![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-179_AAGAGGCA-TATCCTCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2027320 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
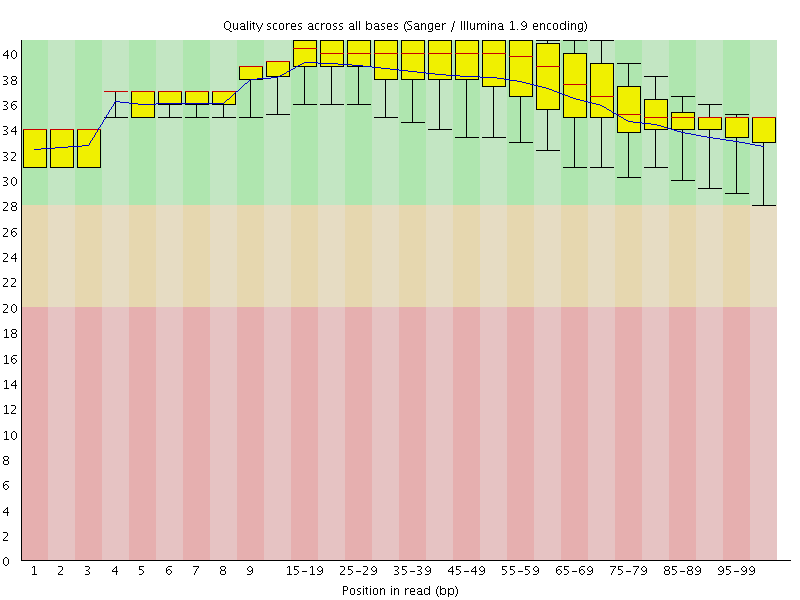
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
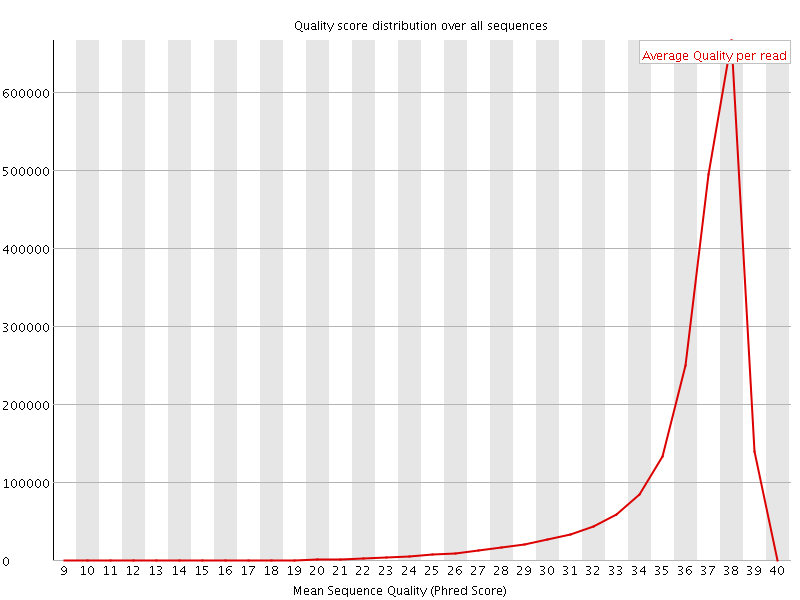
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
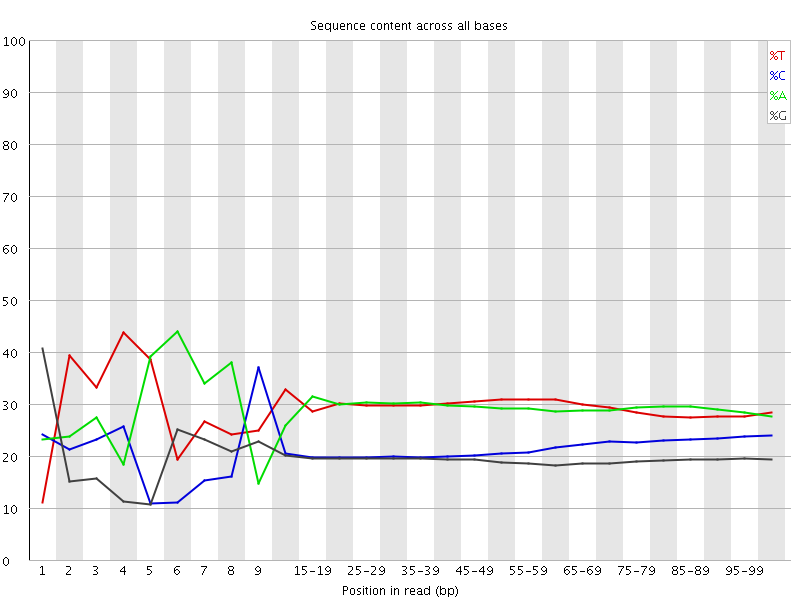
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
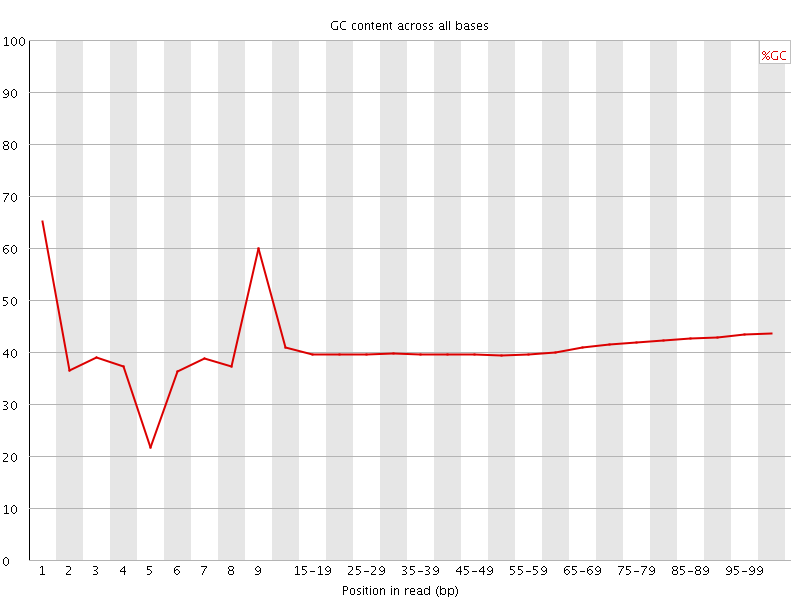
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
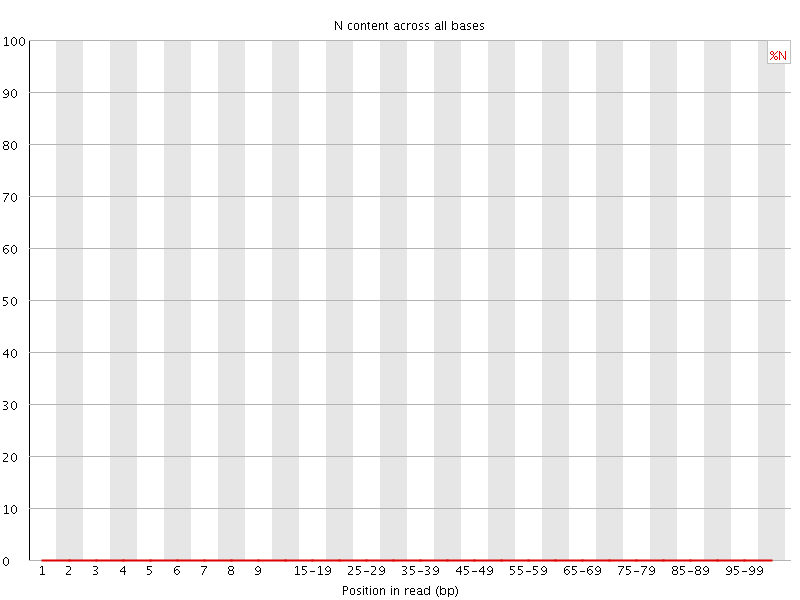
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
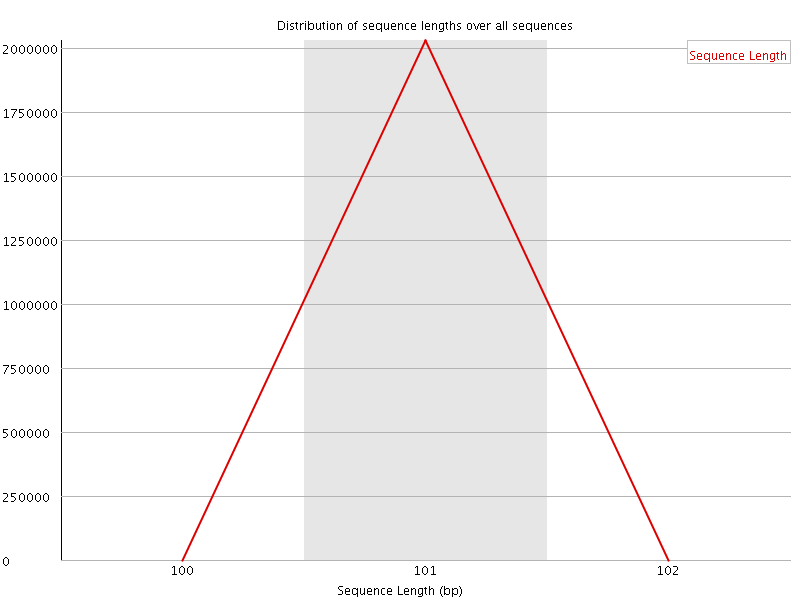
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
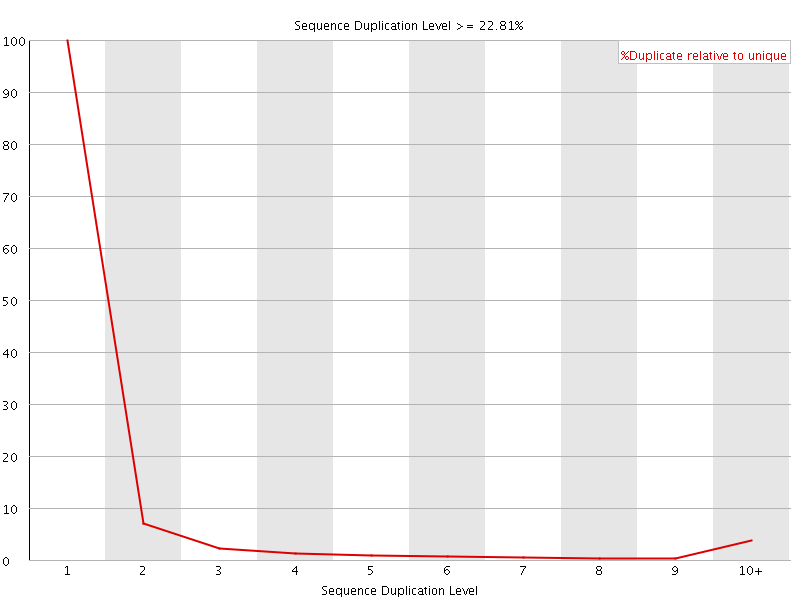
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
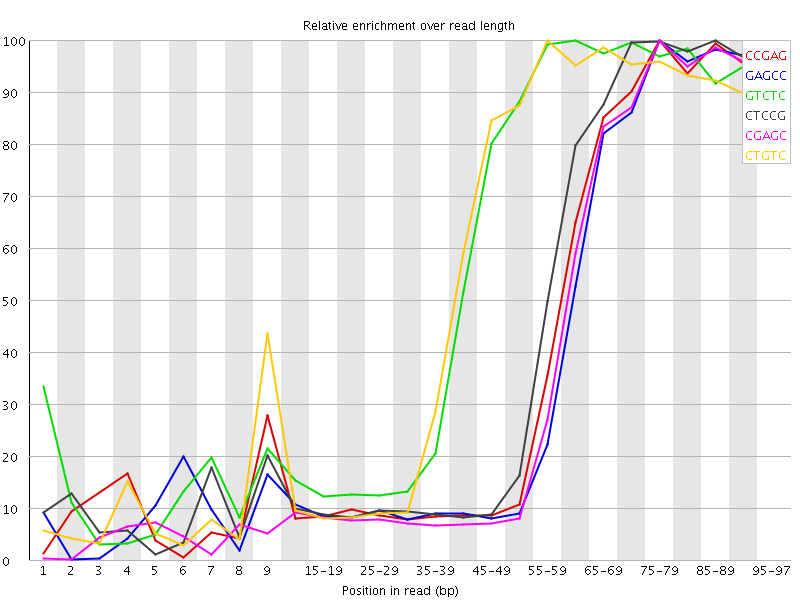
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 516535 | 5.1005235 | 12.0673 | 75-79 |
| GAGCC | 496250 | 4.9002194 | 12.054852 | 75-79 |
| GTCTC | 752830 | 4.8606267 | 8.100113 | 60-64 |
