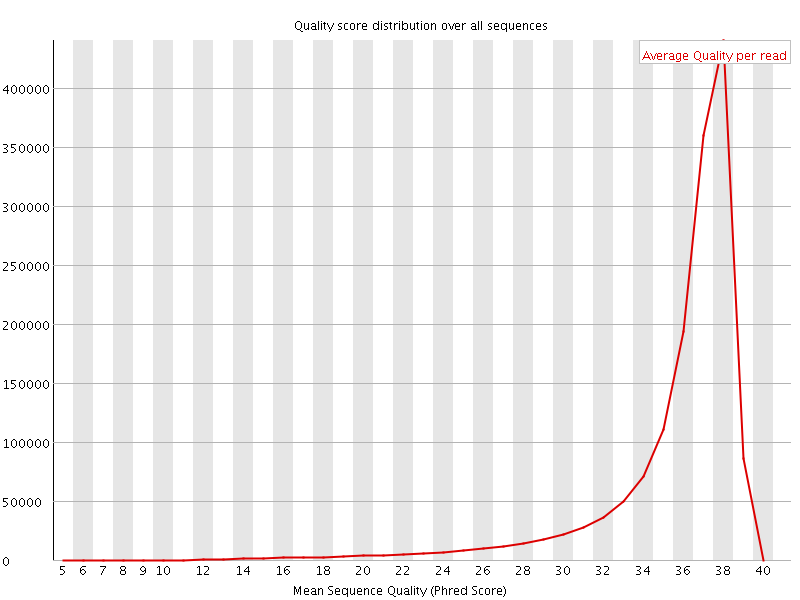
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
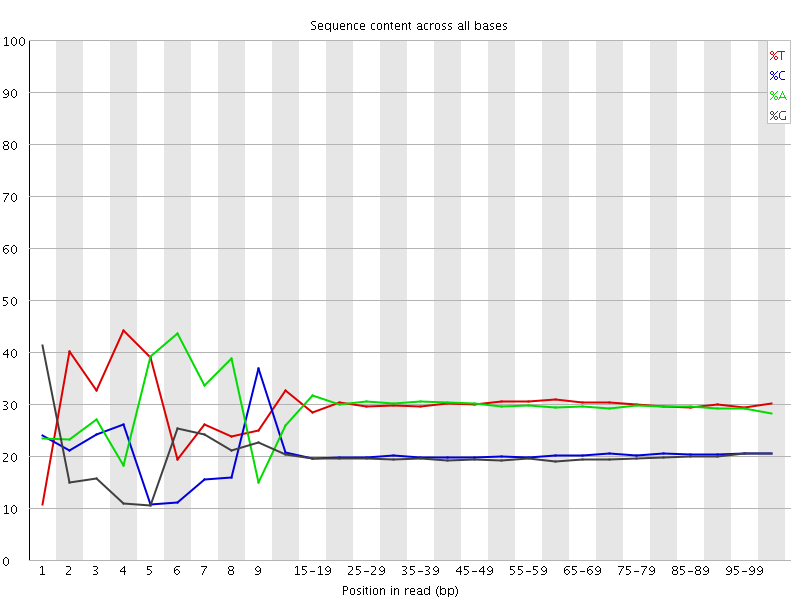
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
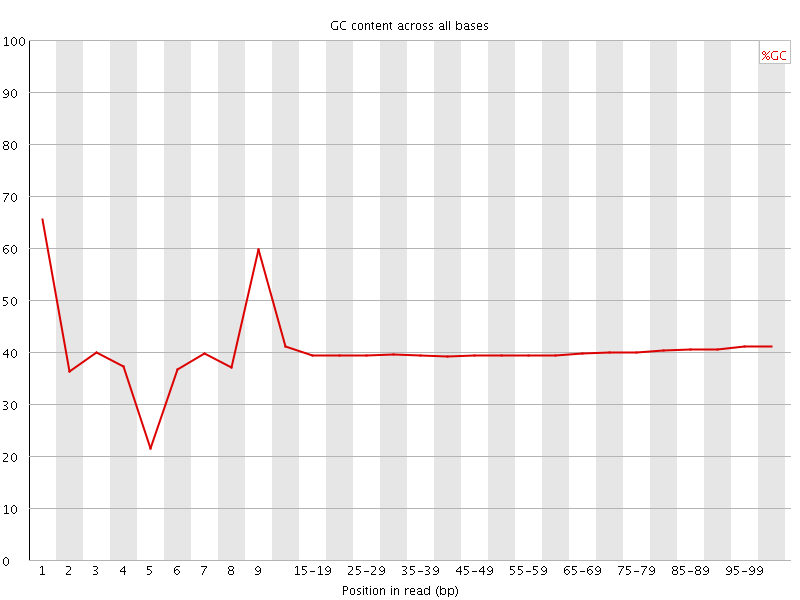
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
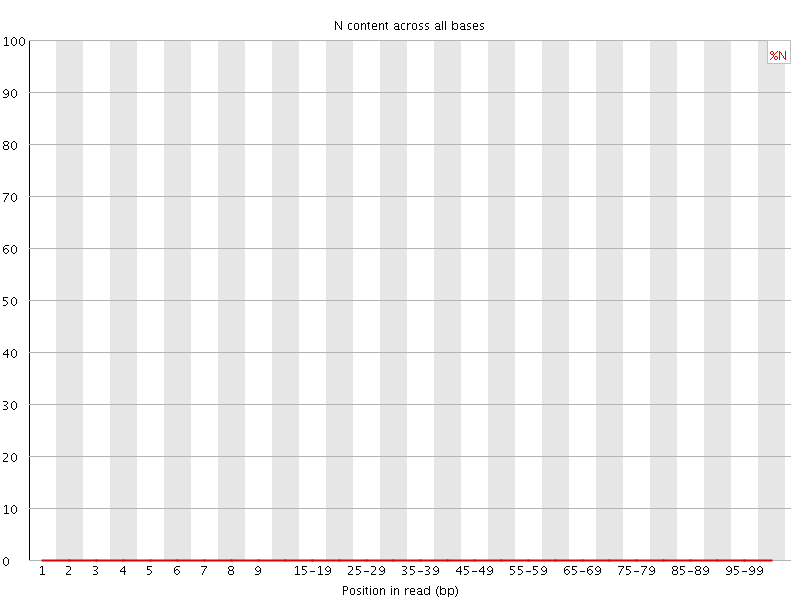
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
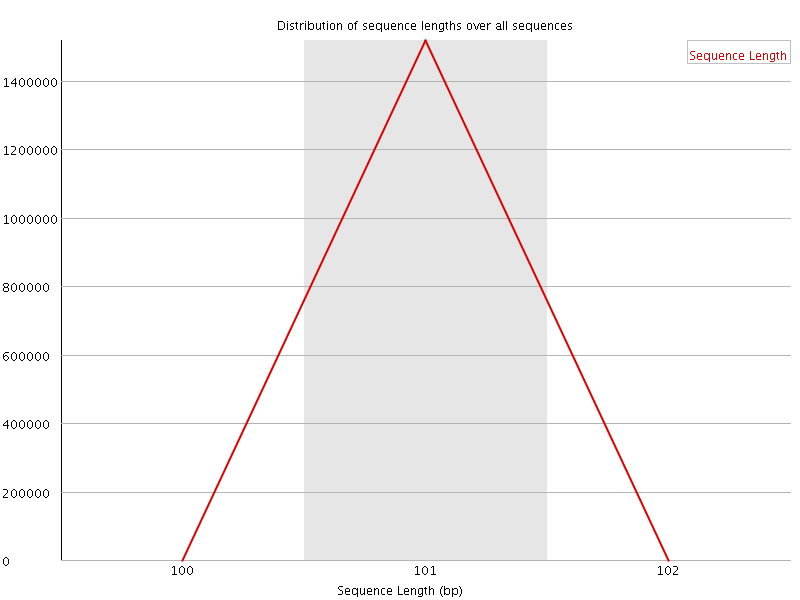
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
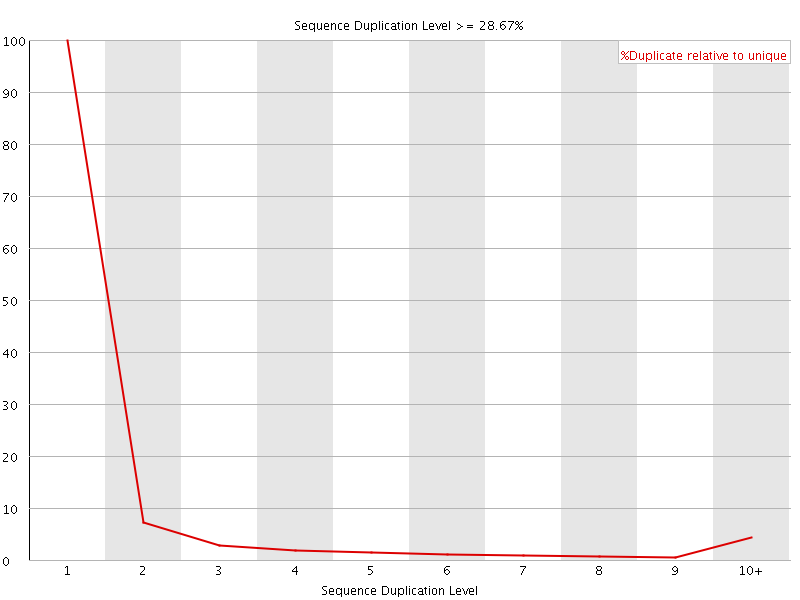
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
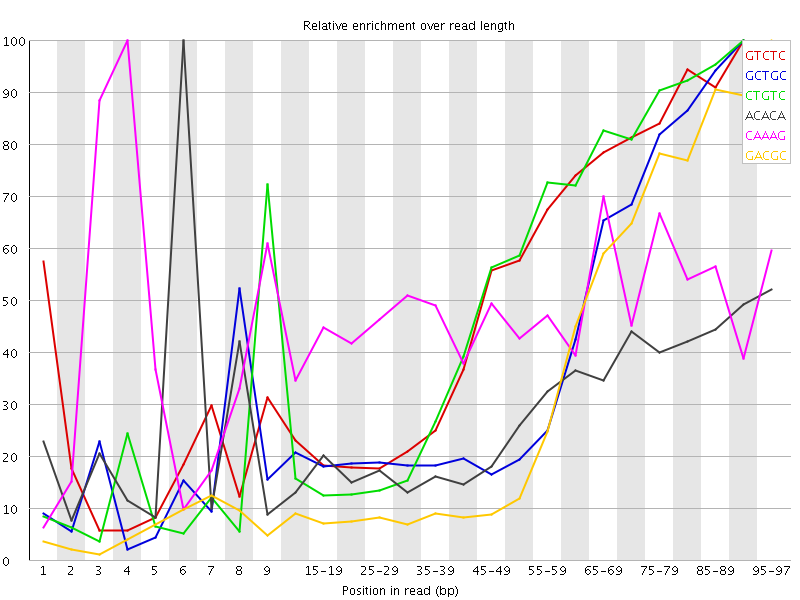
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 321825 | 2.9536328 | 5.5015755 | 90-94 |
| GCTGC | 198305 | 2.7583735 | 6.5275044 | 90-94 |
| CTGTC | 296120 | 2.7177184 | 5.1365843 | 90-94 |
