![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-184_AAGAGGCA-CTAAGCCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2577568 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
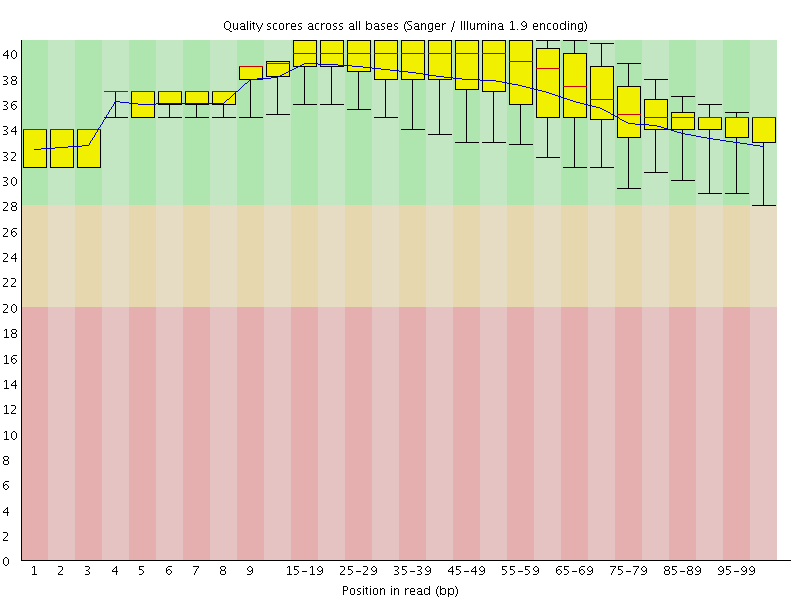
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
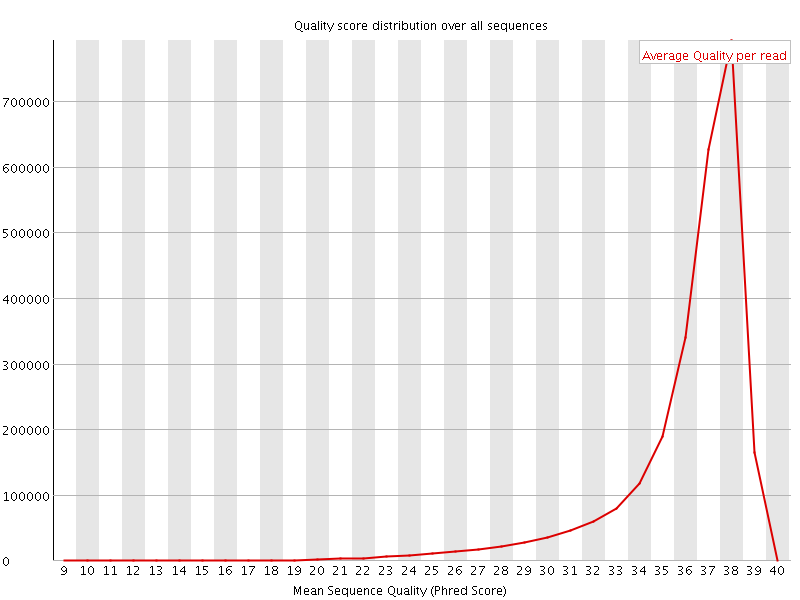
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
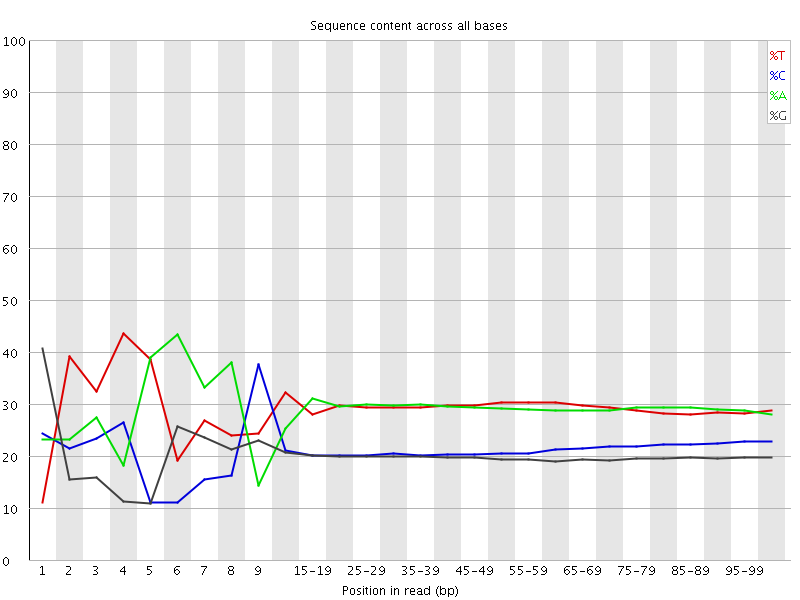
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
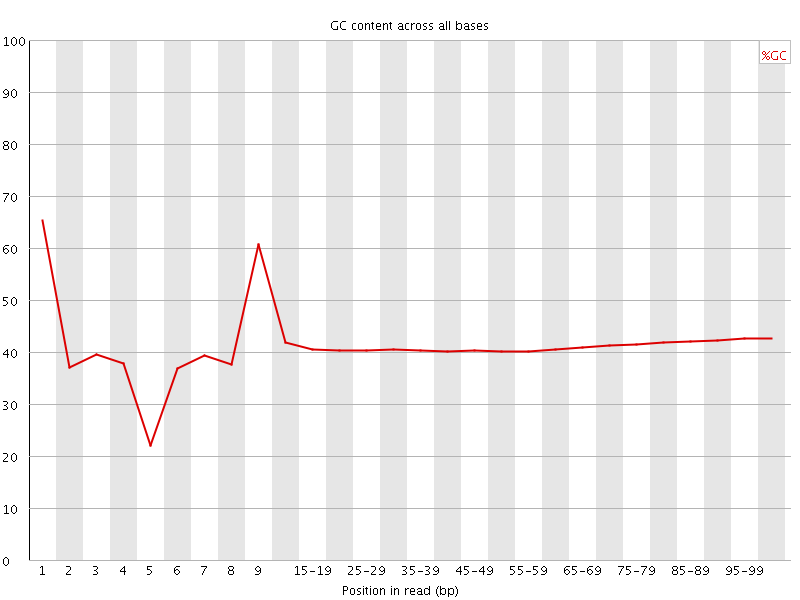
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
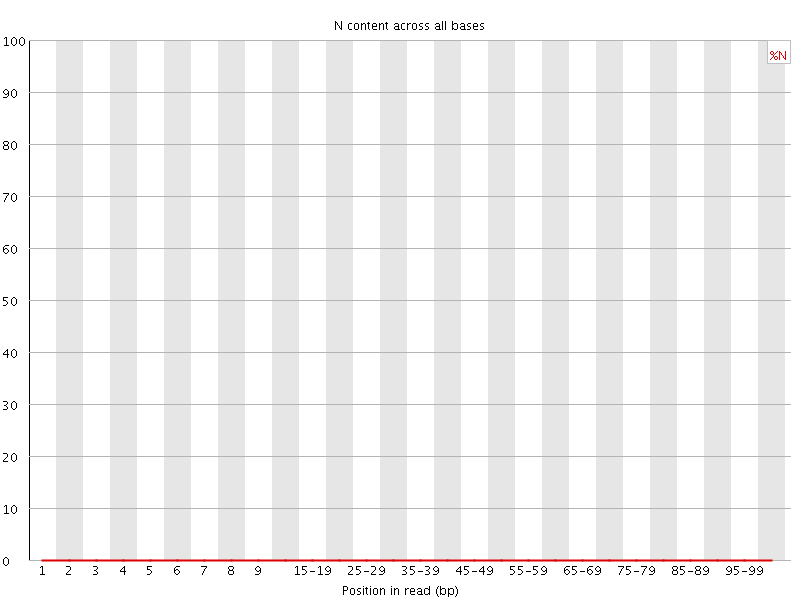
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
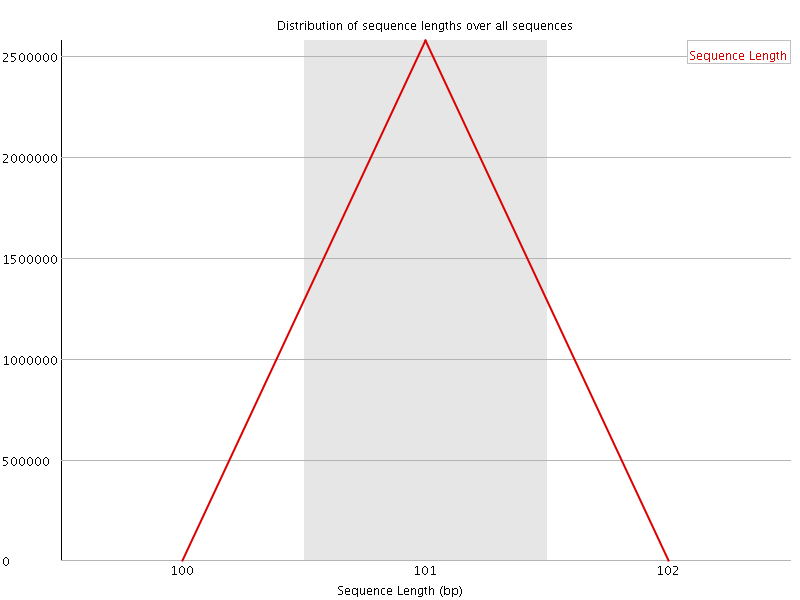
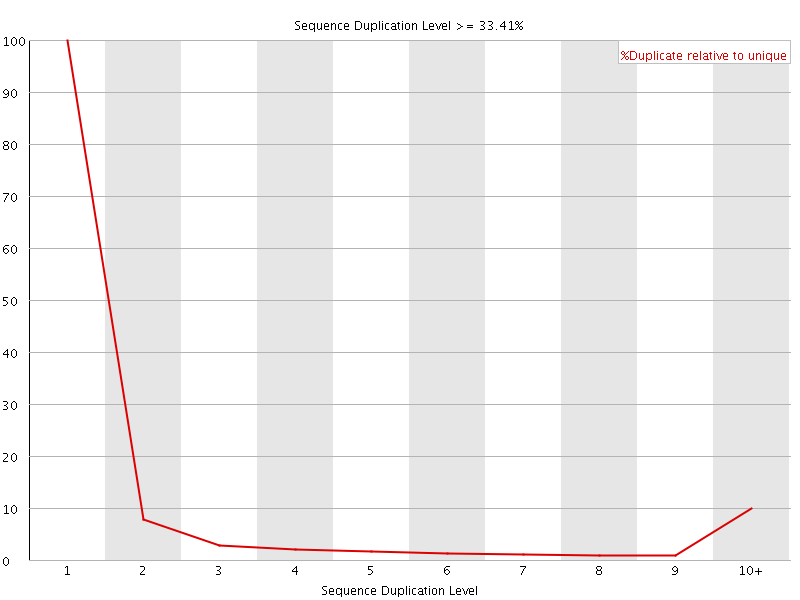
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
