![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-178_AAGAGGCA-CTCTCTAT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1430012 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
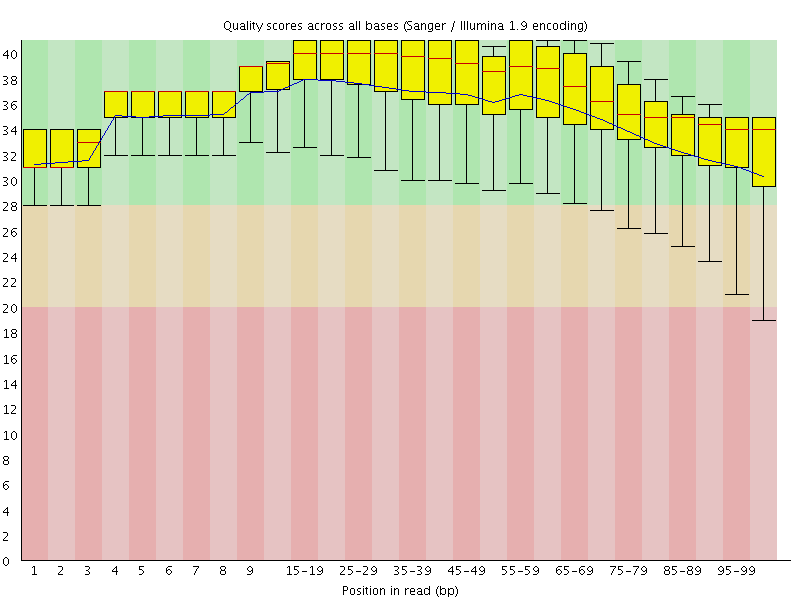
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
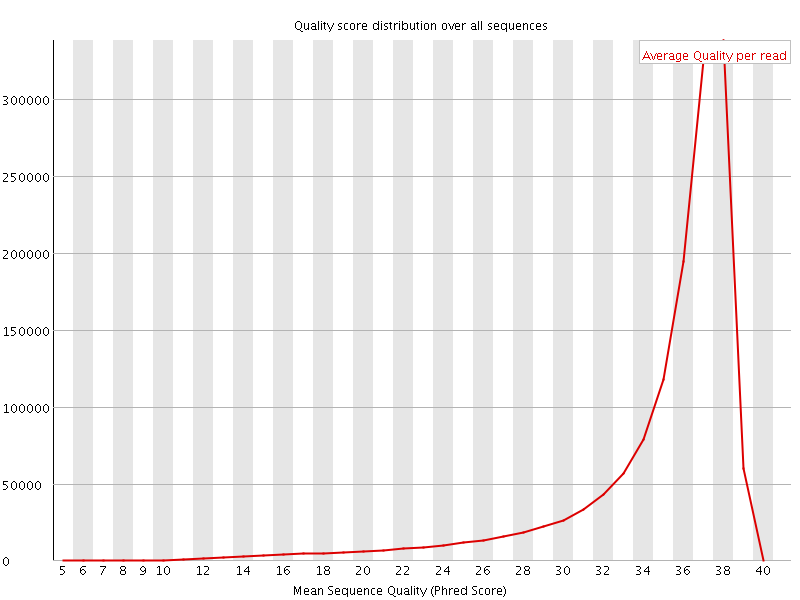
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
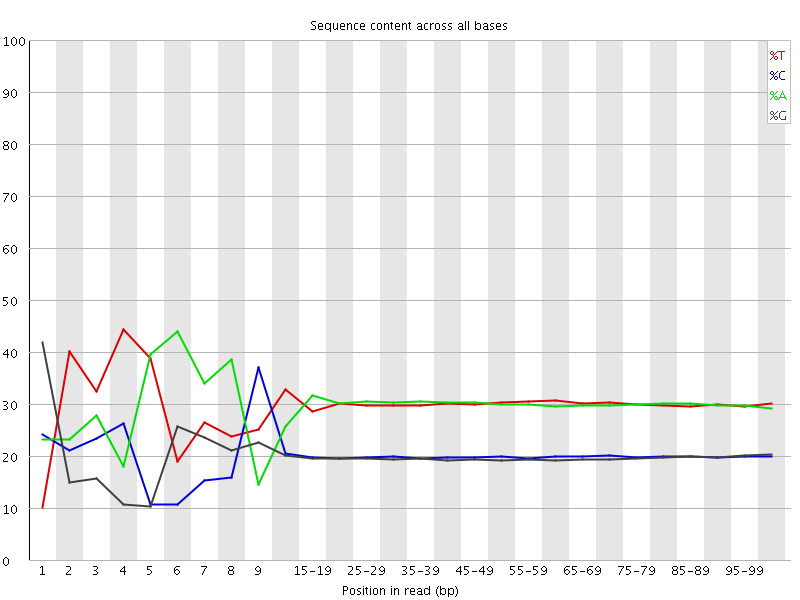
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
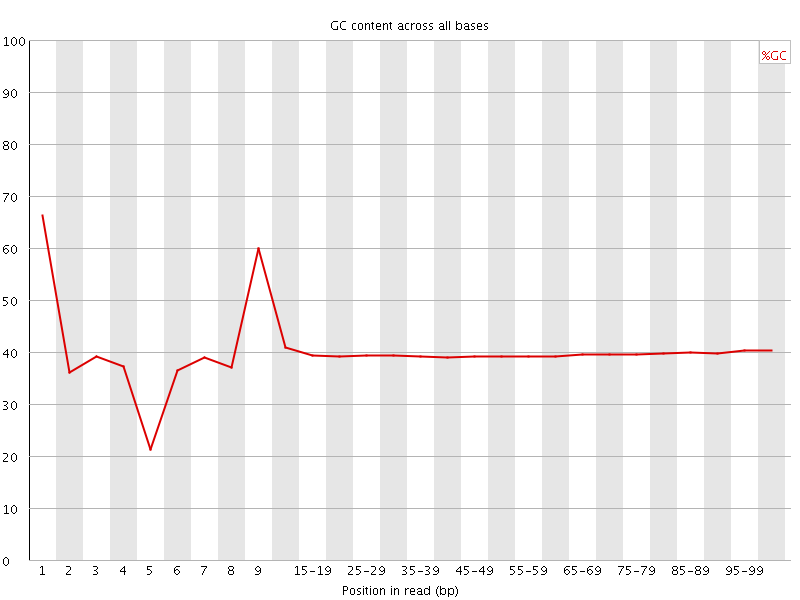
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
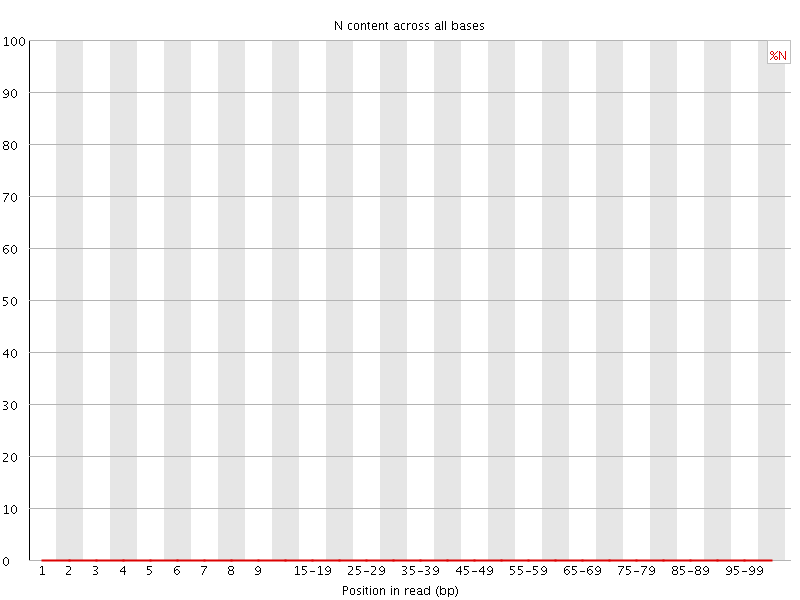
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