| ACACA | 403380 | 2.5310855 | 8.830467 | 6 |
| CAAAG | 374840 | 2.4010112 | 5.027337 | 4 |
| GACGC | 165635 | 2.331405 | 6.669856 | 95-97 |
| CGCTG | 166365 | 2.314096 | 6.2427216 | 95-97 |
| CTGCC | 169635 | 2.3114212 | 6.1395373 | 95-97 |
| CTTTG | 368345 | 2.277008 | 9.2214365 | 9 |
| GCCGA | 160345 | 2.256945 | 6.313138 | 90-94 |
| TACAC | 363255 | 2.252464 | 11.2764635 | 5 |
| GAGAG | 228895 | 2.2416975 | 5.609098 | 7 |
| CGACG | 153650 | 2.1627092 | 6.3580956 | 95-97 |
| CACAT | 344150 | 2.1339982 | 6.2367377 | 7 |
| TGCCG | 148170 | 2.0610082 | 6.0898023 | 95-97 |
| CCGAC | 149340 | 2.0591397 | 5.9050922 | 95-97 |
| GCTTT | 308455 | 1.9067845 | 5.3676305 | 1 |
| CTGAC | 204070 | 1.8952305 | 5.0149245 | 95-97 |
| ATACA | 423610 | 1.7903244 | 5.3343806 | 4 |
| GCCAA | 187140 | 1.7587155 | 5.3428917 | 1 |