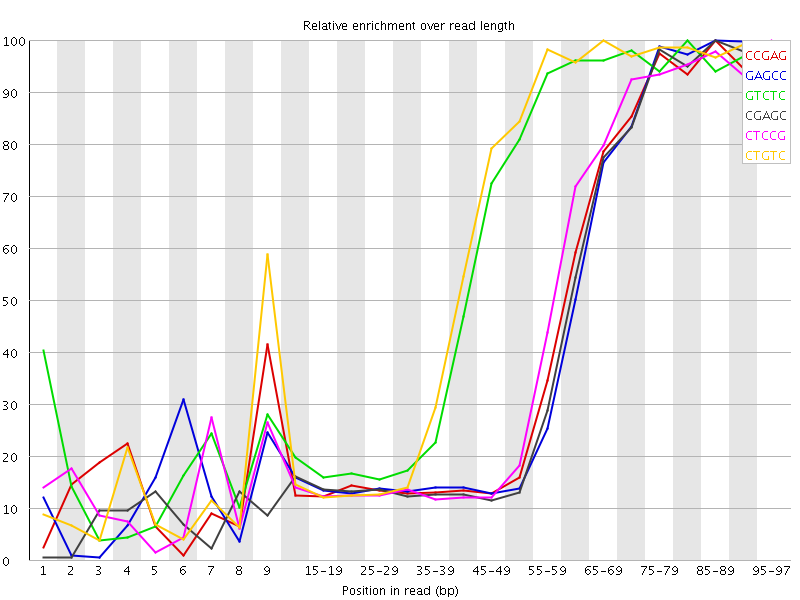
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 485545 | 3.6776097 | 8.394883 | 85-89 |
| GAGCC | 469540 | 3.556384 | 8.252384 | 85-89 |
| GTCTC | 697780 | 3.5488865 | 5.9001255 | 80-84 |
| CGAGC | 465500 | 3.5257847 | 8.272951 | 85-89 |
| CTCCG | 495415 | 3.4918094 | 7.770243 | 95-97 |
| CTGTC | 670785 | 3.4115908 | 5.5972095 | 65-69 |
| TCTCT | 935955 | 3.2221553 | 4.800406 | 70-74 |
| AGCCC | 444590 | 3.1587949 | 7.6507683 | 90-94 |
| GCCCA | 441615 | 3.1376572 | 7.76032 | 90-94 |
| CTCTT | 882395 | 3.0377676 | 4.616122 | 70-74 |
| TCTCC | 629905 | 3.005206 | 5.789335 | 95-97 |
| ATCTC | 811010 | 2.8144796 | 7.16536 | 95-97 |
| CCCAC | 421815 | 2.8113143 | 7.2731 | 90-94 |
| CCACG | 391930 | 2.7846475 | 7.550173 | 90-94 |
