![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-183_AAGAGGCA-AAGGAGTA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2532424 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
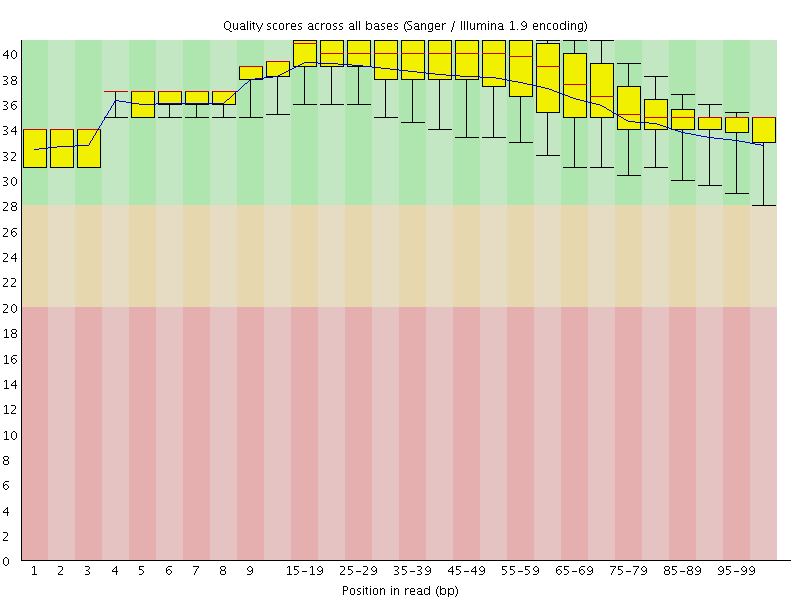
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
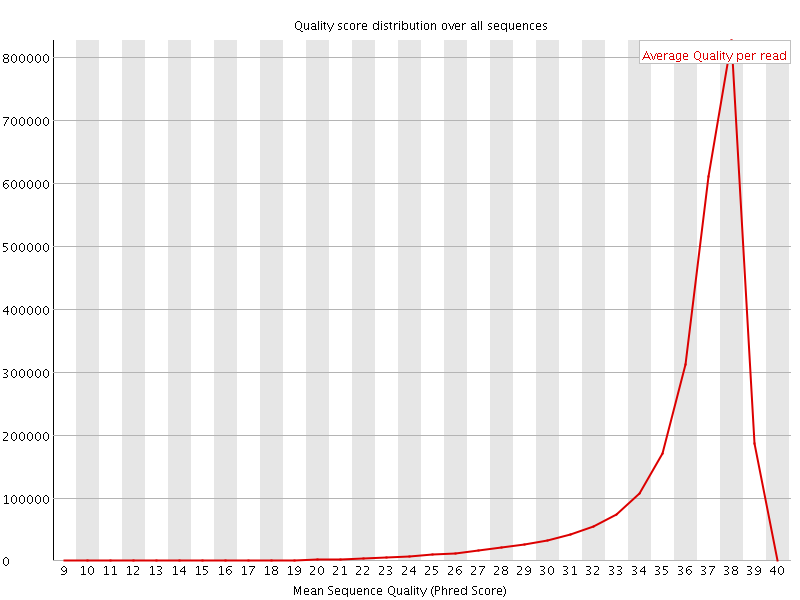
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
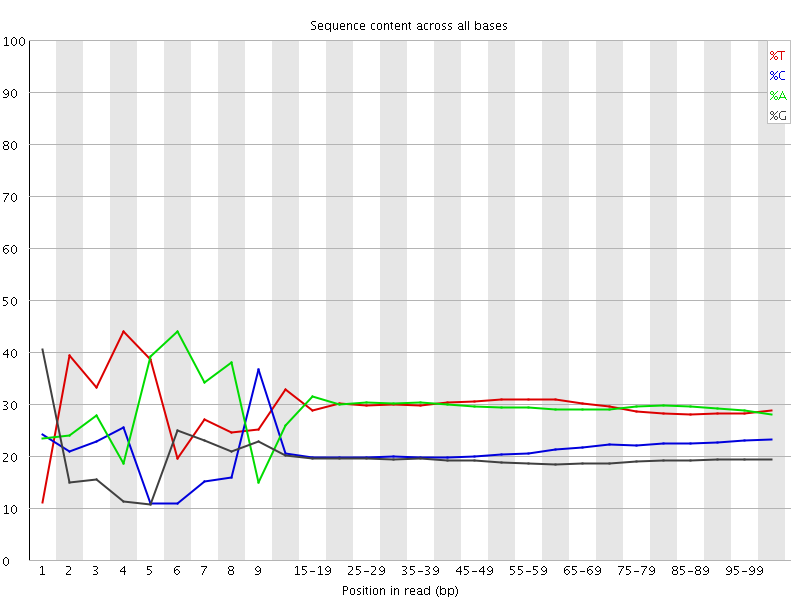
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
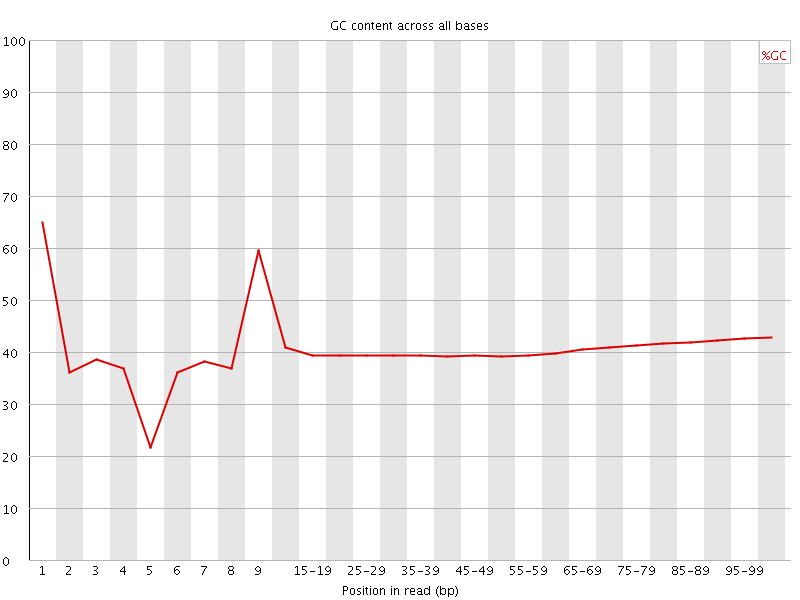
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
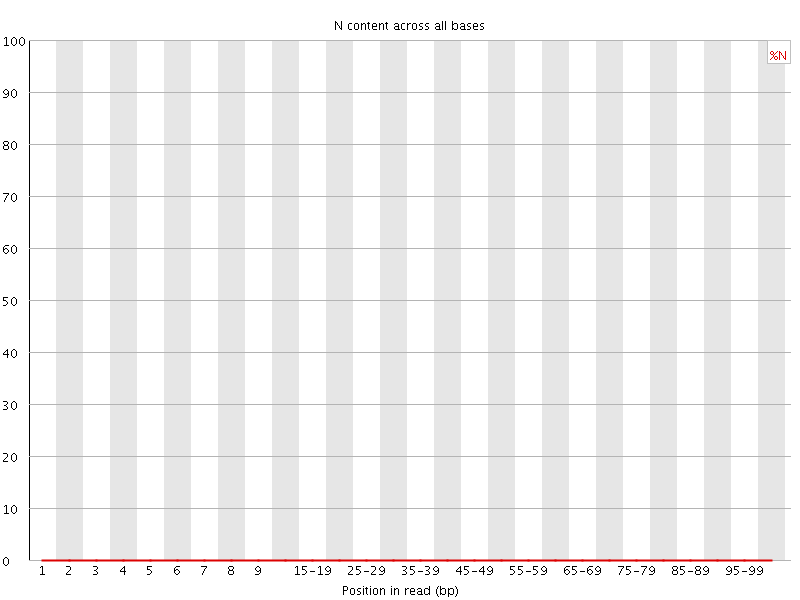
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
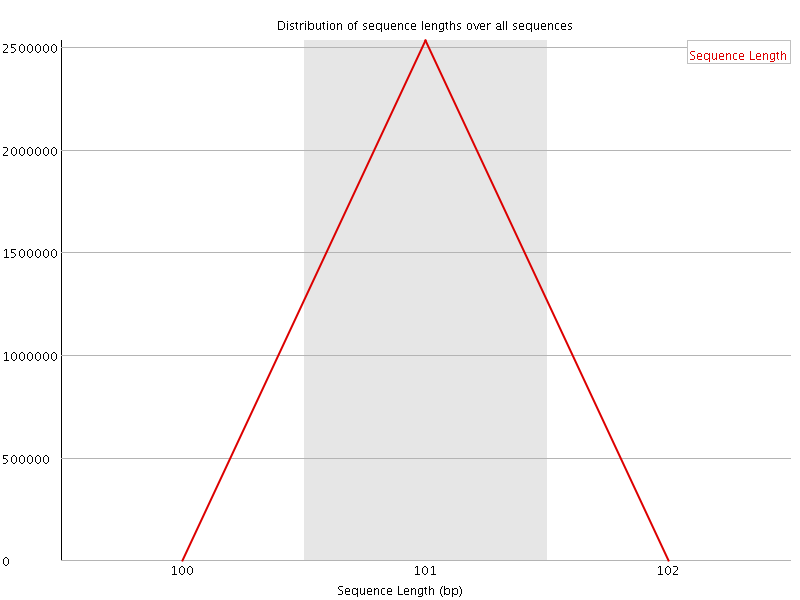
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
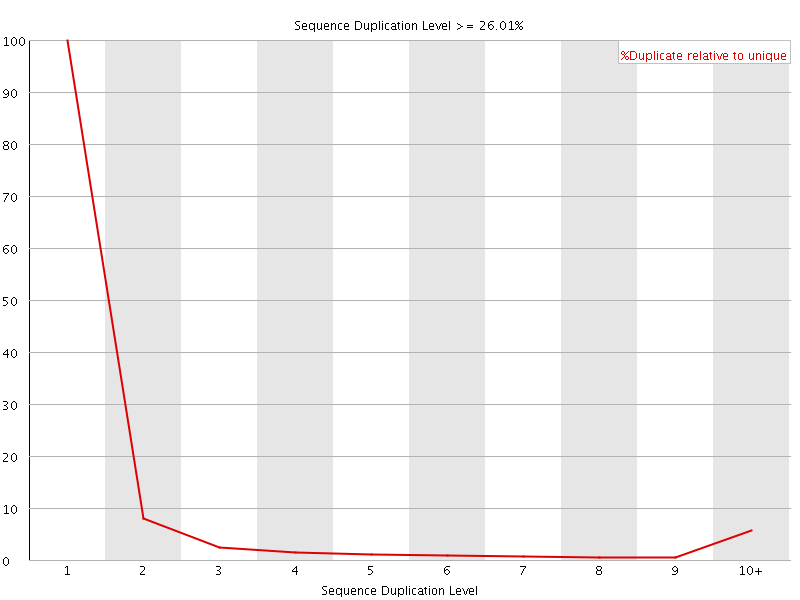
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
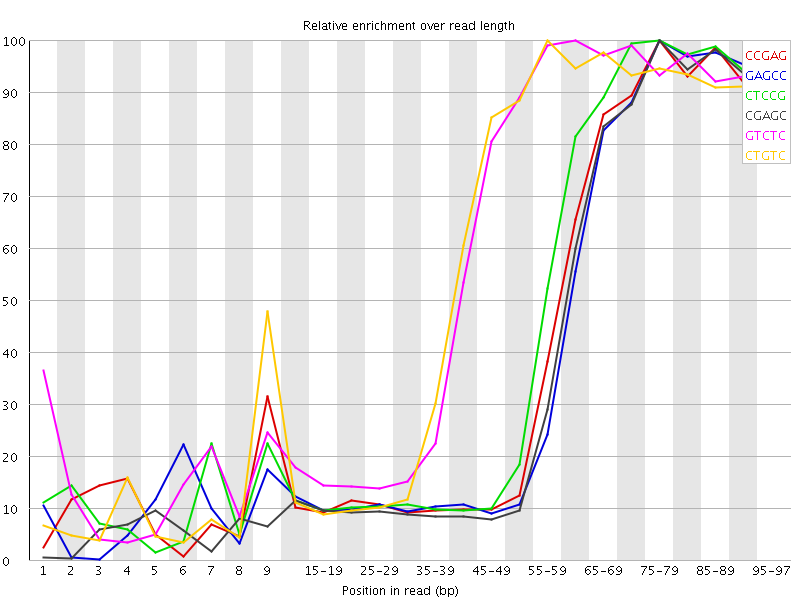
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 574170 | 4.680569 | 10.904313 | 75-79 |
