![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-174_CGAGGCTG-ACTGCATA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 3154287 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
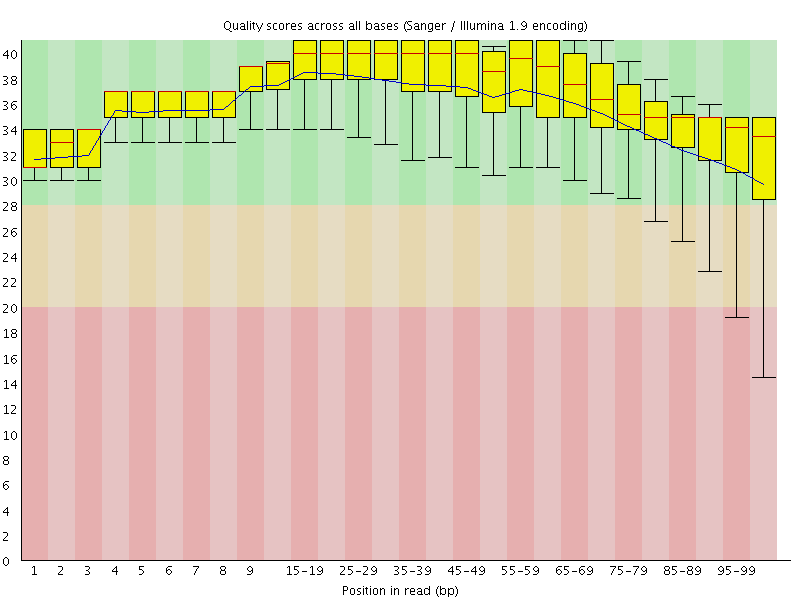
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
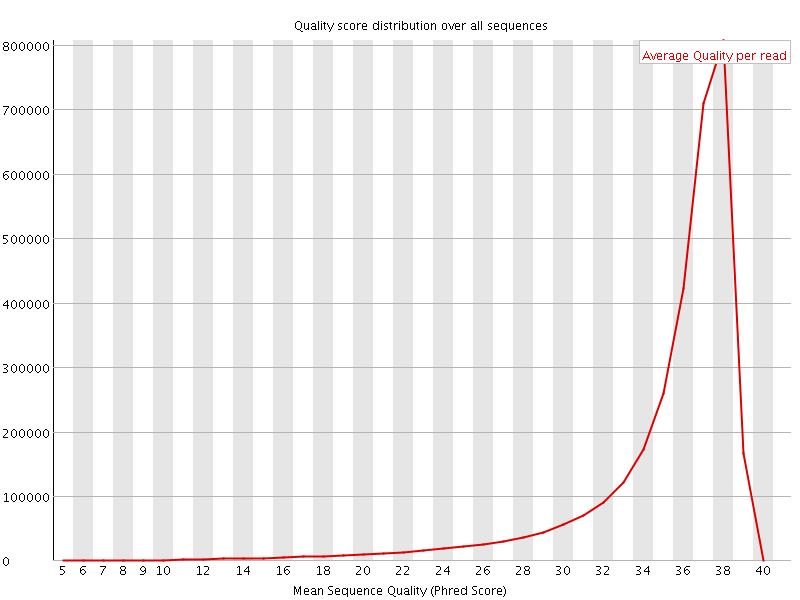
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
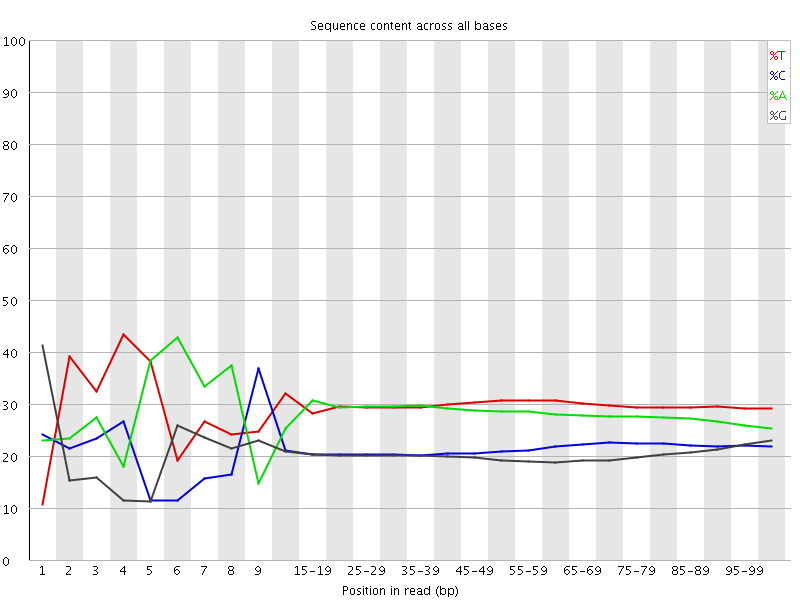
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
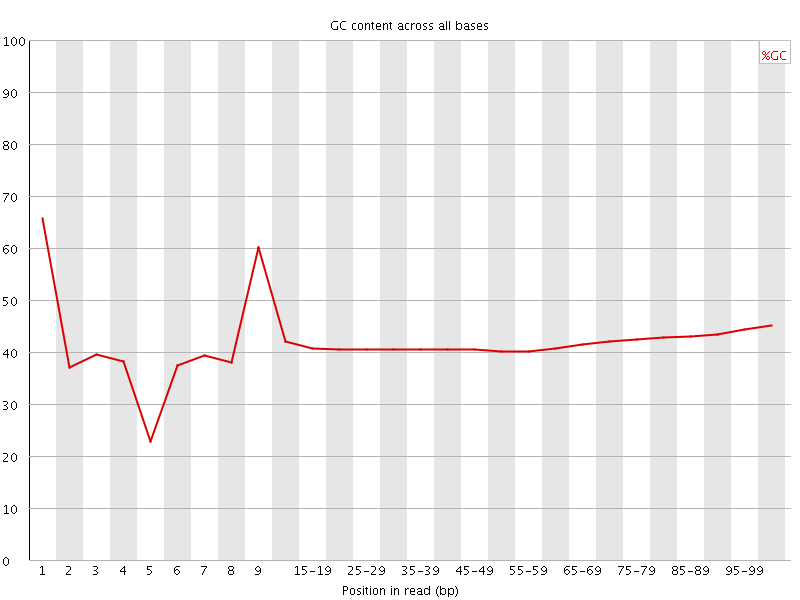
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
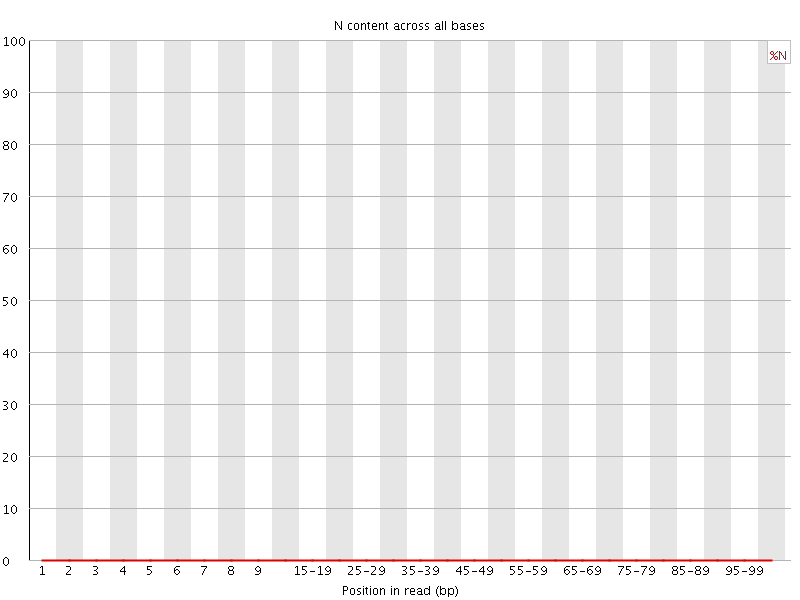
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
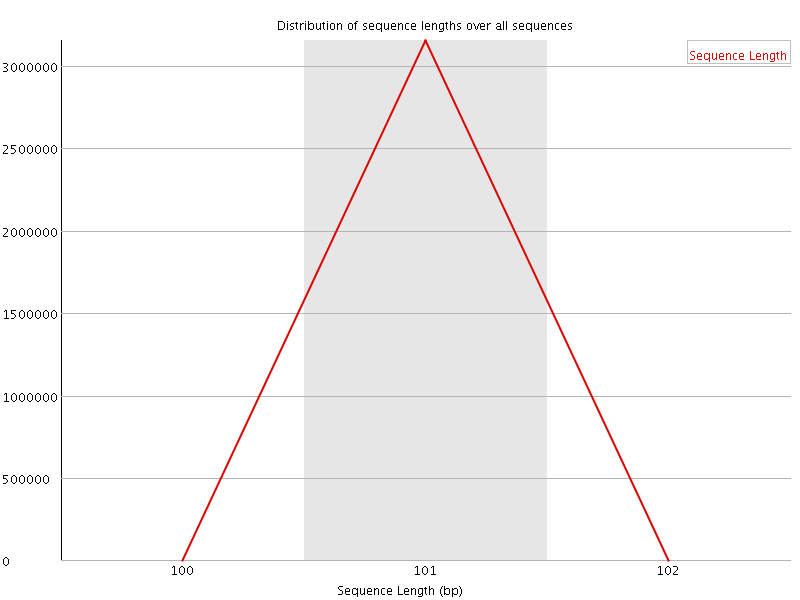
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
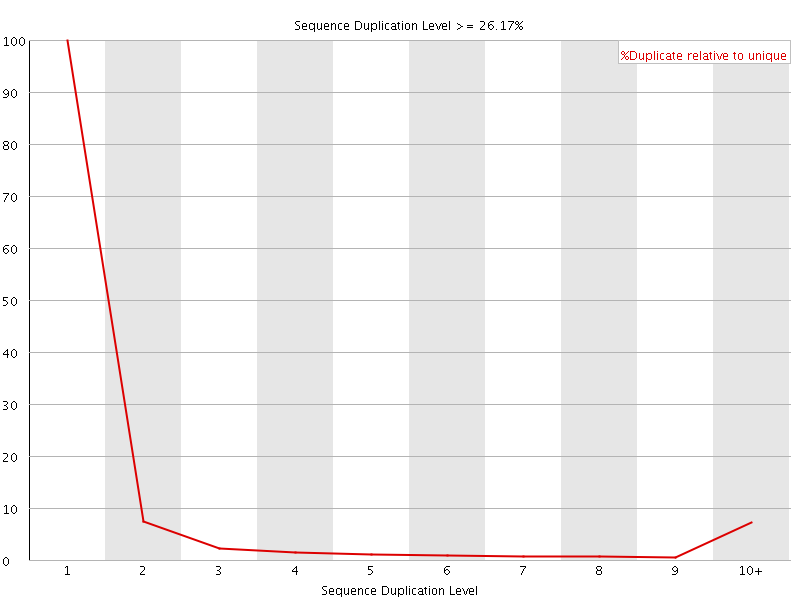
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
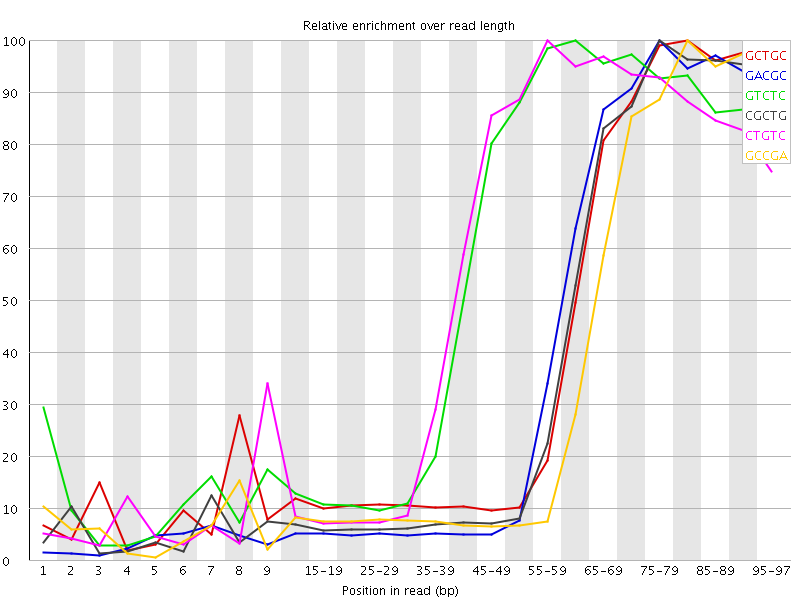
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 943910 | 5.447707 | 13.233231 | 80-84 |