| CTCCG | 538810 | 4.8035593 | 10.624851 | 85-89 |
| CGAGC | 486190 | 4.8008823 | 11.91026 | 75-79 |
| CTGTC | 723830 | 4.673389 | 7.978107 | 55-59 |
| AGCCC | 471235 | 4.230601 | 10.639269 | 90-94 |
| GCCCA | 466270 | 4.1860275 | 10.802951 | 90-94 |
| TCTCT | 969270 | 4.120572 | 6.406086 | 60-64 |
| TGTCT | 839235 | 3.924161 | 6.2720942 | 65-69 |
| TCTCC | 662460 | 3.8887007 | 7.707244 | 70-74 |
| CTCTT | 899580 | 3.8243053 | 6.1374125 | 60-64 |
| CCACG | 422745 | 3.7952733 | 10.724143 | 80-84 |
| ATCTC | 861130 | 3.6865363 | 9.787437 | 95-97 |
| CCCAC | 450755 | 3.679211 | 9.811117 | 80-84 |
| TCCGA | 556865 | 3.6206148 | 8.0399885 | 85-89 |
| GAGGC | 320870 | 3.4849346 | 13.056389 | 90-94 |
| ACACA | 768710 | 3.337232 | 6.1253085 | 6 |
| GAGAC | 457375 | 3.2937634 | 9.105621 | 95-97 |
| CGAGA | 455045 | 3.276984 | 9.010644 | 85-89 |
| AGAGG | 406200 | 3.2174392 | 10.042802 | 90-94 |
| ACGAG | 441245 | 3.177604 | 8.860026 | 95-97 |
| TACAC | 715890 | 3.0862644 | 7.474729 | 5 |
| CATCT | 714000 | 3.0566664 | 5.796075 | 70-74 |
| CACAT | 707940 | 3.0519915 | 5.5527506 | 80-84 |
