![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-182_AAGAGGCA-ACTGCATA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2797648 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
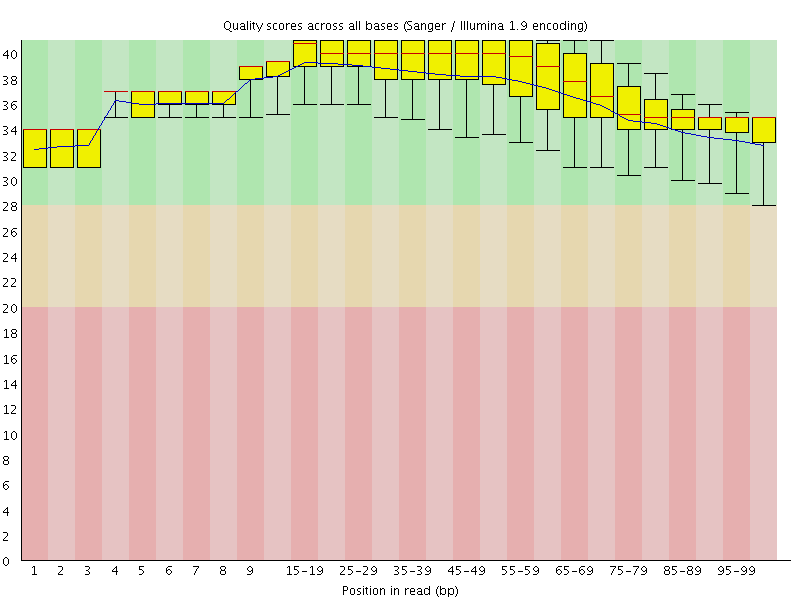
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
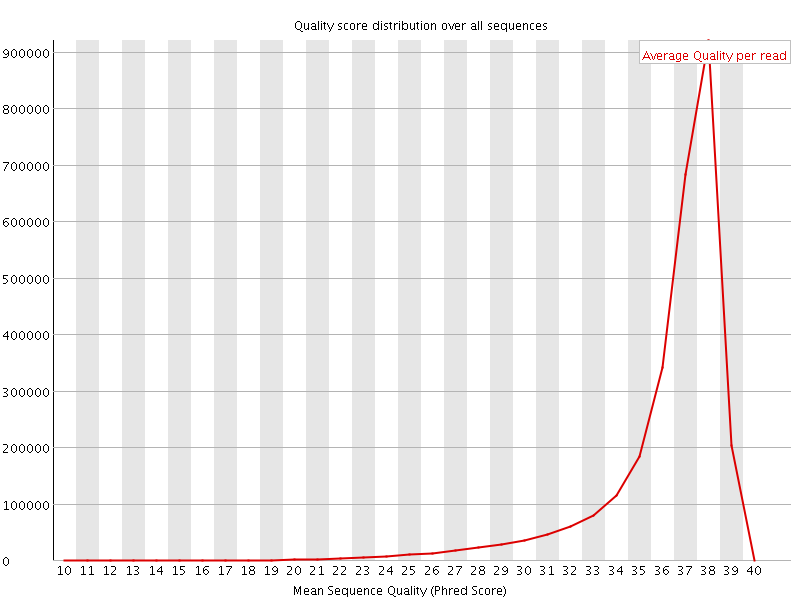
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
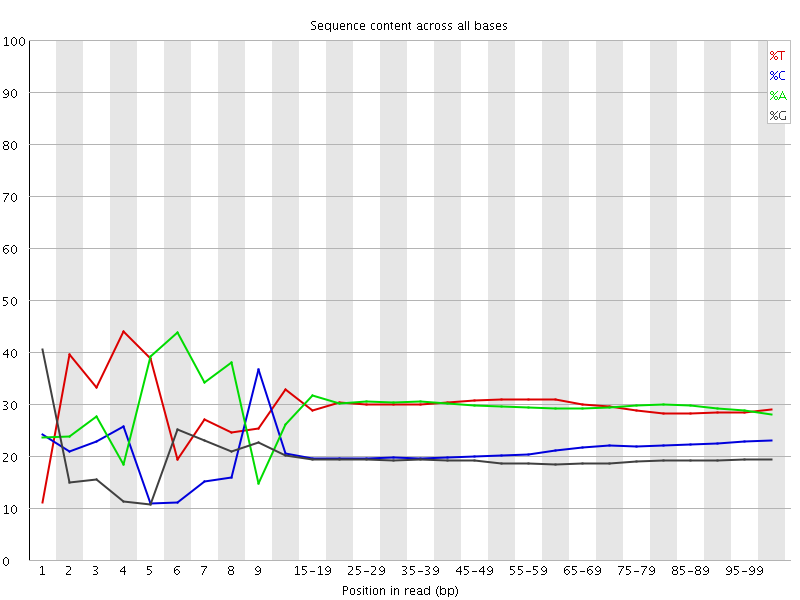
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
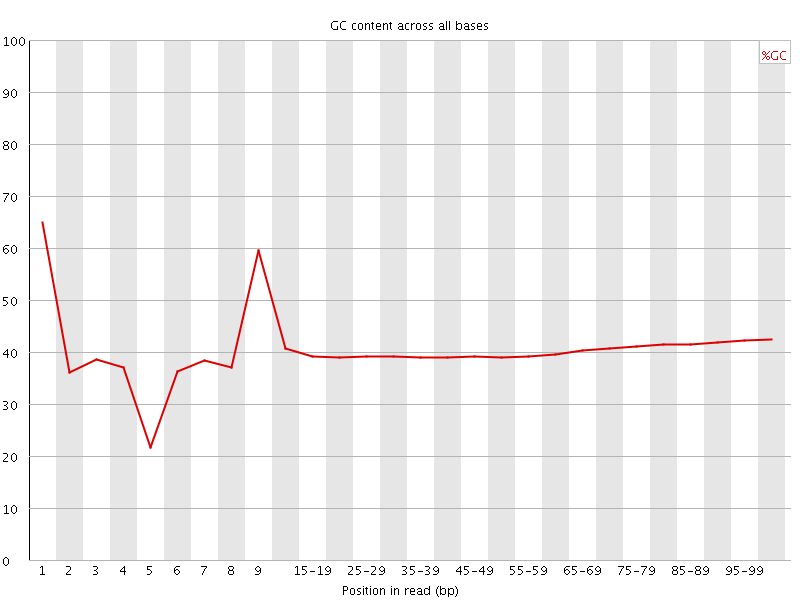
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
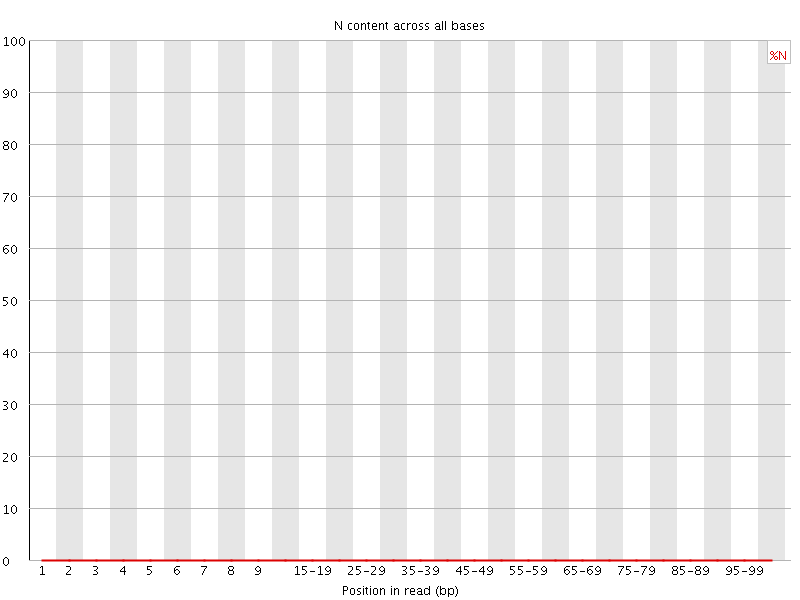
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
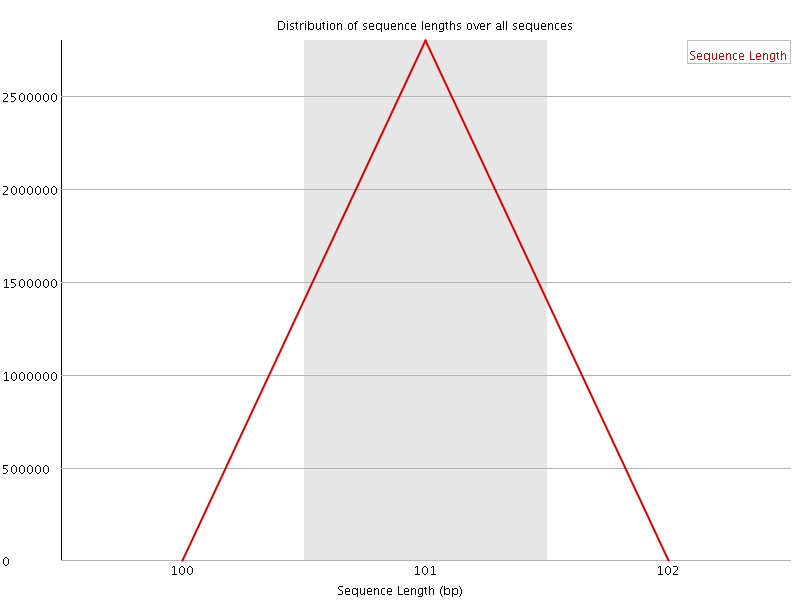
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
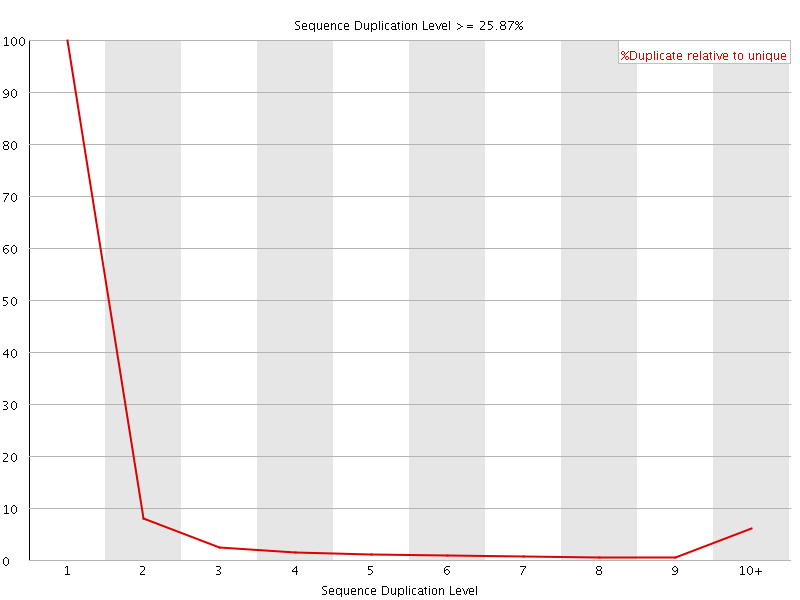
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
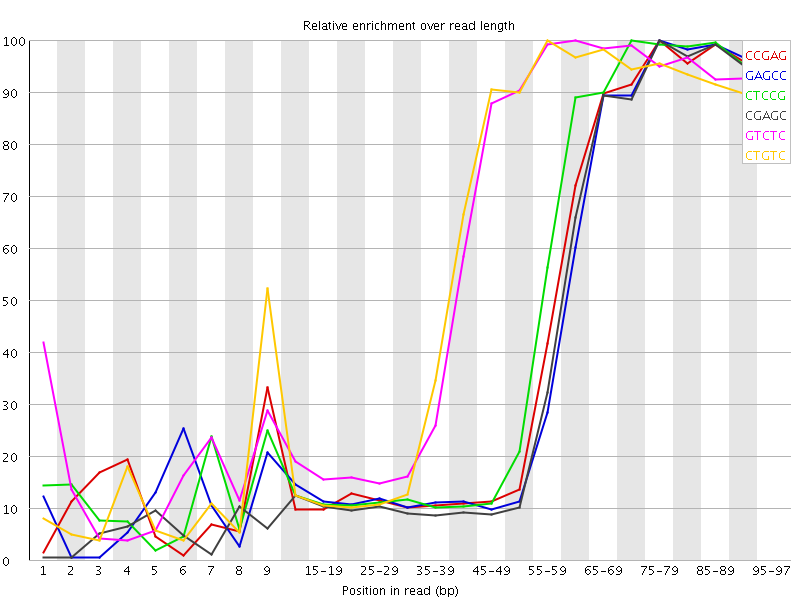
