![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-175_CGAGGCTG-AAGGAGTA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2365119 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
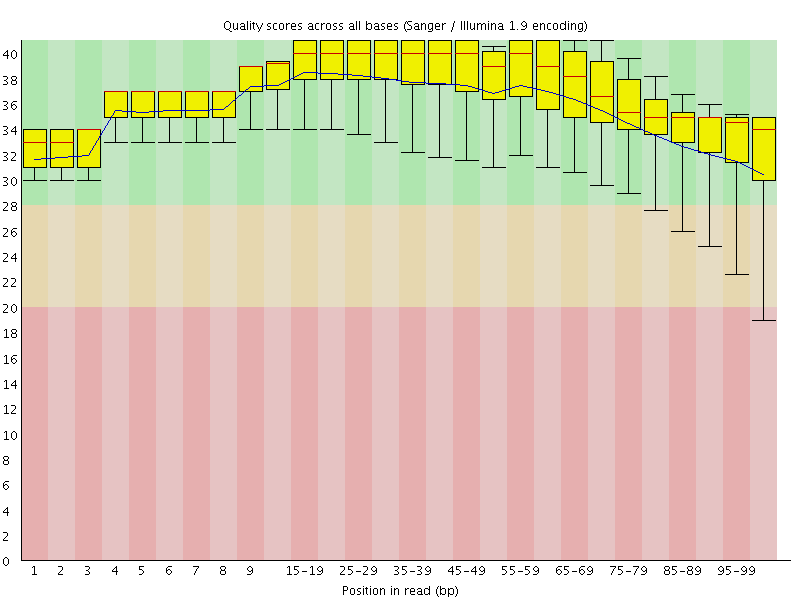
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
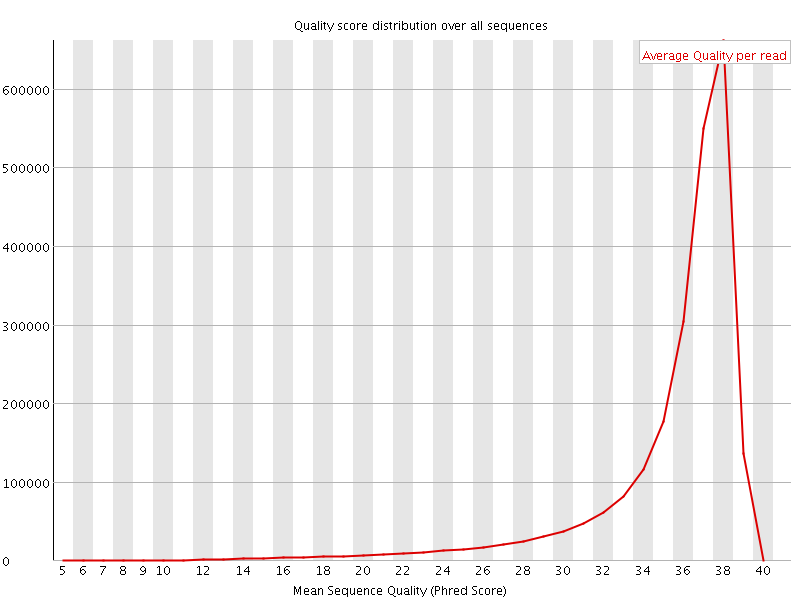
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
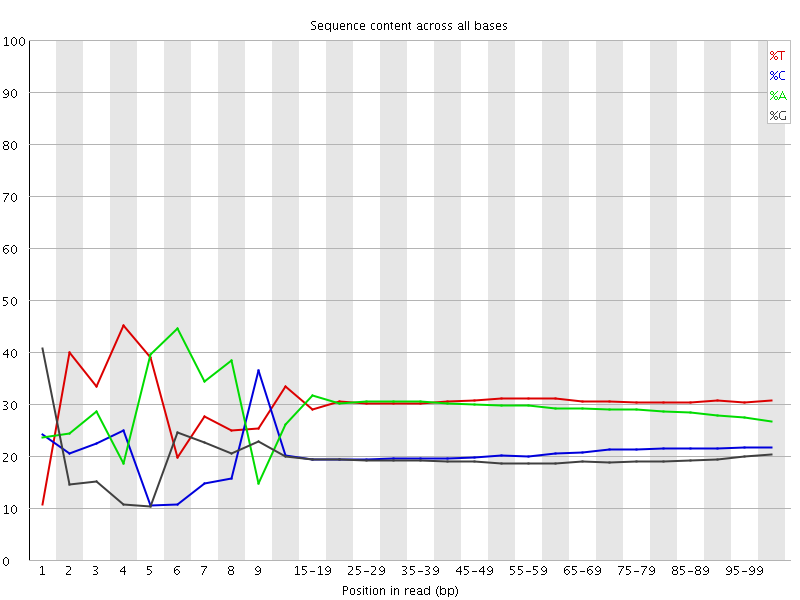
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
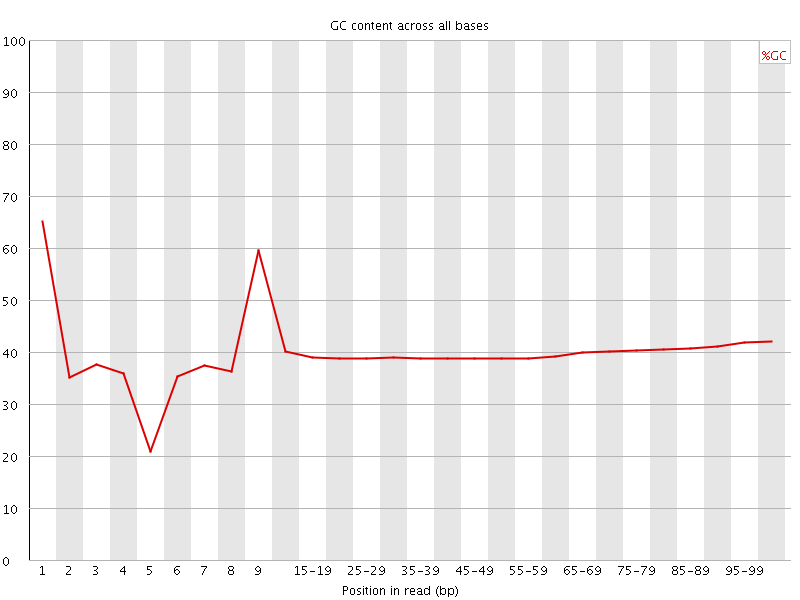
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
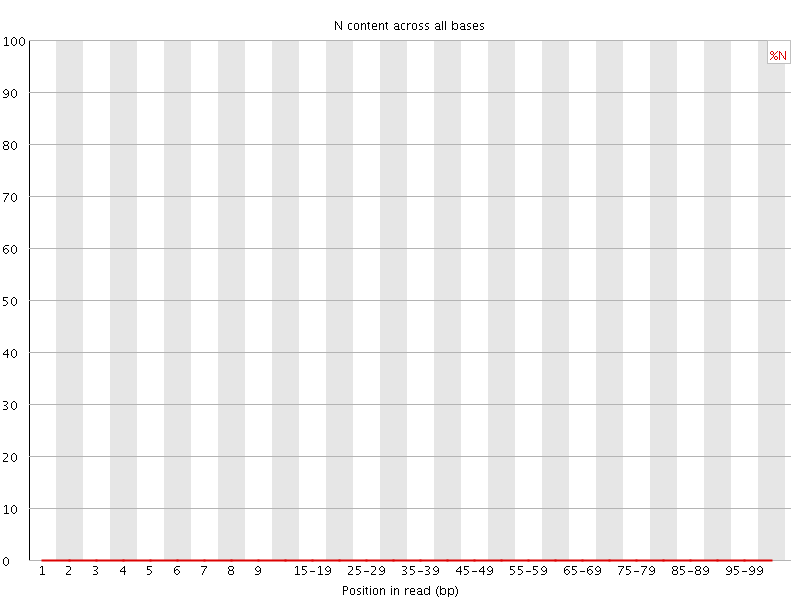
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
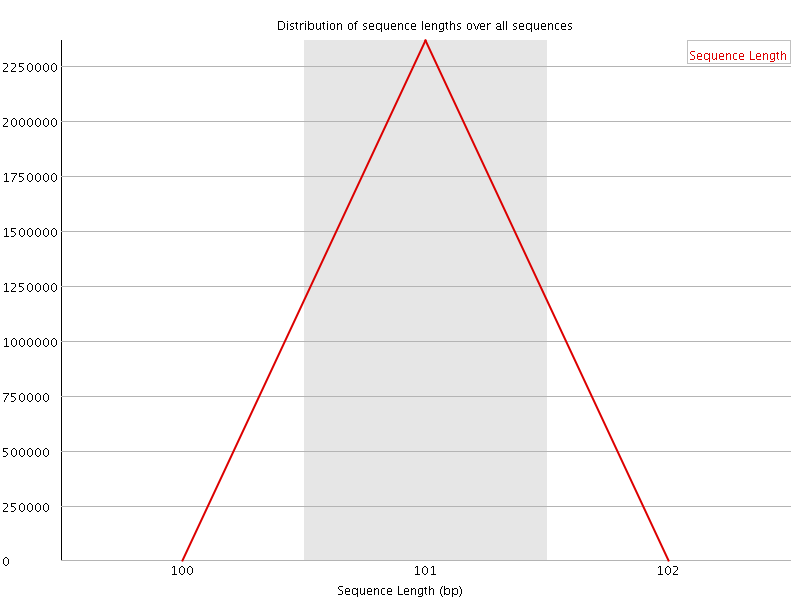
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
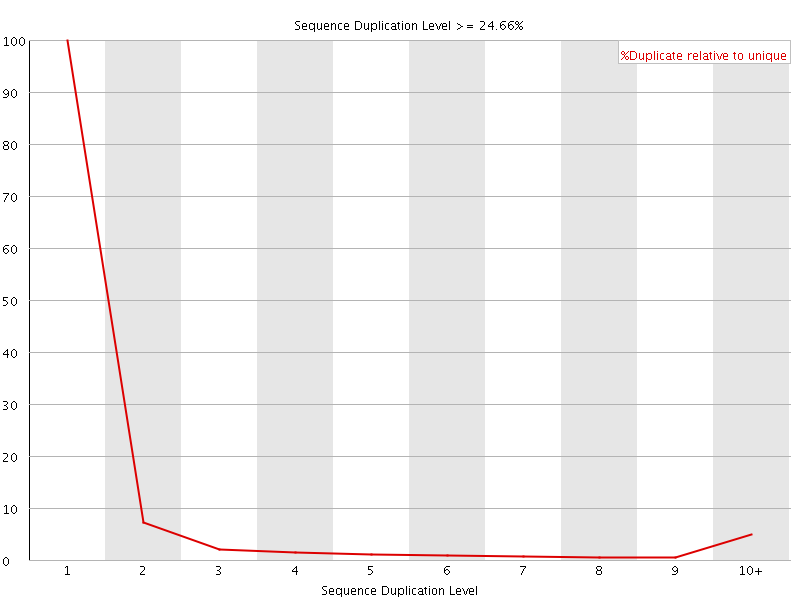