| ACATC | 685530 | 2.95538 | 5.64778 | 70-74 |
| AAGAG | 554425 | 2.9118412 | 7.2870116 | 90-94 |
| CACGA | 436015 | 2.8547685 | 7.9499855 | 80-84 |
| CAAGA | 571880 | 2.7307334 | 6.6460385 | 90-94 |
| ACAAG | 540990 | 2.5832334 | 6.3534164 | 85-89 |
| AGAAG | 488045 | 2.563213 | 5.1973143 | 5 |
| AGACA | 511805 | 2.4438746 | 6.2043056 | 85-89 |
| GACAA | 491280 | 2.3458672 | 6.1806884 | 85-89 |
| TATAC | 741815 | 2.3160627 | 5.367902 | 5 |
| AGGCA | 320265 | 2.3063722 | 8.616759 | 95-97 |
| GGCAA | 317655 | 2.2875767 | 8.763418 | 95-97 |
| CAAAG | 408890 | 1.9524543 | 5.28325 | 4 |
| AGAGA | 364035 | 1.9119124 | 5.546179 | 8 |
| CTTTG | 402405 | 1.8815969 | 8.052807 | 9 |
| GAGAG | 233390 | 1.8486414 | 6.432745 | 7 |
| TCTCG | 277195 | 1.7897019 | 6.840444 | 95-97 |
| CAATC | 402090 | 1.7334453 | 5.621021 | 95-97 |
| GCAAT | 343475 | 1.6286676 | 5.918115 | 95-97 |
| GCTTT | 336940 | 1.5754905 | 5.2014384 | 1 |
| CTCGT | 239710 | 1.5476811 | 6.573296 | 95-97 |
| TTGAG | 283175 | 1.4665797 | 5.3511276 | 9 |
| GTATG | 279055 | 1.4452419 | 5.7105603 | 95-97 |