| GACGC | 894665 | 5.432268 | 13.63038 | 75-79 |
| GTCTC | 1344155 | 5.2795544 | 9.205545 | 60-64 |
| CGCTG | 899825 | 5.193274 | 13.259406 | 75-79 |
| CTGTC | 1310650 | 5.147954 | 9.105021 | 55-59 |
| GCCGA | 818000 | 4.96677 | 13.830164 | 80-84 |
| CGACG | 788360 | 4.786801 | 13.6593685 | 85-89 |
| CTGCC | 860130 | 4.707976 | 12.487071 | 80-84 |
| TGCCG | 813670 | 4.696037 | 13.120686 | 80-84 |
| CCGAC | 772270 | 4.4471006 | 12.885102 | 80-84 |
| TCTCT | 1646710 | 4.401795 | 7.2076716 | 60-64 |
| CTCTT | 1554025 | 4.1540403 | 6.987018 | 60-64 |
| TGACG | 947080 | 4.1265326 | 9.849523 | 75-79 |
| TGTCT | 1463440 | 4.1247787 | 6.9651 | 55-59 |
| ACACA | 1304700 | 4.06103 | 7.605423 | 65-69 |
| CTGAC | 977260 | 4.038273 | 9.235772 | 75-79 |
| ATCTG | 1260990 | 3.7391684 | 7.697618 | 70-74 |
| GACGA | 813955 | 3.7310975 | 10.635757 | 85-89 |
| ACGCT | 883145 | 3.649367 | 9.394097 | 75-79 |
| TACAC | 1229360 | 3.6371984 | 7.020193 | 65-69 |
| CACAT | 1225690 | 3.6263404 | 6.9861 | 70-74 |