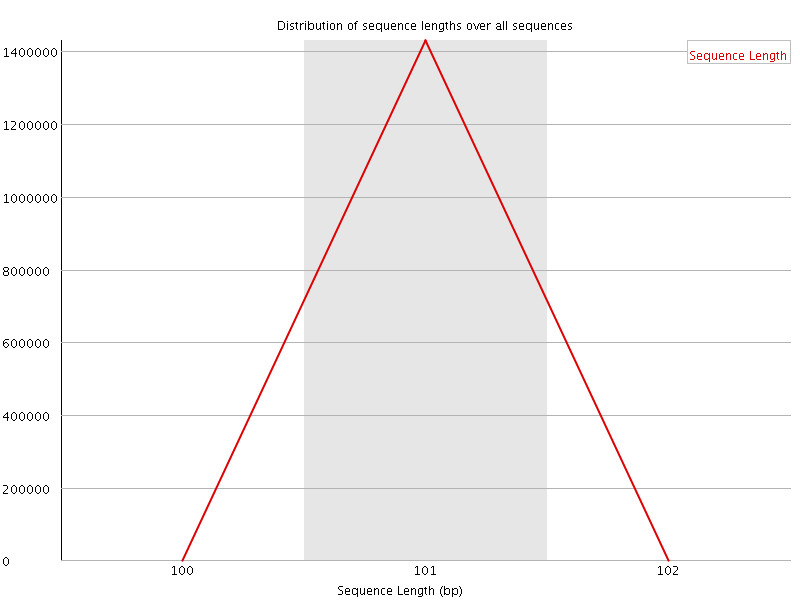
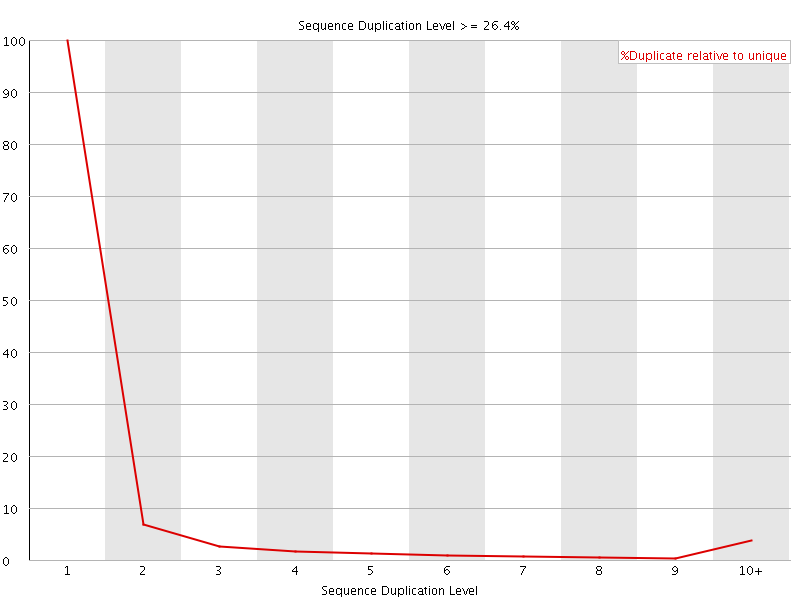
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
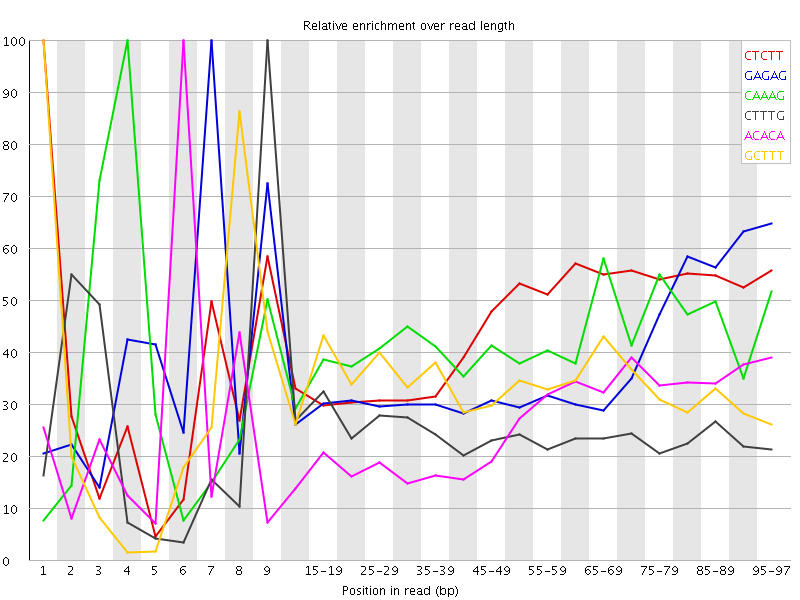
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CTCTT | 351910 | 2.2983491 | 5.1798773 | 1 |
| GAGAG | 215825 | 2.2277563 | 5.931733 | 7 |
| CAAAG | 315595 | 2.1245577 | 5.1006927 | 4 |
| CTTTG | 307700 | 2.0377316 | 8.208792 | 9 |
| ACACA | 298475 | 1.9815782 | 7.48861 | 6 |
| GCTTT | 261735 | 1.7333302 | 5.139926 | 1 |
| CACAT | 255890 | 1.689599 | 5.1841135 | 7 |
| TACAC | 252145 | 1.6648712 | 9.426062 | 5 |
| TGGCT | 153295 | 1.5435219 | 5.029331 | 6 |