| GAGCC | 549765 | 4.4816227 | 10.764395 | 75-79 |
| CTCCG | 591300 | 4.406412 | 9.577985 | 75-79 |
| CGAGC | 539590 | 4.3986773 | 10.709849 | 75-79 |
| GTCTC | 830980 | 4.381131 | 7.2405815 | 60-64 |
| CTGTC | 791035 | 4.1705313 | 7.06598 | 55-59 |
| AGCCC | 520955 | 3.915521 | 9.627443 | 80-84 |
| GCCCA | 514820 | 3.8694096 | 9.724379 | 80-84 |
| TCTCT | 1091410 | 3.7534714 | 5.706927 | 60-64 |
| TCTCC | 741275 | 3.6033473 | 7.032811 | 70-74 |
| CCACG | 466750 | 3.5081134 | 9.692309 | 80-84 |
| CTCTT | 1018910 | 3.5041363 | 5.4839115 | 60-64 |
| TGTCT | 937495 | 3.4968982 | 5.4518394 | 55-59 |
| CCCAC | 496235 | 3.438807 | 9.00889 | 80-84 |
| ATCTC | 969870 | 3.3641145 | 8.608794 | 95-97 |
| TCCGA | 619155 | 3.2923586 | 7.119997 | 85-89 |
| GAGGC | 362660 | 3.2064667 | 11.51948 | 90-94 |
| ACACA | 865085 | 3.0523908 | 5.9328837 | 6 |
| CGAGA | 515440 | 2.9982426 | 7.893548 | 85-89 |
| GAGAC | 514255 | 2.99135 | 7.997344 | 85-89 |