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 601415 | 4.509418 | 10.10378 | 75-79 |
| GAGCC | 582755 | 4.369505 | 10.073241 | 75-79 |
| CTCCG | 621750 | 4.2798133 | 9.034197 | 70-74 |
| CGAGC | 564505 | 4.232667 | 9.964178 | 75-79 |
| GTCTC | 873785 | 4.215232 | 6.7974863 | 60-64 |
| CTGTC | 826985 | 3.9894638 | 6.5856304 | 55-59 |
| AGCCC | 551890 | 3.8273396 | 9.059975 | 80-84 |
| GCCCA | 547390 | 3.7961323 | 9.257018 | 80-84 |
| TCTCT | 1189355 | 3.7190716 | 5.5705385 | 60-64 |
| TCTCC | 795550 | 3.5496264 | 6.643842 | 70-74 |
| CTCTT | 1094590 | 3.4227445 | 5.20779 | 60-64 |
| CCACG | 489675 | 3.39588 | 9.115792 | 80-84 |
| CCCAC | 526430 | 3.376628 | 8.5005045 | 90-94 |
| TGTCT | 988790 | 3.3429422 | 5.1274085 | 55-59 |
| ATCTC | 1043255 | 3.2866154 | 8.1247425 | 95-97 |
| TCCGA | 652925 | 3.1733334 | 6.6830764 | 75-79 |
| GAGGC | 387105 | 3.138173 | 10.783207 | 95-97 |