| ACATC | 1199625 | 3.5492241 | 7.129312 | 70-74 |
| CATCT | 1251270 | 3.5188549 | 6.983625 | 70-74 |
| TCTGA | 1159590 | 3.4384902 | 7.1346993 | 70-74 |
| ATACA | 1373170 | 3.067098 | 5.4978933 | 65-69 |
| TCTTA | 1493120 | 3.0131645 | 5.1158686 | 60-64 |
| TGCAG | 677960 | 2.9539468 | 9.8970175 | 90-94 |
| TATAC | 1307645 | 2.776231 | 5.640547 | 5 |
| GCAGT | 562130 | 2.4492626 | 9.377611 | 90-94 |
| ACGAT | 769760 | 2.4013546 | 7.151703 | 85-89 |
| ATGCA | 765530 | 2.3881586 | 7.23315 | 90-94 |
| CAGTG | 543135 | 2.3664994 | 9.263006 | 95-97 |
| GTGTG | 535750 | 2.3395708 | 9.496339 | 95-97 |
| AGAAG | 656980 | 2.273546 | 5.1971498 | 5 |
| CGATA | 701915 | 2.1897044 | 7.0212865 | 85-89 |
| ATATG | 973155 | 2.1785161 | 5.639059 | 90-94 |
| TATGC | 683735 | 2.0274546 | 6.701836 | 90-94 |
| GATAT | 874835 | 1.9584159 | 5.421449 | 85-89 |
| GGTGG | 301890 | 1.9371264 | 9.171455 | 95-97 |
| TGTGT | 650095 | 1.9320377 | 6.2619667 | 95-97 |
| AGTGT | 612855 | 1.9161702 | 6.700643 | 95-97 |