| TGGCT | 181520 | 1.7006583 | 5.370366 | 6 |
| CACCA | 176845 | 1.6280432 | 5.388685 | 7 |
| GACTC | 171200 | 1.5899614 | 5.940596 | 7 |
| TTGAG | 248570 | 1.5873055 | 6.298907 | 9 |
| GAGTC | 163265 | 1.5478605 | 7.9678235 | 9 |
| TATAC | 357370 | 1.4925796 | 6.0110197 | 5 |
| ATGGA | 227080 | 1.4673607 | 5.0716314 | 4 |
| CCATG | 157180 | 1.4597555 | 6.566633 | 9 |
| GTGTA | 228085 | 1.4564935 | 12.251184 | 1 |
| AAGAC | 227275 | 1.4557939 | 6.027832 | 5 |
| CATGA | 229165 | 1.450609 | 6.078126 | 9 |
| GTCTT | 225005 | 1.3909194 | 6.7350273 | 1 |
| TATGA | 323910 | 1.3810189 | 6.98835 | 4 |
| GACTT | 220235 | 1.3776609 | 7.3777165 | 7 |
| TCCAT | 223690 | 1.3707135 | 5.1155143 | 2 |
| TCATA | 325220 | 1.358303 | 7.34 | 2 |
| TGAGT | 210915 | 1.3468502 | 5.722957 | 8 |
| ACTCA | 215320 | 1.3351517 | 5.635371 | 8 |