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
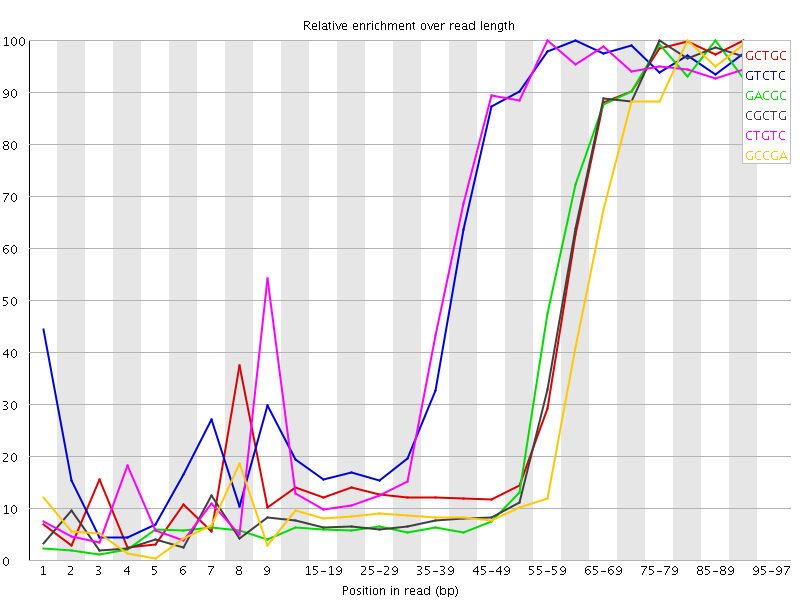
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 468390 | 4.210119 | 9.526098 | 90-94 |
| GTCTC | 710805 | 4.0458374 | 6.398896 | 60-64 |
| GACGC | 431790 | 4.0355697 | 9.647521 | 85-89 |
| CGCTG | 429215 | 3.8579946 | 9.326593 | 75-79 |
| CTGTC | 667150 | 3.7973568 | 6.1684675 | 55-59 |
| GCCGA | 402125 | 3.758316 | 9.838961 | 80-84 |
| CGACG | 394060 | 3.6829398 | 9.932279 | 95-97 |
| CTGCC | 422860 | 3.5934553 | 8.983994 | 90-94 |
| TGCCG | 389225 | 3.4985452 | 9.278487 | 90-94 |
| TCTCT | 965900 | 3.4814591 | 5.064555 | 60-64 |
| CCGAC | 385620 | 3.4073815 | 9.088585 | 80-84 |
| CTCTT | 891255 | 3.2124112 | 4.7792673 | 60-64 |
| ACACA | 768155 | 3.1125379 | 6.814395 | 6 |