| GTCTT | 296530 | 1.3865383 | 6.196831 | 1 |
| TGCCG | 128510 | 1.2601287 | 9.409388 | 95-97 |
| GCCGT | 127810 | 1.2532648 | 8.995735 | 95-97 |
| GAGTC | 173835 | 1.2431401 | 6.095549 | 9 |
| GTGTT | 241670 | 1.2429007 | 6.177411 | 1 |
| AAGAC | 260170 | 1.2423147 | 5.7810144 | 5 |
| ATGCC | 189335 | 1.231015 | 6.6688166 | 95-97 |
| GATTG | 228305 | 1.1824048 | 5.84079 | 7 |
| CCATG | 178790 | 1.1624537 | 5.0943046 | 9 |
| CCGTC | 129795 | 1.1571387 | 7.5043783 | 95-97 |
| TATGA | 335645 | 1.1526178 | 5.742612 | 4 |
| GACTT | 239705 | 1.128697 | 5.538628 | 7 |
| GTTCT | 240130 | 1.1228187 | 5.2694607 | 1 |
| TCATA | 335320 | 1.0469215 | 5.2792645 | 2 |
| TATGC | 222080 | 1.0457063 | 5.030579 | 95-97 |
| GTGTA | 197190 | 1.0212585 | 9.854772 | 1 |
| TGGAC | 137355 | 0.982262 | 6.8063498 | 5 |
| AGACT | 196285 | 0.9307316 | 5.1705637 | 6 |
| GGACT | 126990 | 0.9081392 | 6.067811 | 6 |
| CGTCT | 140390 | 0.9064243 | 5.077895 | 95-97 |
| GAGTA | 173490 | 0.90482 | 5.035306 | 1 |
| GTCCA | 138720 | 0.90192723 | 6.192139 | 1 |
| GTATA | 199655 | 0.685623 | 6.722499 | 1 |