| ATGAG | 201395 | 1.3013877 | 5.3367276 | 7 |
| AACAC | 203870 | 1.2792215 | 5.1864862 | 5 |
| GATTG | 199355 | 1.2730309 | 6.9027843 | 7 |
| GTGTT | 201680 | 1.2727073 | 5.9294596 | 1 |
| GTTCT | 197150 | 1.2187275 | 5.2836676 | 1 |
| ACACC | 131910 | 1.2143697 | 6.0180655 | 6 |
| ACCAT | 192265 | 1.1921929 | 5.506065 | 8 |
| TGGAC | 124430 | 1.1796789 | 8.075593 | 5 |
| ACATG | 184065 | 1.1651272 | 6.418934 | 8 |
| GTCCA | 125090 | 1.1617306 | 8.510098 | 1 |
| ACACT | 179440 | 1.1126679 | 5.4878597 | 6 |
| GGACT | 115210 | 1.0922673 | 7.7699685 | 6 |
| AGACT | 164075 | 1.0385908 | 5.8999634 | 6 |
| TGTAT | 245850 | 1.0358558 | 5.673576 | 2 |
| GTATG | 160600 | 1.0255512 | 6.545086 | 3 |
| GAGTA | 154920 | 1.0010724 | 5.225326 | 1 |
| CATAC | 156710 | 0.97172403 | 6.289329 | 3 |
| GTATA | 182250 | 0.77703893 | 7.338015 | 1 |