![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-98_TAAGGCGA-CTCTCTAT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1517524 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
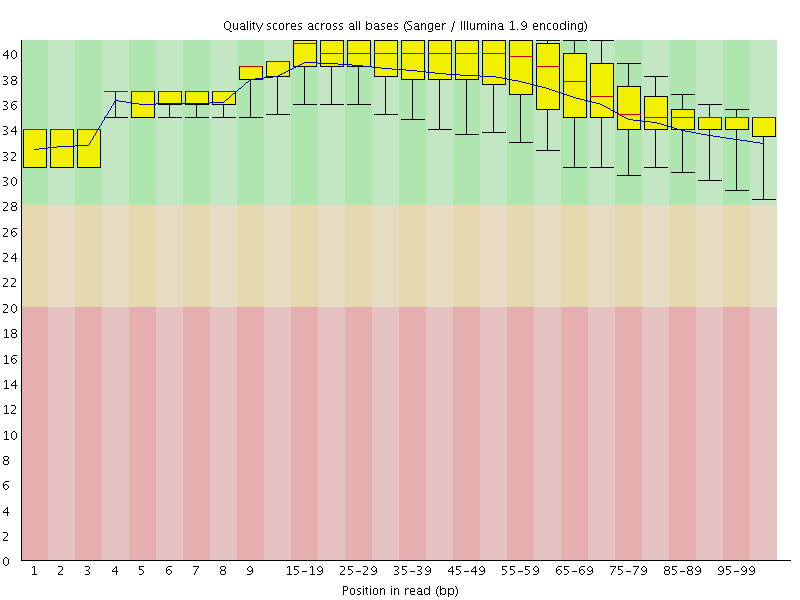
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
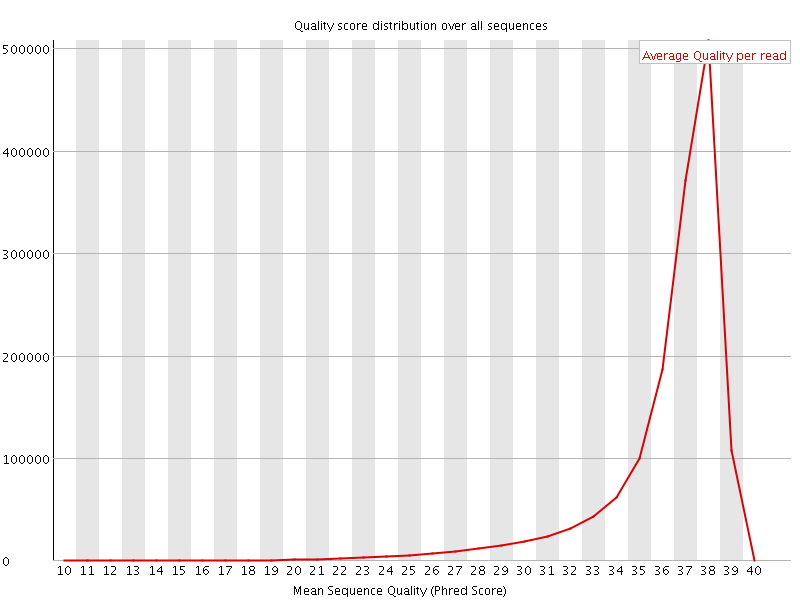
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
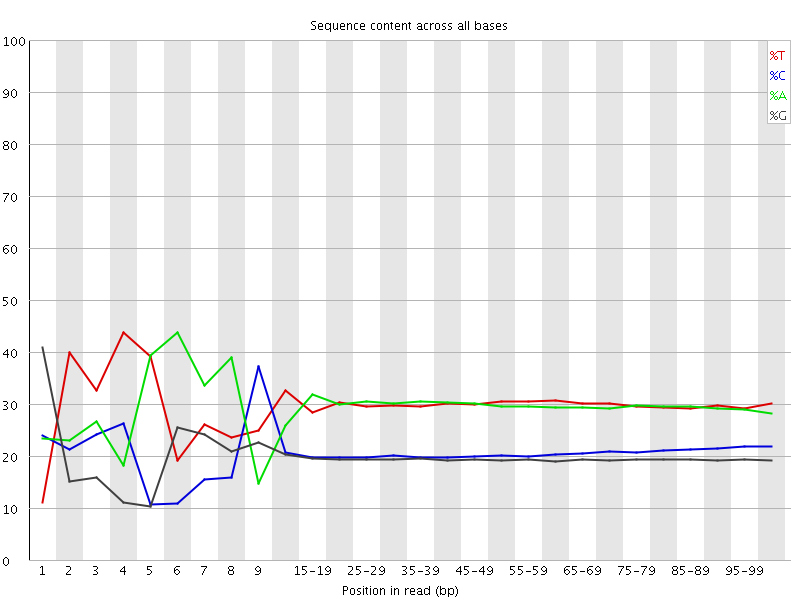
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
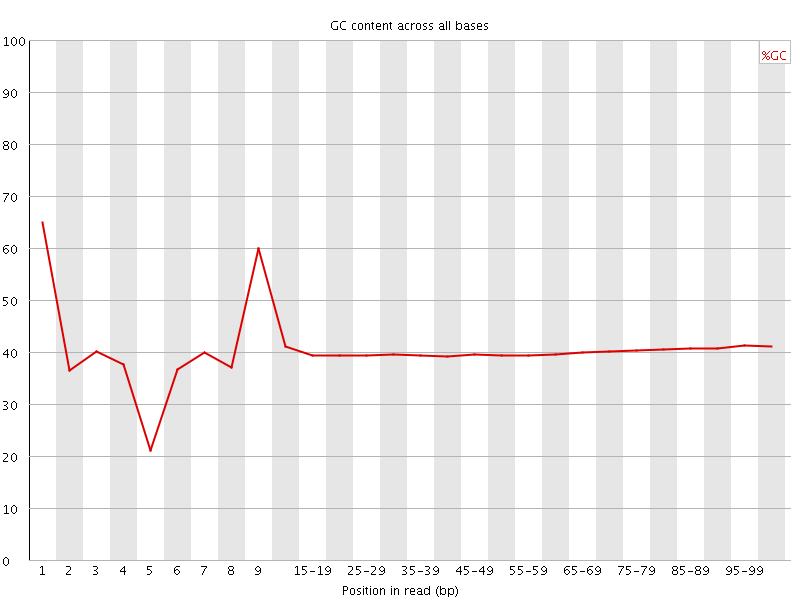
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
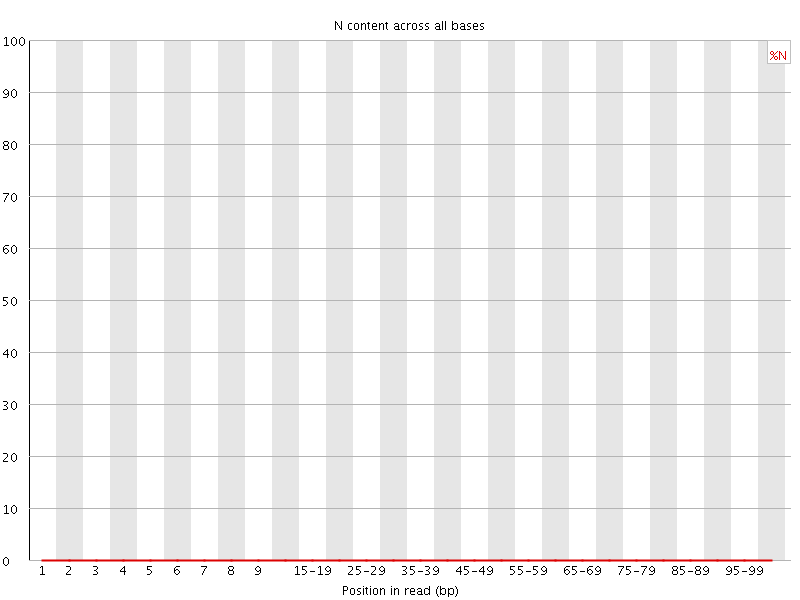
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
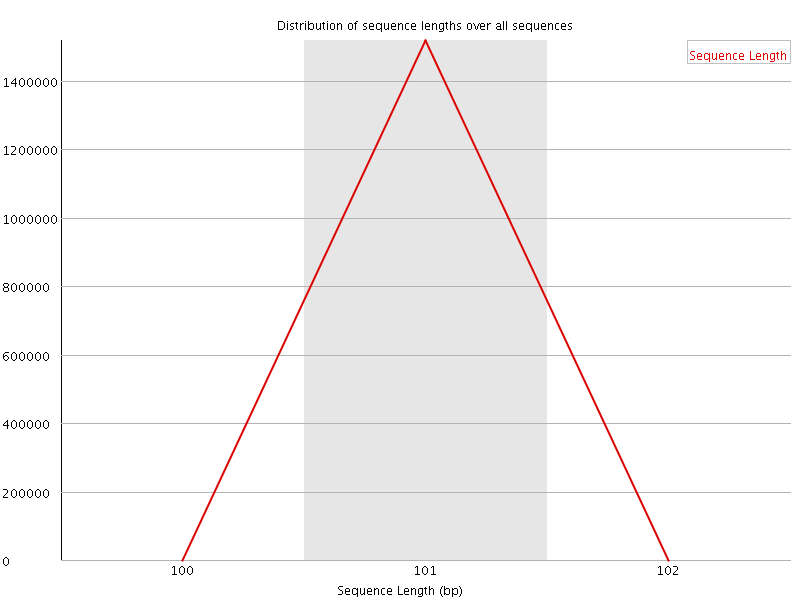
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
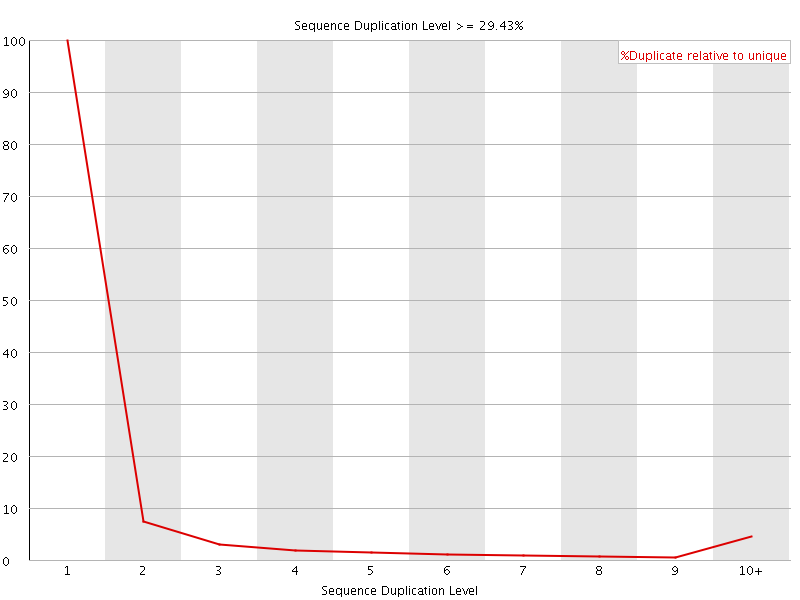
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
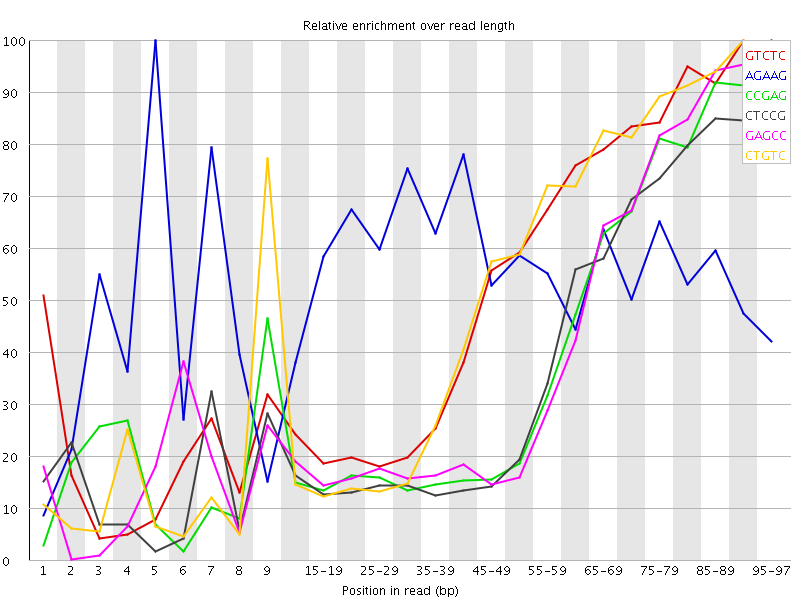
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 322560 | 2.9143403 | 5.3797417 | 90-94 |