| TCCGA | 521535 | 2.6738527 | 5.865024 | 95-97 |
| ACACA | 753460 | 2.6570077 | 6.3483562 | 6 |
| GAGGC | 313985 | 2.5352416 | 8.517837 | 95-97 |
| TACAC | 692705 | 2.423263 | 7.8598084 | 5 |
| CGAGA | 436095 | 2.4026463 | 6.280408 | 95-97 |
| AGAGG | 406835 | 2.389469 | 6.6645365 | 90-94 |
| GAGAC | 429645 | 2.36711 | 6.3026686 | 95-97 |
| ACGAG | 415495 | 2.2891512 | 6.09252 | 95-97 |
| AAGAG | 569820 | 2.2835948 | 5.0864935 | 90-94 |
| CACGA | 402630 | 2.0808487 | 5.5405083 | 95-97 |
| CTTTG | 513085 | 1.8830211 | 8.044593 | 9 |
| TATAC | 723370 | 1.8260179 | 6.1978626 | 5 |
| AGGCA | 315655 | 1.7390872 | 5.7514744 | 95-97 |
| GGCAA | 315050 | 1.735754 | 5.8984003 | 95-97 |
| GAGAG | 265700 | 1.560539 | 5.4931192 | 7 |
| GTCTT | 345615 | 1.2684065 | 5.7515707 | 1 |
