![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-174_CGAGGCTG-ACTGCATA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 3154287 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 42 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
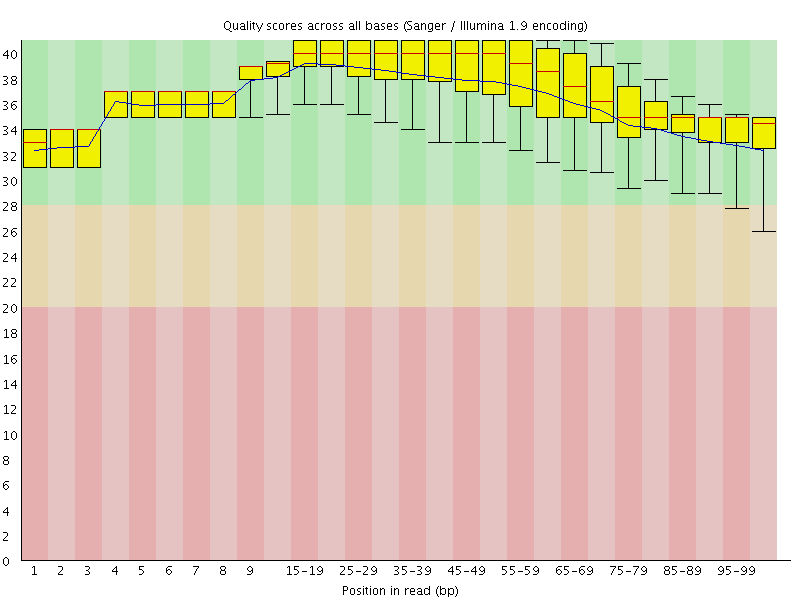
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
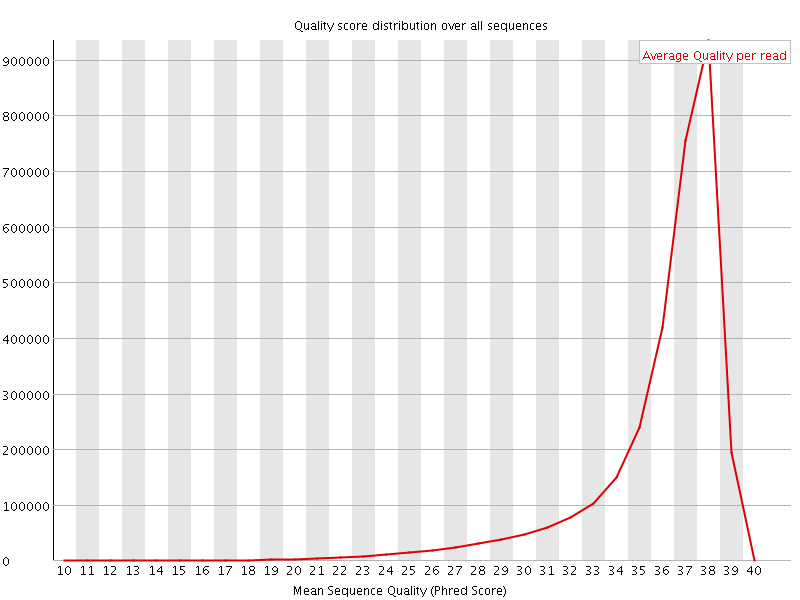
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
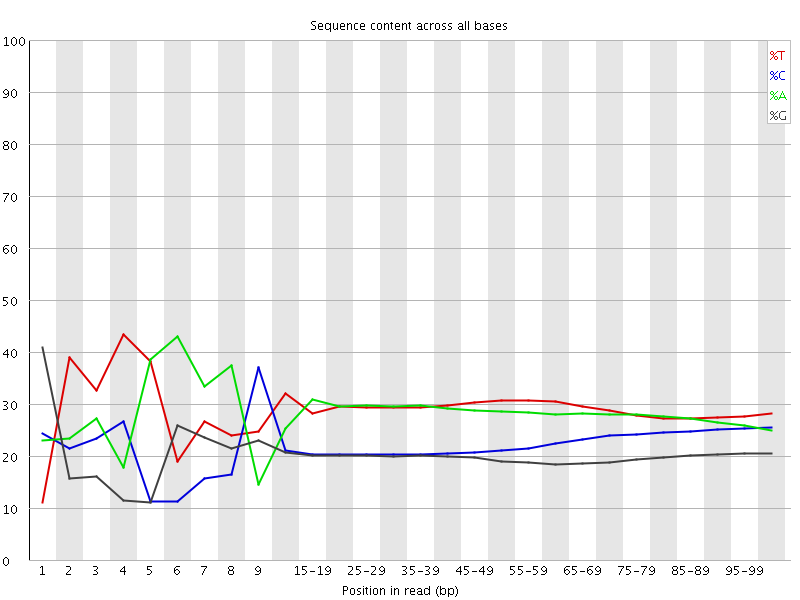
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
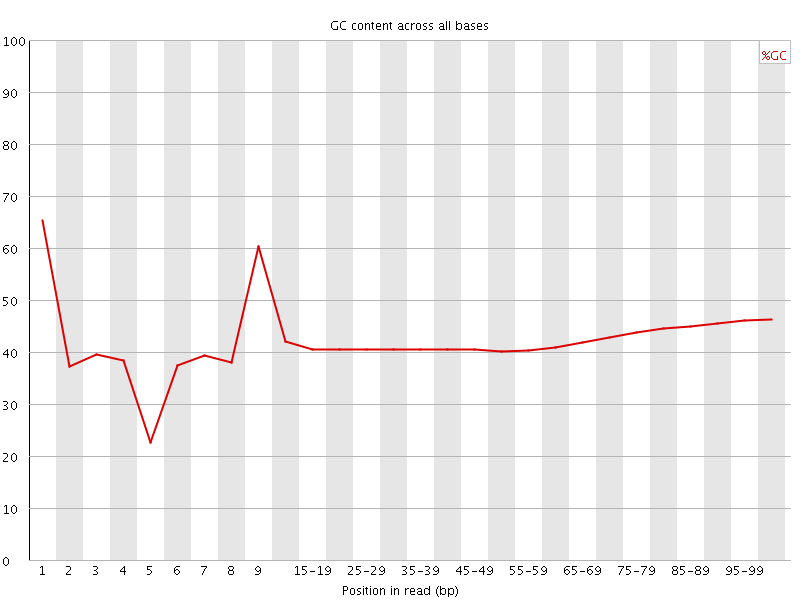
