![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-176_CGAGGCTG-CTAAGCCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1485139 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
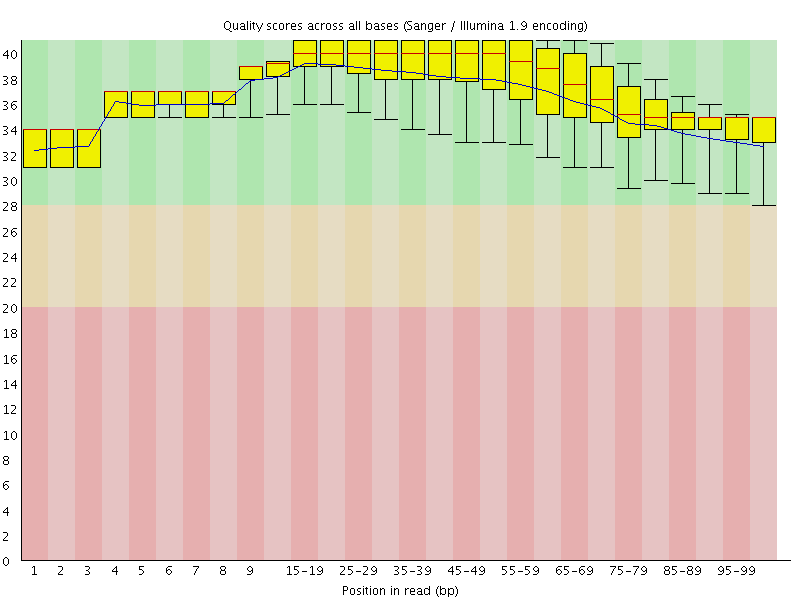
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
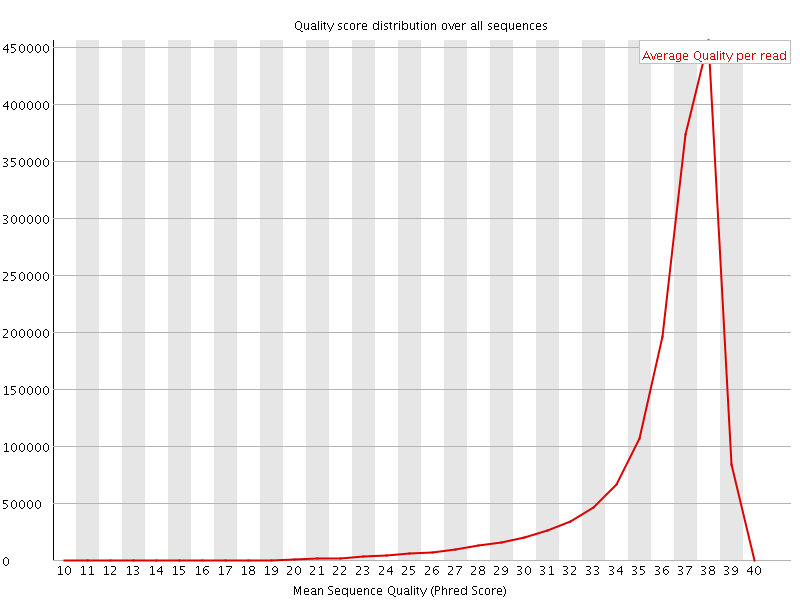
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
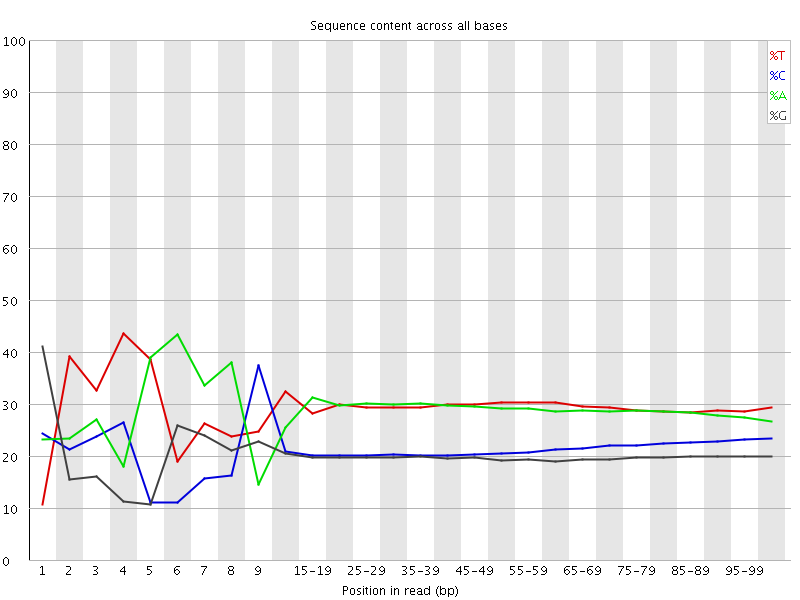
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
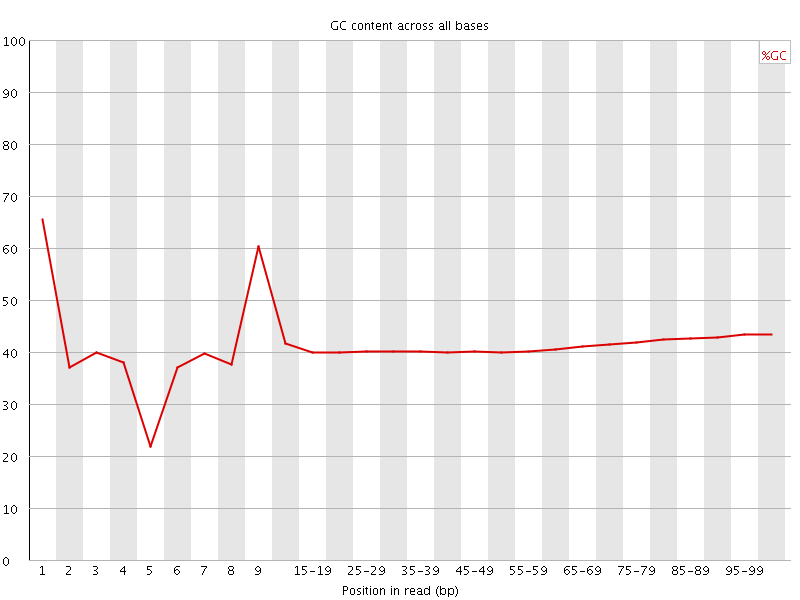
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
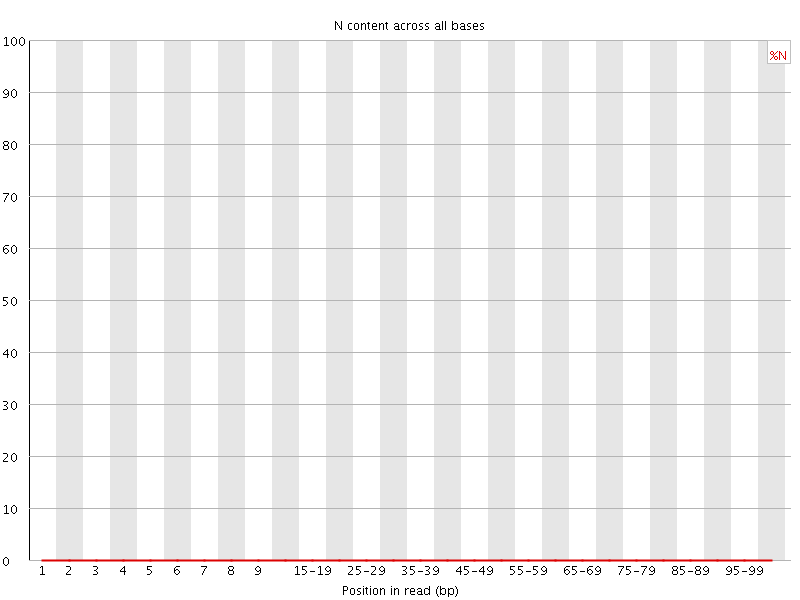
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
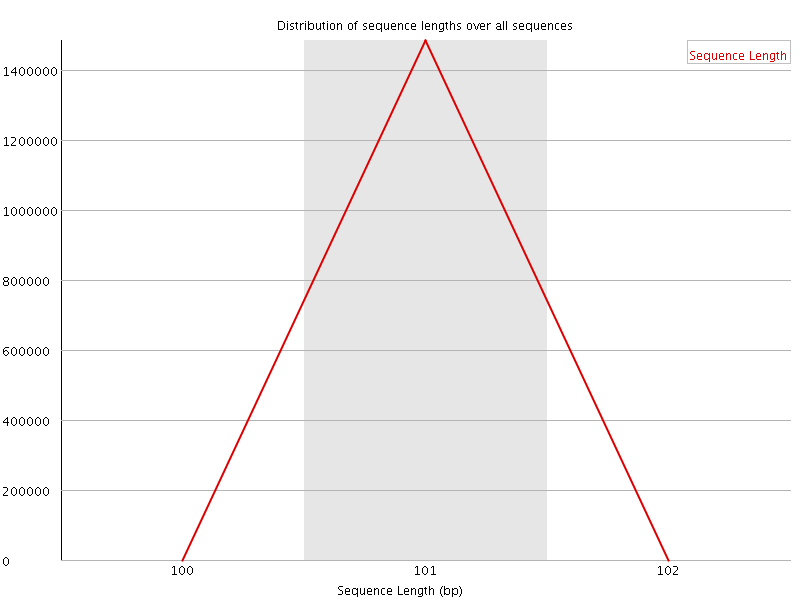
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
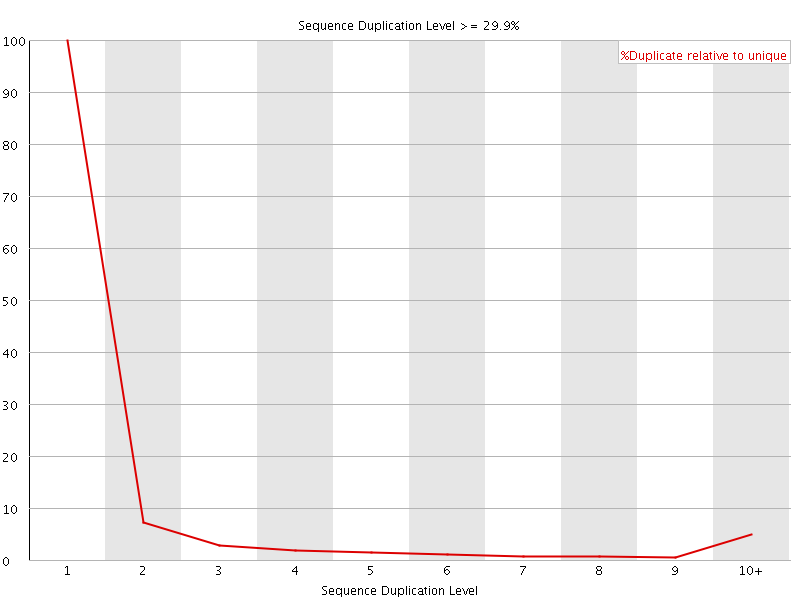
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
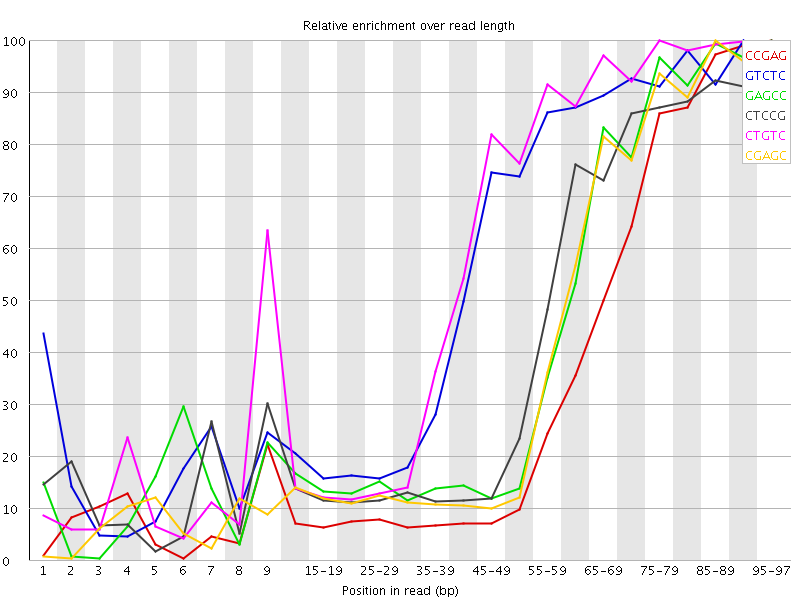
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 398150 | 5.2386084 | 14.8374 | 95-97 |