| GAGAC | 555350 | 2.9400613 | 7.528058 | 85-89 |
| ACACA | 913545 | 2.921186 | 5.7084885 | 6 |
| CGAGA | 551130 | 2.9177206 | 7.4561815 | 85-89 |
| AGAGG | 508390 | 2.9099696 | 8.206238 | 90-94 |
| ACGAG | 524740 | 2.7780097 | 7.1077647 | 80-84 |
| AAGAG | 718695 | 2.6864426 | 5.906742 | 90-94 |
| TACAC | 832365 | 2.6418467 | 6.9218364 | 5 |
| CAAGA | 730830 | 2.526664 | 5.5647745 | 90-94 |
| CACGA | 510820 | 2.5012417 | 6.5559793 | 80-84 |
| AGAAG | 658270 | 2.4605772 | 5.2594266 | 5 |
| ACAAG | 674240 | 2.331018 | 5.2001204 | 85-89 |
| AGACA | 635600 | 2.1974294 | 5.129377 | 85-89 |
| GACAA | 611030 | 2.1124852 | 5.090821 | 85-89 |
| AGGCA | 393995 | 2.0858366 | 7.066706 | 95-97 |
| GGCAA | 389200 | 2.0604517 | 7.1514153 | 95-97 |
| TATAC | 884805 | 1.9681097 | 5.530416 | 5 |
| AGAGA | 522220 | 1.9520297 | 5.413476 | 8 |
| GAGAG | 329735 | 1.8873676 | 6.2886786 | 7 |
| CTTTG | 534005 | 1.8053867 | 7.222332 | 9 |
