![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-184_AAGAGGCA-CTAAGCCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2577568 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
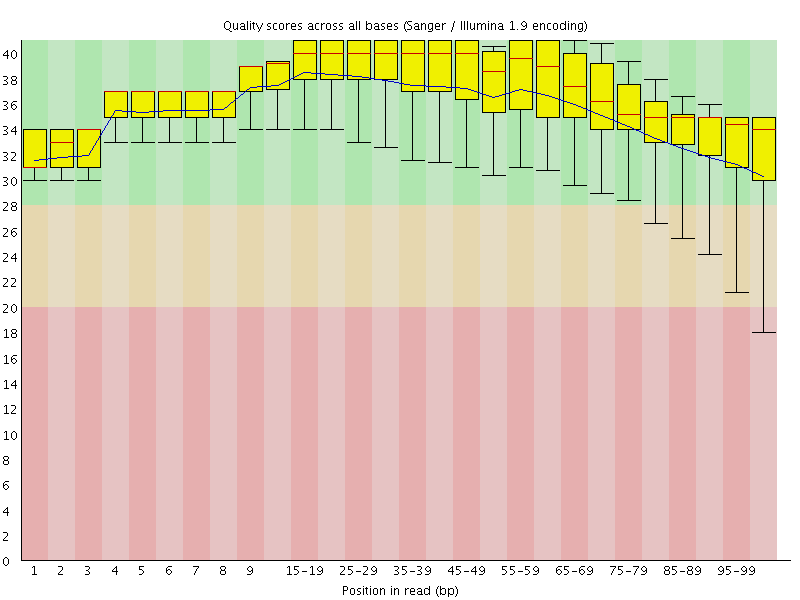
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
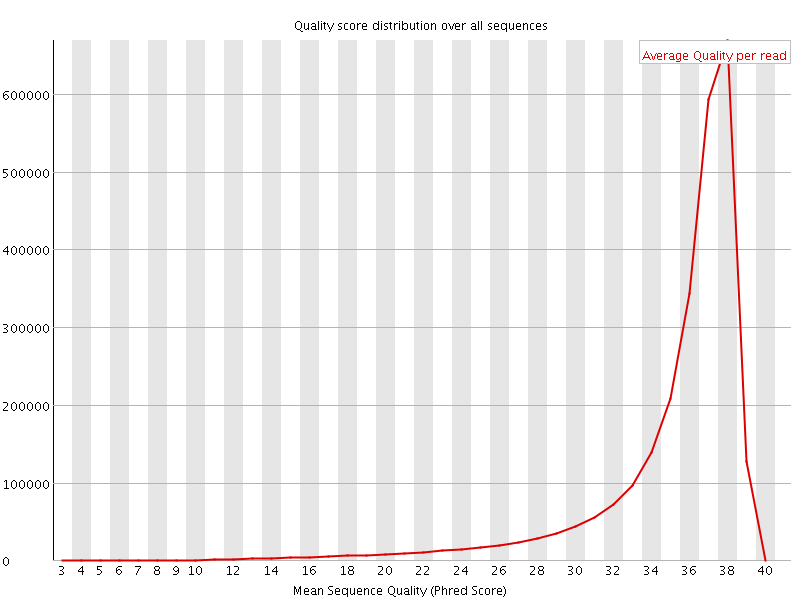
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
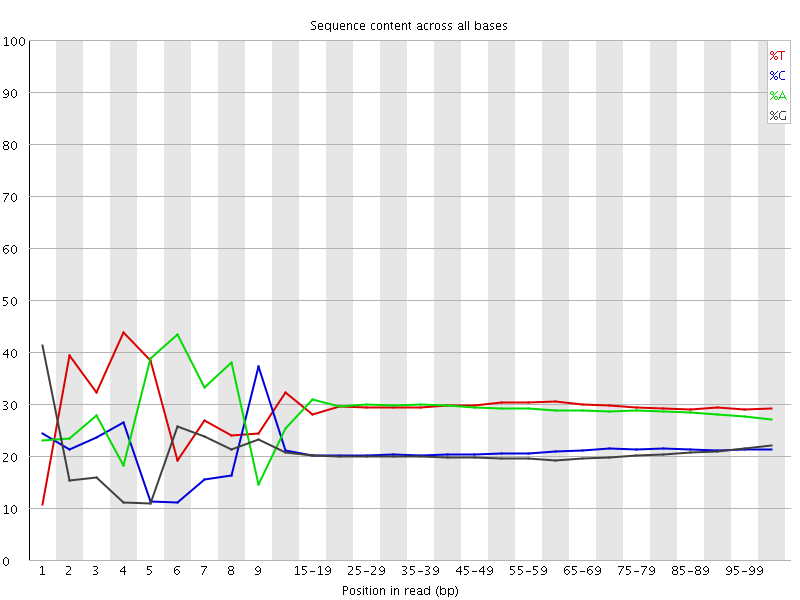
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
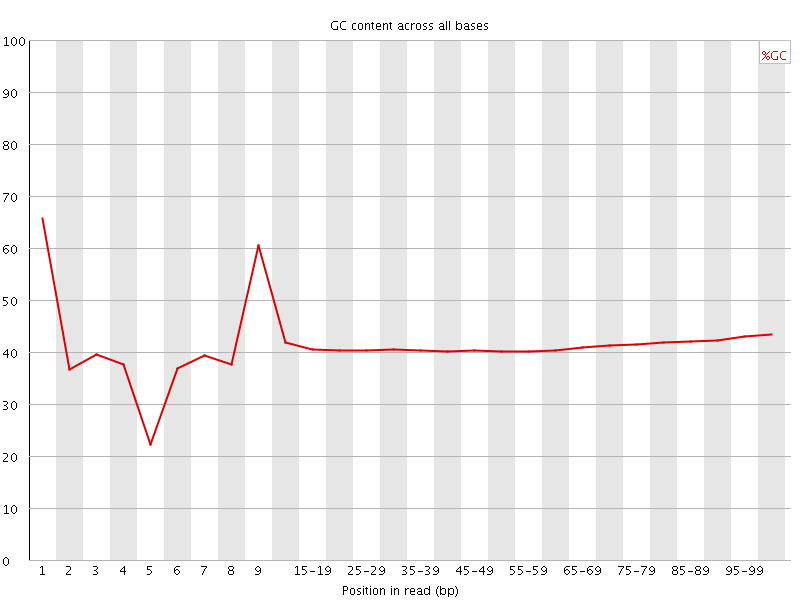
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
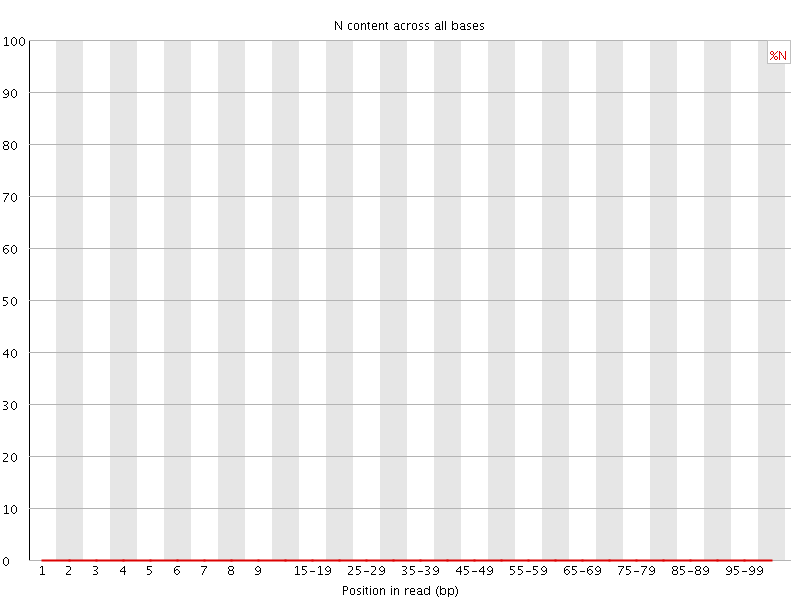
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
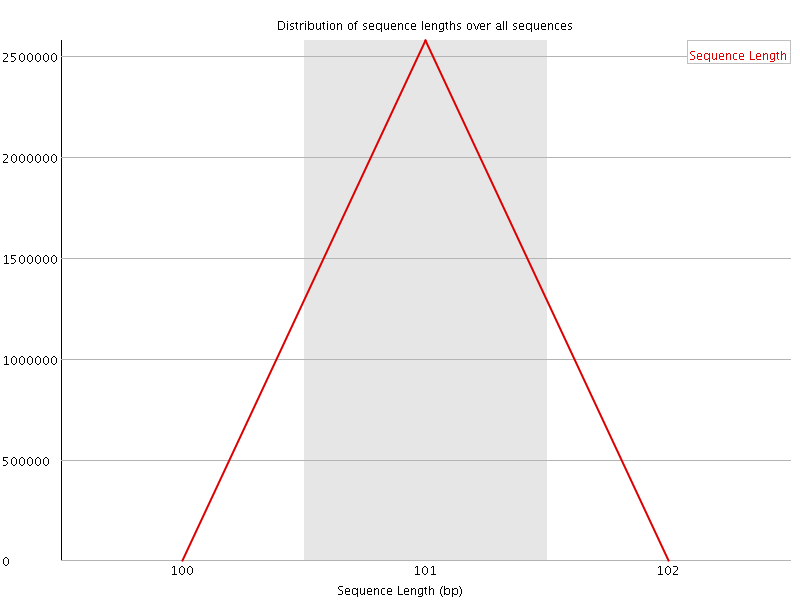
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