| GTCTC | 411990 | 3.5803611 | 6.1085596 | 90-94 |
| GAGCC | 266390 | 3.504993 | 8.091774 | 95-97 |
| CTCCG | 288005 | 3.4708793 | 7.949246 | 95-97 |
| CTGTC | 389725 | 3.3868692 | 5.6355367 | 75-79 |
| CGAGC | 256400 | 3.3735504 | 8.115406 | 85-89 |
| TCTCT | 558240 | 3.266164 | 4.8717885 | 90-94 |
| AGCCC | 253890 | 3.118825 | 7.326575 | 90-94 |
| GCCCA | 253780 | 3.1174736 | 7.661334 | 90-94 |
| CTCTT | 522470 | 3.05688 | 4.666774 | 80-84 |
| TCTCC | 370375 | 3.0050936 | 5.8593493 | 95-97 |
| ATCTC | 477860 | 2.849861 | 7.064599 | 95-97 |
| ACACA | 454145 | 2.814034 | 7.162476 | 6 |
| CCCAC | 240250 | 2.755402 | 7.131742 | 90-94 |
| TCCGA | 306015 | 2.710746 | 6.0434 | 95-97 |
| AGAAG | 380350 | 2.703746 | 5.152829 | 5 |
| CCACG | 219435 | 2.695574 | 7.402818 | 90-94 |
| CGAGG | 190160 | 2.6798604 | 8.25214 | 90-94 |
| GGCTG | 189825 | 2.6244633 | 8.3150835 | 95-97 |
| TACAC | 423640 | 2.5752883 | 9.039924 | 5 |
| GAGGC | 181440 | 2.5569727 | 8.161939 | 90-94 |