| AGAAG | 426900 | 2.8691797 | 5.0866723 | 5 |
| CCGAG | 202945 | 2.8390994 | 7.03855 | 95-97 |
| CTCCG | 208590 | 2.7488234 | 7.037125 | 95-97 |
| GAGCC | 195410 | 2.7336884 | 6.665482 | 95-97 |
| CTGTC | 298190 | 2.6941564 | 5.097612 | 90-94 |
| CGAGC | 184560 | 2.5819023 | 6.55017 | 95-97 |
| TCTCC | 295100 | 2.5391505 | 5.255306 | 95-97 |
| ACACA | 404360 | 2.4647837 | 9.259132 | 6 |
| GCCCA | 183575 | 2.4457119 | 6.1511064 | 90-94 |
| CAAAG | 378455 | 2.4223413 | 5.03041 | 4 |
| AGCCC | 181285 | 2.4152029 | 6.010281 | 90-94 |
| CTTTG | 367995 | 2.2795436 | 9.472551 | 9 |
| ATCTC | 374580 | 2.2339761 | 5.2648683 | 95-97 |
| TACAC | 365655 | 2.2046714 | 11.716539 | 5 |
| CCCAC | 173330 | 2.1991506 | 5.8062344 | 90-94 |
| CACAT | 344015 | 2.0741954 | 6.653403 | 7 |
| CCACG | 154310 | 2.0558236 | 5.8048563 | 90-94 |
| GCTTT | 309105 | 1.9147499 | 5.578076 | 1 |
| ATACA | 429310 | 1.7941468 | 5.7625647 | 4 |
