![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-175_CGAGGCTG-AAGGAGTA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2365119 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
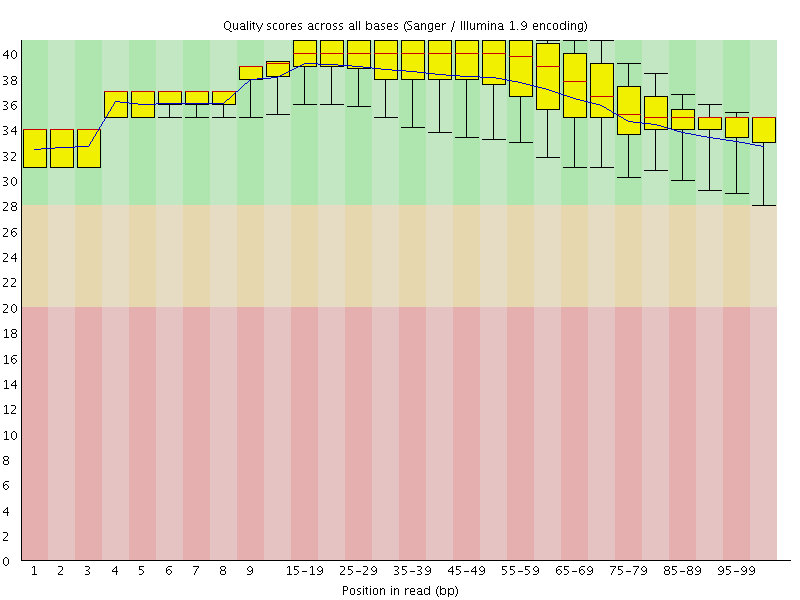
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
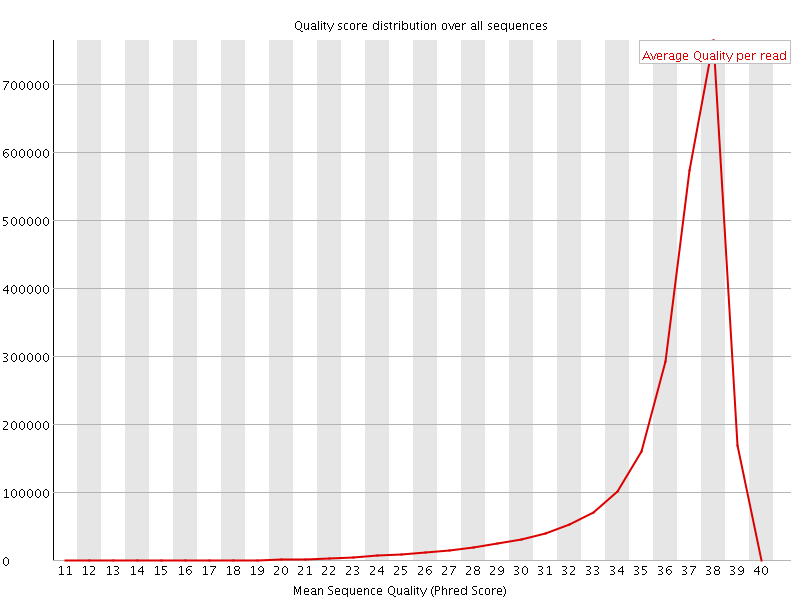
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
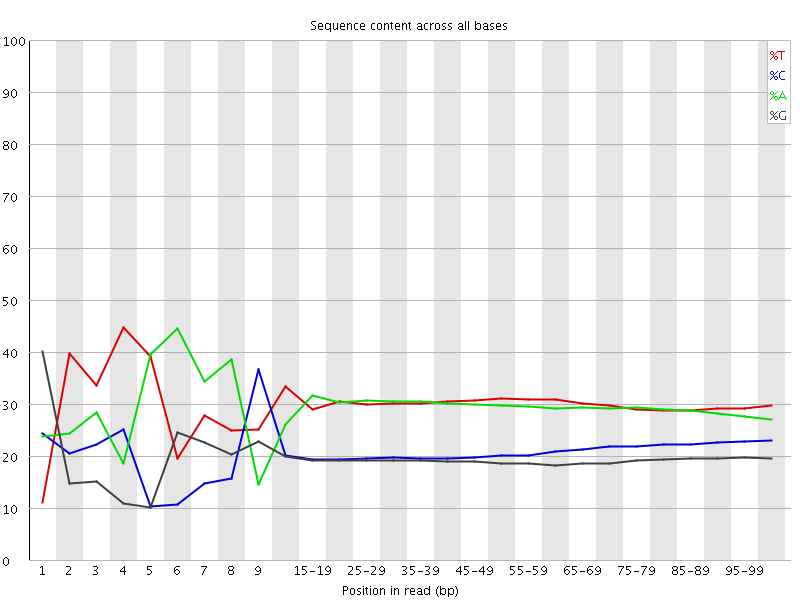
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
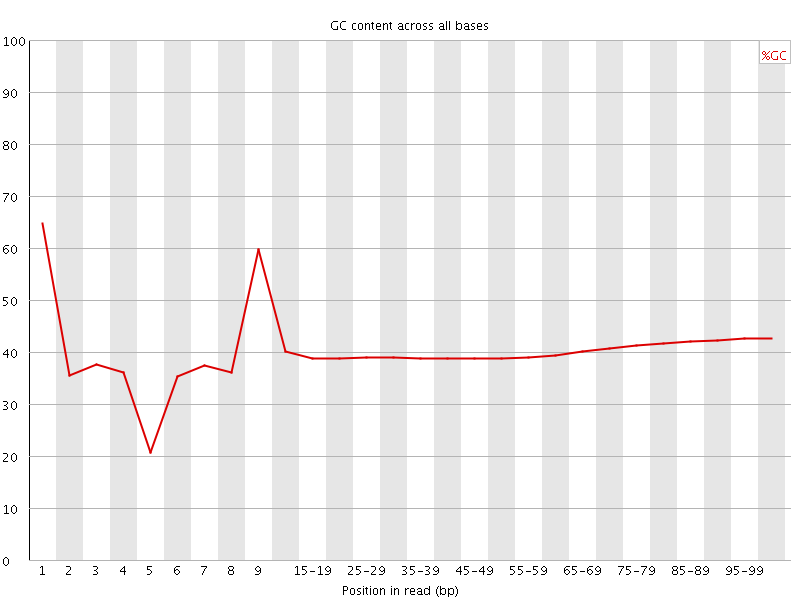
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
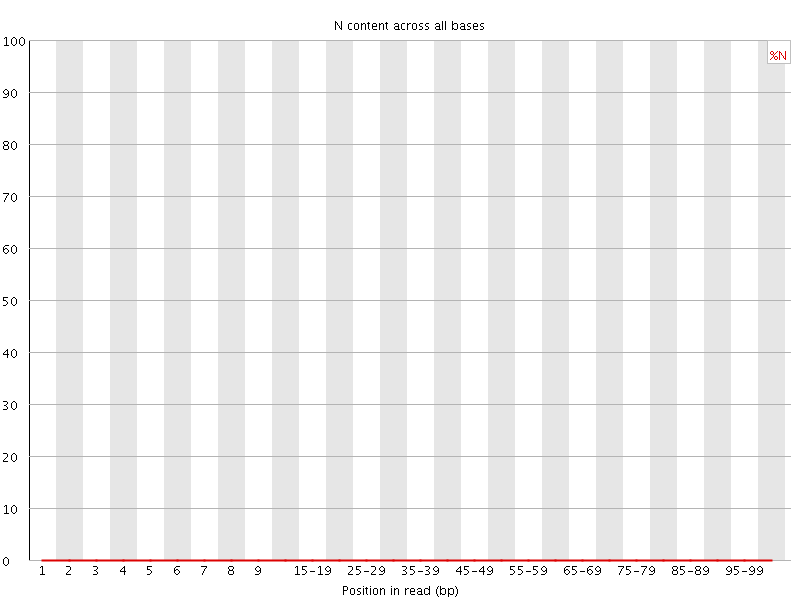
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
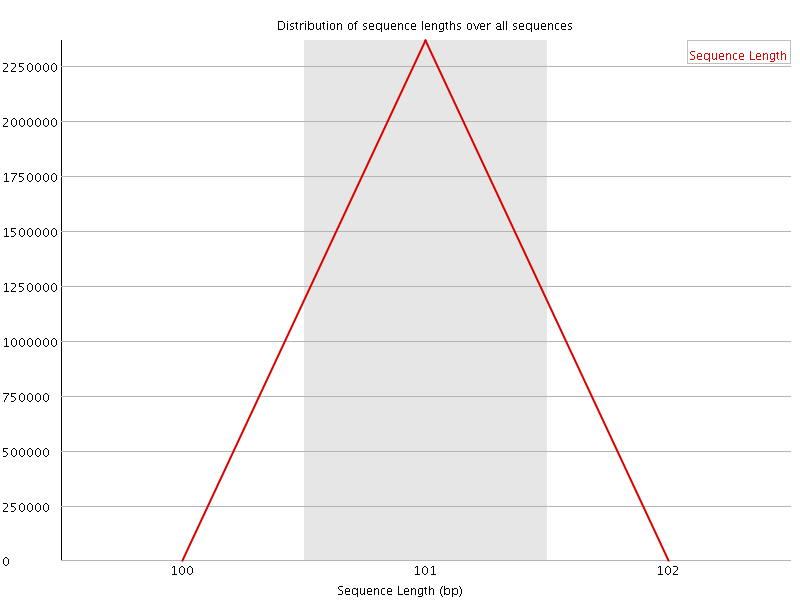
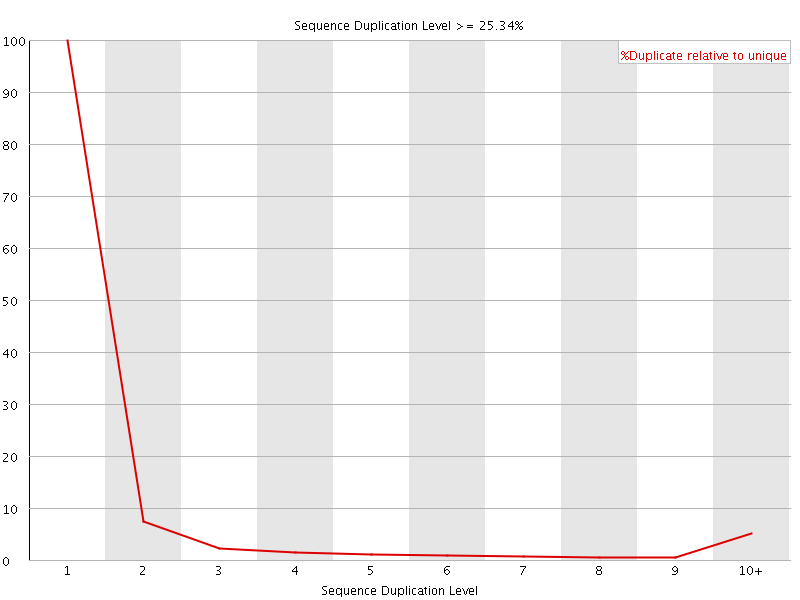
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
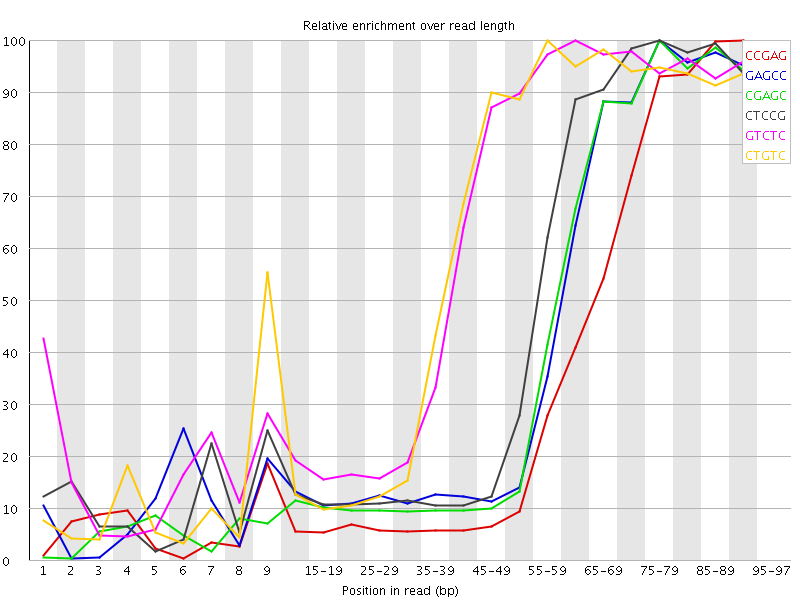
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 723630 | 6.523958 | 17.805094 | 90-94 |