| CCATG | 241465 | 1.2379645 | 5.0783906 | 9 |
| GAGTC | 225475 | 1.2323292 | 5.919912 | 9 |
| TATGA | 449350 | 1.2092154 | 5.7421427 | 4 |
| AAGAC | 320770 | 1.2058707 | 5.415186 | 5 |
| GATTG | 300710 | 1.1859558 | 5.474516 | 7 |
| GTGTT | 294200 | 1.1510202 | 5.50158 | 1 |
| TCATA | 452030 | 1.1410687 | 5.672177 | 2 |
| TGCCG | 150975 | 1.1343857 | 6.0501294 | 95-97 |
| GACTT | 305300 | 1.1294657 | 5.599933 | 7 |
| GCCGT | 148190 | 1.1134598 | 5.717384 | 95-97 |
| GTTCT | 303335 | 1.1132389 | 5.0397425 | 1 |
| TGGAC | 192620 | 1.0527608 | 7.183922 | 5 |
| GTGTA | 253490 | 0.999727 | 9.274949 | 1 |
| GTCCA | 193905 | 0.9941296 | 6.8017287 | 1 |
| GGACT | 172100 | 0.9406091 | 6.2697 | 6 |
| GTATA | 263830 | 0.70997506 | 6.9706297 | 1 |
| ATACT | 280785 | 0.7087913 | 5.140407 | 6 |