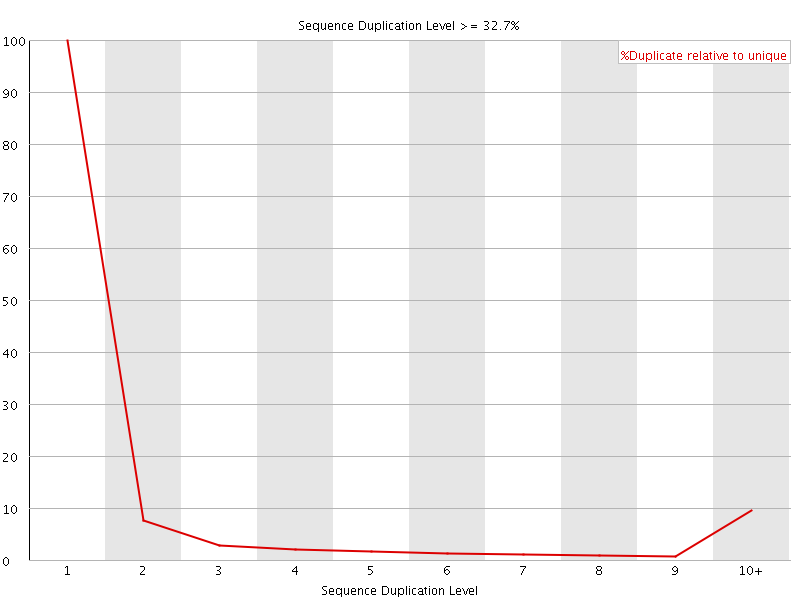
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
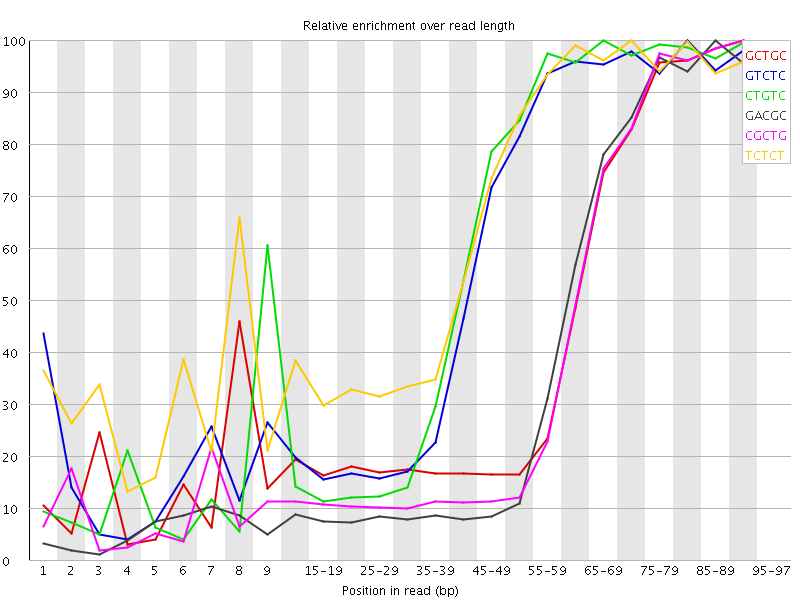
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 496175 | 3.6943593 | 8.34513 | 90-94 |
| GTCTC | 696885 | 3.5472434 | 5.896762 | 80-84 |
| CTGTC | 667985 | 3.4001384 | 5.588844 | 65-69 |
| GACGC | 430165 | 3.2816327 | 8.107093 | 85-89 |
| CGCTG | 440380 | 3.2789278 | 7.9999146 | 90-94 |
| TCTCT | 931535 | 3.241565 | 4.8387957 | 70-74 |
| GCCGA | 415105 | 3.1667435 | 8.287659 | 90-94 |
| CTGCC | 434760 | 3.1476967 | 7.812719 | 90-94 |
| CTCTT | 880550 | 3.064147 | 4.6862226 | 70-74 |
| CGACG | 394915 | 3.0127182 | 8.385633 | 95-97 |
| TGCCG | 397385 | 2.958801 | 7.9110827 | 90-94 |
| CCGAC | 381905 | 2.8330173 | 7.825469 | 95-97 |