| GAGCC | 469705 | 4.2346725 | 9.671886 | 75-79 |
| CGAGC | 454915 | 4.1013317 | 9.554736 | 75-79 |
| CTCCG | 500320 | 4.090456 | 8.53853 | 75-79 |
| GTCTC | 711040 | 4.0245237 | 6.407486 | 60-64 |
| CTGTC | 668510 | 3.7838016 | 6.17206 | 55-59 |
| AGCCC | 442075 | 3.6911583 | 8.582165 | 90-94 |
| GCCCA | 437185 | 3.6503286 | 8.838035 | 90-94 |
| TCTCT | 968030 | 3.5129972 | 5.1109076 | 60-64 |
| TCTCC | 647250 | 3.392849 | 6.2306566 | 70-74 |
| CCACG | 394945 | 3.2976408 | 8.66359 | 80-84 |
| CCCAC | 424515 | 3.2827046 | 8.167588 | 80-84 |
| CGAGG | 336445 | 3.2751915 | 10.316409 | 90-94 |
| CTCTT | 891815 | 3.236412 | 4.8322334 | 60-64 |
| GGCTG | 336150 | 3.2041502 | 10.577597 | 95-97 |
| TGTCT | 814850 | 3.1929686 | 4.766993 | 65-69 |
| ATCTC | 860370 | 3.1887255 | 7.4936376 | 95-97 |
| GAGGC | 327005 | 3.1832955 | 10.321129 | 90-94 |
| TCCGA | 527215 | 3.047551 | 6.36643 | 85-89 |
| GACCG | 332035 | 2.9934947 | 9.236175 | 85-89 |