| AGGCG | 119165 | 1.750497 | 5.643438 | 95-97 |
| GCCAA | 187900 | 1.7351348 | 5.028665 | 1 |
| TGGCT | 182630 | 1.7326558 | 5.763556 | 6 |
| GGCGA | 117900 | 1.7319149 | 5.7170377 | 95-97 |
| TTGGC | 180525 | 1.7126851 | 5.1103835 | 5 |
| GAGAG | 163880 | 1.6686041 | 5.6573267 | 7 |
| TTGAG | 247885 | 1.6300657 | 6.4817386 | 9 |
| GACTC | 173845 | 1.5879259 | 5.726191 | 7 |
| AGACT | 249855 | 1.5818701 | 5.7094593 | 6 |
| GAGTC | 164805 | 1.580698 | 8.147266 | 9 |
| CACCA | 179570 | 1.5791732 | 5.721999 | 7 |
| ATGGA | 224415 | 1.4919186 | 5.195861 | 4 |
| TATAC | 360640 | 1.4908108 | 5.974614 | 5 |
| AAGAC | 228830 | 1.4646508 | 5.8155394 | 5 |
| GTCTT | 236150 | 1.4628305 | 6.7195253 | 1 |
| CATGA | 229960 | 1.455912 | 6.1852484 | 9 |
| CCATG | 157400 | 1.437715 | 6.6119137 | 9 |
| TATGA | 322655 | 1.4005463 | 7.2017555 | 4 |
| TGAGT | 212630 | 1.3982327 | 5.923793 | 8 |
| GACTT | 218620 | 1.3690971 | 7.308357 | 7 |
| TCATA | 328395 | 1.3575165 | 7.7455497 | 2 |