| TGTCT | 812420 | 3.0972826 | 4.642667 | 65-69 |
| TGACG | 472295 | 2.9565685 | 6.6561723 | 85-89 |
| CTGAC | 492695 | 2.9159606 | 6.1195693 | 85-89 |
| GACGA | 435035 | 2.831681 | 7.27406 | 95-97 |
| TACAC | 699575 | 2.72618 | 7.848485 | 5 |
| AGAAG | 574740 | 2.6054301 | 5.268695 | 5 |
| ACGCT | 440035 | 2.6042984 | 6.211049 | 75-79 |
| CCTTG | 412965 | 2.350559 | 6.096944 | 90-94 |
| CTCCT | 431765 | 2.323455 | 5.9599705 | 90-94 |
| ACTCC | 403235 | 2.2562675 | 5.8698378 | 90-94 |
| CAAAG | 483870 | 2.0737936 | 5.143335 | 4 |
| GGTGG | 197430 | 1.9853729 | 7.5959973 | 95-97 |
| AGAGA | 432330 | 1.9598523 | 5.550975 | 8 |
| TATAC | 731015 | 1.908046 | 5.756994 | 5 |
| GAGAG | 270255 | 1.8606511 | 6.263467 | 7 |
| CTTTG | 480645 | 1.8324184 | 8.160794 | 9 |
| GTGTA | 383010 | 1.6059315 | 9.107923 | 1 |