| AGAGG | 467695 | 2.9506679 | 8.796789 | 90-94 |
| ACGAG | 493635 | 2.8714058 | 7.6772385 | 95-97 |
| TACAC | 800480 | 2.8003983 | 7.2359495 | 5 |
| CATCT | 803670 | 2.7876291 | 5.1350536 | 70-74 |
| AAGAG | 647725 | 2.6884983 | 6.3239884 | 90-94 |
| CACGA | 480805 | 2.5786283 | 7.027862 | 80-84 |
| CAAGA | 664450 | 2.542802 | 5.8411117 | 90-94 |
| AGAAG | 602165 | 2.4993937 | 5.153878 | 5 |
| ACAAG | 621705 | 2.37922 | 5.5895495 | 85-89 |
| AGACA | 591130 | 2.2622118 | 5.4915724 | 85-89 |
| GACAA | 563695 | 2.1572201 | 5.455944 | 85-89 |
| AGGCA | 364955 | 2.1228926 | 7.499551 | 95-97 |
| GGCAA | 363795 | 2.1161447 | 7.653735 | 95-97 |
| TATAC | 837400 | 2.0726247 | 5.590953 | 5 |
| CTTTG | 507240 | 1.892028 | 7.9972043 | 9 |
| AGAGA | 441890 | 1.8341435 | 5.1880894 | 8 |
| GAGAG | 281470 | 1.7757822 | 6.203416 | 7 |
| TCTCG | 326065 | 1.719095 | 6.20346 | 95-97 |
| CAATC | 475235 | 1.6625615 | 5.0814896 | 95-97 |
| TGGAG | 262835 | 1.6441017 | 5.058793 | 5 |