| AGAGA | 528355 | 1.8284262 | 5.380481 | 8 |
| GTGTA | 541265 | 1.6923348 | 7.4273925 | 1 |
| CTCGG | 289505 | 1.6708567 | 9.0460415 | 95-97 |
| GAGAG | 345145 | 1.6682104 | 6.0637307 | 7 |
| AGATC | 524845 | 1.6373142 | 5.3364854 | 95-97 |
| GATCT | 536245 | 1.590108 | 5.2292104 | 95-97 |
| TGTAG | 506445 | 1.5834658 | 5.853965 | 95-97 |
| CGCCG | 195685 | 1.5738486 | 5.138661 | 95-97 |
| CTTTG | 532260 | 1.5002013 | 6.3409047 | 9 |
| GTAGA | 454870 | 1.4962401 | 5.813548 | 95-97 |
| CGGTG | 240660 | 1.4645362 | 8.585959 | 95-97 |
| TCTCG | 350480 | 1.3766108 | 6.2065163 | 95-97 |
| AAGAC | 351655 | 1.1541317 | 5.500464 | 5 |
| TCGGT | 252595 | 1.0461302 | 5.8392205 | 95-97 |
| GTCTT | 351705 | 0.9912981 | 5.370653 | 1 |
| GTGTT | 329255 | 0.97852325 | 5.1741867 | 1 |
| TGGAC | 207815 | 0.905473 | 5.7213635 | 5 |
| GTCCA | 204825 | 0.8463861 | 5.953494 | 1 |
| GGACT | 182705 | 0.79606587 | 5.030904 | 6 |
| GTATA | 295910 | 0.66242766 | 6.9218273 | 1 |