| ACACA | 750410 | 2.8087044 | 6.7774925 | 6 |
| GAAGG | 485035 | 2.7411084 | 6.668976 | 85-89 |
| CTGAC | 486465 | 2.537068 | 5.776575 | 95-97 |
| TACAC | 688670 | 2.5157528 | 8.100142 | 5 |
| TGACG | 464035 | 2.4888127 | 5.8894553 | 95-97 |
| GACGA | 423835 | 2.3291044 | 6.2803826 | 95-97 |
| GGCTT | 429390 | 2.2477229 | 5.8801255 | 95-97 |
| CGAAG | 404690 | 2.2238967 | 6.0351734 | 95-97 |
| ACGCT | 424395 | 2.2133534 | 5.477558 | 85-89 |
| ATACA | 808045 | 2.1263244 | 5.136262 | 6 |
| AAGGC | 380580 | 2.0914047 | 5.711941 | 90-94 |
| AGGCT | 380420 | 2.040351 | 5.6997056 | 90-94 |
| TATAC | 721635 | 1.8533651 | 5.9664063 | 5 |
| CTTTG | 513070 | 1.8360865 | 7.7940598 | 9 |
| GGTGG | 219845 | 1.7311821 | 5.326646 | 95-97 |
| AGGTG | 284255 | 1.5678719 | 5.263052 | 95-97 |
| GAGAG | 267940 | 1.514226 | 5.17158 | 7 |
| GTGTA | 351570 | 1.3256836 | 8.917418 | 1 |
| AAGAC | 321670 | 1.2381662 | 5.388767 | 5 |
| GAGTC | 219515 | 1.1773503 | 5.971968 | 9 |
| TATGA | 443535 | 1.1714729 | 5.8142467 | 4 |
| TCATA | 452195 | 1.1613661 | 5.788272 | 2 |
| GATTG | 301835 | 1.1381453 | 5.443838 | 7 |
| GTCTT | 317585 | 1.1365185 | 5.8052273 | 1 |
| GACTT | 301885 | 1.1069008 | 5.503336 | 7 |
| GTGTT | 293500 | 1.0801537 | 5.2541704 | 1 |
| ACATG | 275425 | 1.034716 | 5.084922 | 8 |
| TGGAC | 192635 | 1.0331817 | 6.6032495 | 5 |
| GTCCA | 192110 | 1.0019141 | 6.9852734 | 1 |
| GGACT | 172805 | 0.92682517 | 5.97954 | 6 |
| TATAA | 431370 | 0.79805076 | 5.0570917 | 2 |
| GTATA | 263795 | 0.6967403 | 7.0841446 | 1 |