| TCTCG | 354145 | 1.7084336 | 5.8452687 | 95-97 |
| GCTTT | 448145 | 1.5151072 | 5.0510626 | 1 |
| CTCGT | 306630 | 1.4792161 | 5.655929 | 90-94 |
| AGGAG | 253815 | 1.4528097 | 5.012071 | 7 |
| GTCTT | 395310 | 1.3364804 | 5.8248706 | 1 |
| GCCGT | 165590 | 1.2323811 | 8.386259 | 95-97 |
| TGCCG | 163625 | 1.2177567 | 8.679754 | 95-97 |
| AAGAC | 346615 | 1.1983354 | 5.3723855 | 5 |
| ATGCC | 246535 | 1.1982045 | 5.9526033 | 95-97 |
| CCGTC | 172820 | 1.1896057 | 7.1835613 | 95-97 |
| GAGTC | 226050 | 1.1878421 | 5.177093 | 9 |
| GTGTT | 324475 | 1.1860635 | 5.858203 | 1 |
| GATTG | 319595 | 1.1769611 | 5.128078 | 7 |
| GACTT | 317045 | 1.0798942 | 5.1063175 | 7 |
| TGGAC | 188505 | 0.9905515 | 6.507035 | 5 |
| GTGTA | 258835 | 0.95320225 | 8.757473 | 1 |
| GTCCA | 190895 | 0.9277841 | 6.1959686 | 1 |
| GGACT | 169055 | 0.8883461 | 5.5133996 | 6 |
| GTATA | 285330 | 0.68620044 | 6.622801 | 1 |