| CGAGA | 256160 | 2.4773486 | 6.441667 | 95-97 |
| GAGAC | 254140 | 2.4578133 | 6.622991 | 95-97 |
| GACCG | 183870 | 2.4192464 | 7.508028 | 85-89 |
| ACGAG | 240955 | 2.3302996 | 6.1493616 | 95-97 |
| CAAAG | 332105 | 2.204114 | 5.274237 | 4 |
| ATACA | 480095 | 2.145175 | 5.067431 | 6 |
| CACGA | 231325 | 2.0886927 | 5.50436 | 90-94 |
| AGGCT | 214275 | 2.033019 | 5.837321 | 90-94 |
| CTTTG | 323735 | 2.0287592 | 8.951498 | 9 |
| AGACC | 224540 | 2.0274296 | 5.8331895 | 95-97 |
| TATAC | 431400 | 1.8910792 | 6.0044074 | 5 |
| GCTGA | 190285 | 1.8054042 | 5.539078 | 95-97 |
| ACCGA | 197285 | 1.7813368 | 5.0753307 | 85-89 |
| GAGAG | 168465 | 1.7450559 | 5.544863 | 7 |
| GCTTT | 273775 | 1.7156736 | 5.117343 | 1 |
| TTGAG | 220660 | 1.5097123 | 5.8053813 | 9 |
| GTCTT | 221720 | 1.389459 | 6.0593715 | 1 |
| GAGTC | 145485 | 1.3803467 | 6.560117 | 9 |
| AAGAC | 206285 | 1.3690721 | 5.8405995 | 5 |
| CCATG | 148940 | 1.3193421 | 5.433238 | 9 |
| GTATG | 191430 | 1.3097265 | 5.4504237 | 3 |