| ACACA | 772735 | 2.9870887 | 6.4548182 | 6 |
| GAGAC | 454000 | 2.893942 | 7.244254 | 85-89 |
| CGAGA | 447145 | 2.8502464 | 7.2096047 | 95-97 |
| ACGAG | 426605 | 2.719318 | 6.9458637 | 95-97 |
| TACAC | 704010 | 2.6647317 | 8.118823 | 5 |
| AGAAG | 577925 | 2.6046236 | 5.5218167 | 5 |
| CACGA | 412765 | 2.4367385 | 6.280007 | 80-84 |
| AGACC | 401200 | 2.3684654 | 6.423446 | 85-89 |
| AGGCT | 369045 | 2.303406 | 6.9395375 | 90-94 |
| GCTGA | 338545 | 2.1130395 | 6.7598066 | 95-97 |
| ACCGA | 355630 | 2.0994449 | 6.1067986 | 90-94 |
| AGAGA | 430895 | 1.9419806 | 5.515261 | 8 |
| TATAC | 735040 | 1.9261149 | 5.930692 | 5 |
| CTTTG | 481625 | 1.8872352 | 8.48285 | 9 |
| GAGAG | 268710 | 1.8494635 | 6.5139594 | 7 |
| TGGAG | 250665 | 1.6893235 | 5.005889 | 5 |
| TCTCG | 295075 | 1.6701397 | 5.390104 | 90-94 |
| GCTTT | 400900 | 1.5709163 | 5.0866733 | 1 |
| CTCGT | 254715 | 1.4417002 | 5.3039284 | 90-94 |
| AGGAG | 208425 | 1.4345369 | 5.2125025 | 7 |