| CTCGG | 178350 | 1.6030972 | 7.000296 | 95-97 |
| CGGTG | 142630 | 1.3560282 | 6.8494554 | 95-97 |
| AAGAC | 309690 | 1.3272848 | 5.877207 | 5 |
| GAGTC | 204975 | 1.2831442 | 6.514847 | 9 |
| CGCCG | 92350 | 1.2393138 | 5.471955 | 95-97 |
| GTCTT | 309340 | 1.1793327 | 6.080656 | 1 |
| GATTG | 281015 | 1.1782744 | 5.5611544 | 7 |
| TATGA | 416695 | 1.1504081 | 5.5897756 | 4 |
| GACTT | 289675 | 1.1483039 | 5.8901353 | 7 |
| GTGTT | 282465 | 1.1390321 | 5.6081214 | 1 |
| TCATA | 417810 | 1.0905395 | 5.337912 | 2 |
| TGGAC | 169510 | 1.0611333 | 6.99625 | 5 |
| GTCCA | 166735 | 0.9868026 | 7.2548275 | 1 |
| GGACT | 154315 | 0.9660125 | 6.5752983 | 6 |
| AGACT | 229255 | 0.9449529 | 5.0570154 | 6 |
| GAGTA | 208655 | 0.9096855 | 5.111607 | 1 |
| GTATA | 254505 | 0.70263535 | 7.2117014 | 1 |