| TATGA | 276615 | 1.2987607 | 6.2650414 | 4 |
| TGAGT | 186370 | 1.2751069 | 5.2381124 | 8 |
| GACTT | 196960 | 1.258128 | 6.7797656 | 7 |
| TCATA | 280970 | 1.2316563 | 6.3344283 | 2 |
| GTGTT | 183325 | 1.2305135 | 5.9953685 | 1 |
| GATTG | 177930 | 1.217362 | 6.087357 | 7 |
| GTGTA | 171370 | 1.17248 | 10.898521 | 1 |
| GTTCT | 183230 | 1.1482526 | 5.235856 | 1 |
| ACATG | 166845 | 1.0863403 | 5.4142575 | 8 |
| TGGAC | 112960 | 1.0717528 | 7.323777 | 5 |
| GCCGT | 82610 | 1.0663406 | 6.179712 | 95-97 |
| CCGTC | 86635 | 1.0440779 | 5.0995164 | 95-97 |
| GTCCA | 114495 | 1.0142211 | 6.894182 | 1 |
| GGACT | 105440 | 1.0004036 | 6.932746 | 6 |
| AGACT | 153535 | 0.999678 | 5.5310674 | 6 |
| GAGTA | 143335 | 0.9996059 | 5.5866075 | 1 |
| TGCCG | 76405 | 0.9862456 | 6.440503 | 95-97 |
| TGTAT | 213605 | 0.9839179 | 5.050212 | 2 |
| ACACT | 159770 | 0.97123456 | 5.205251 | 6 |
| GTATA | 164920 | 0.77433115 | 7.2036314 | 1 |
| ATACT | 170955 | 0.7493961 | 5.0628314 | 6 |