| GCAAT | 398945 | 1.5137398 | 5.1796007 | 95-97 |
| CTCGT | 279625 | 1.4742519 | 5.9009566 | 95-97 |
| GTCTT | 361125 | 1.3470125 | 6.1135635 | 1 |
| GCCGT | 153775 | 1.2428875 | 8.272991 | 95-97 |
| GAGTC | 215150 | 1.2408462 | 6.079131 | 9 |
| TGCCG | 151775 | 1.2267227 | 8.496375 | 95-97 |
| AAGAC | 320265 | 1.2256309 | 5.560865 | 5 |
| GTGTT | 302205 | 1.2226006 | 5.902946 | 1 |
| CCGTC | 159205 | 1.1864077 | 7.0074167 | 95-97 |
| GATTG | 289665 | 1.1819282 | 5.4978004 | 7 |
| ATGCC | 221600 | 1.1783587 | 5.9137955 | 95-97 |
| TATGA | 428085 | 1.1491761 | 5.3376703 | 4 |
| GACTT | 299640 | 1.1272647 | 5.7493362 | 7 |
| GTTCT | 299325 | 1.1164956 | 5.086195 | 1 |
| TCATA | 429315 | 1.0625852 | 5.1620336 | 2 |
| TGGAC | 176030 | 1.0152273 | 6.6635556 | 5 |
| GTGTA | 248705 | 1.0147978 | 9.13521 | 1 |
| GGACT | 163410 | 0.9424434 | 6.255298 | 6 |
| GTCCA | 176395 | 0.937981 | 6.4901485 | 1 |
| AGACT | 241120 | 0.9148955 | 5.2026305 | 6 |
| GTATA | 260555 | 0.6994489 | 6.7663827 | 1 |