| GTCTT | 352020 | 1.3793813 | 5.9721384 | 1 |
| AAGAC | 311445 | 1.2999506 | 5.775675 | 5 |
| GAGTC | 204690 | 1.2775792 | 6.5577283 | 9 |
| GCCGT | 141915 | 1.2527955 | 7.917171 | 95-97 |
| ATGAG | 279325 | 1.2326517 | 5.034494 | 7 |
| TGCCG | 137760 | 1.216116 | 8.068409 | 95-97 |
| GATTG | 277360 | 1.1984823 | 5.507845 | 7 |
| CCGTC | 146185 | 1.1951617 | 6.809065 | 95-97 |
| ATGCC | 206560 | 1.1940142 | 5.505582 | 95-97 |
| GTGTT | 280180 | 1.1854467 | 5.9191484 | 1 |
| TATGA | 413810 | 1.170847 | 5.4434485 | 4 |
| GACTT | 286505 | 1.1465474 | 6.127386 | 7 |
| TCATA | 423415 | 1.109526 | 5.4893827 | 2 |
| TGGAC | 167745 | 1.0469857 | 7.0842834 | 5 |
| GTGTA | 240005 | 1.0370699 | 9.114765 | 1 |
| GTCCA | 169230 | 0.97822917 | 6.909533 | 1 |
| GGACT | 152150 | 0.94964904 | 6.5547023 | 6 |
| AGACT | 230750 | 0.943071 | 5.387846 | 6 |
| GAGTA | 207495 | 0.91566837 | 5.178625 | 1 |
| GTATA | 255345 | 0.7224812 | 7.0495567 | 1 |