| TTGAG | 216555 | 1.4621633 | 5.3634033 | 9 |
| ATACA | 323365 | 1.4317466 | 5.0372024 | 6 |
| GACTC | 140780 | 1.4056058 | 5.12762 | 7 |
| CCATG | 138970 | 1.3875341 | 5.786146 | 9 |
| GAGTC | 133870 | 1.3553175 | 6.27953 | 9 |
| AAGAC | 198965 | 1.3394148 | 5.7929764 | 5 |
| GTCTT | 196825 | 1.3034663 | 6.678692 | 1 |
| GACTT | 184325 | 1.2273735 | 6.167699 | 7 |
| AGAAC | 181470 | 1.22164 | 5.0125246 | 5 |
| GTGTA | 179630 | 1.2128484 | 10.467712 | 1 |
| TATGA | 271570 | 1.2125989 | 5.787374 | 4 |
| GTTCT | 183015 | 1.21201 | 5.3230357 | 1 |
| TCATA | 274795 | 1.210066 | 5.95734 | 2 |
| GATTG | 179185 | 1.2098439 | 5.956042 | 7 |
| GTGTT | 178510 | 1.1987185 | 5.840531 | 1 |
| ACCAT | 175230 | 1.1570145 | 5.011308 | 8 |
| TATAC | 261900 | 1.1532825 | 6.1048098 | 5 |
| ACATG | 167650 | 1.1224554 | 5.2665086 | 8 |
| TGGAC | 110120 | 1.1148694 | 7.410657 | 5 |
| GTCCA | 109735 | 1.0956398 | 7.526463 | 1 |
| ACACT | 153465 | 1.0133038 | 5.1808033 | 6 |
| AGACT | 148730 | 0.9957816 | 5.5649924 | 6 |
| GGACT | 97705 | 0.98917824 | 6.7948847 | 6 |
| GAGTA | 141065 | 0.9576789 | 5.3679147 | 1 |
| GTATG | 139765 | 0.9436829 | 5.272938 | 3 |
| CATAC | 138355 | 0.9135349 | 5.1308804 | 3 |
| GTATA | 166430 | 0.7431337 | 7.3556385 | 1 |
| CCGTG | 49150 | 0.74205893 | 5.0153856 | 9 |