| ATGAG | 202000 | 1.3429029 | 5.2732186 | 7 |
| GTGTT | 203175 | 1.3215598 | 6.3997808 | 1 |
| GATTG | 198545 | 1.305611 | 7.192721 | 7 |
| ACTCA | 216190 | 1.3034906 | 5.1186767 | 8 |
| GTGTA | 195735 | 1.287133 | 12.544571 | 1 |
| AACAC | 202485 | 1.234251 | 5.233937 | 5 |
| GTATG | 187195 | 1.2309748 | 6.791001 | 3 |
| GTTCT | 198255 | 1.22809 | 5.403856 | 1 |
| TGGAC | 123355 | 1.1831377 | 8.658796 | 5 |
| ACATG | 185340 | 1.1734158 | 7.0416665 | 8 |
| ACACC | 132945 | 1.1691439 | 6.0929475 | 6 |
| GTCCA | 127835 | 1.1676639 | 8.030312 | 1 |
| ACCAT | 192835 | 1.1626745 | 5.668255 | 8 |
| ATTGA | 261730 | 1.1360897 | 5.0088186 | 8 |
| GGACT | 115185 | 1.1047766 | 8.054261 | 6 |
| ACACT | 181060 | 1.0916786 | 5.639022 | 6 |
| TGTAT | 243030 | 1.0434716 | 5.7588987 | 2 |
| GAGTA | 156380 | 1.0396196 | 5.6254234 | 1 |
| TGTAG | 150455 | 0.98937625 | 5.1434646 | 2 |
| CATAC | 160065 | 0.96509206 | 6.7206397 | 3 |
| GTATA | 183085 | 0.7947159 | 7.3081155 | 1 |