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
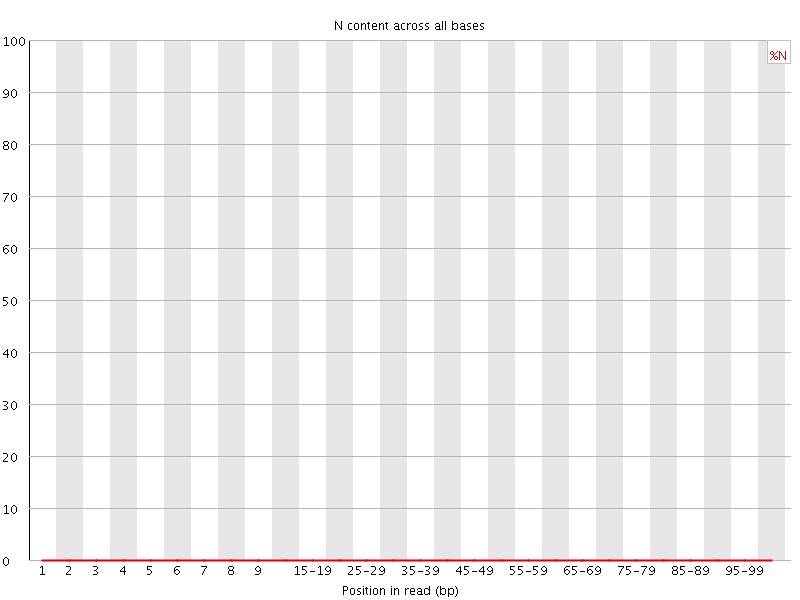
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
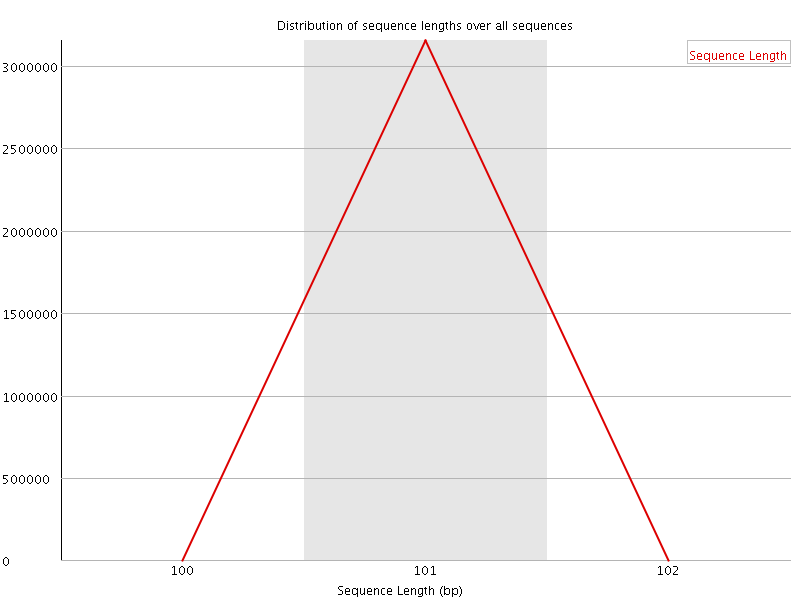
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
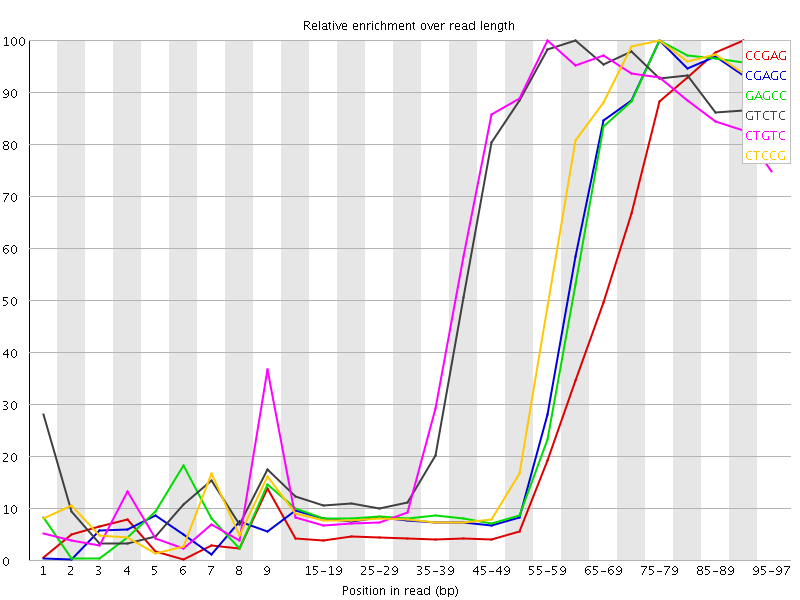
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 1457460 | 8.43904 | 24.970184 | 90-94 |
| CGAGC | 919970 | 5.326845 | 13.2521305 | 75-79 |
| GAGCC | 919185 | 5.3223 | 13.187557 | 75-79 |
| GTCTC | 1347430 | 5.163693 | 8.999582 | 60-64 |
| CTGTC | 1316050 | 5.043437 | 8.911181 | 55-59 |
| CTCCG | 988845 | 4.9748063 | 11.272796 | 75-79 |
| AGCCC | 877190 | 4.547751 | 11.565592 | 80-84 |
| GCCCA | 866175 | 4.490644 | 11.763178 | 80-84 |
| TCTCT | 1654680 | 4.324949 | 7.0883174 | 60-64 |
| TGTCT | 1465220 | 4.2772365 | 7.1998 | 55-59 |
| CGAGG | 642615 | 4.155667 | 14.704657 | 90-94 |
| ATCTC | 1539795 | 4.1474853 | 11.260763 | 95-97 |
| CCACG | 790310 | 4.0973253 | 11.684243 | 80-84 |
| GGCTG | 649420 | 4.075308 | 14.762626 | 95-97 |
| CTCTT | 1555210 | 4.0649576 | 6.8515916 | 60-64 |
| TCTCC | 1171875 | 4.021074 | 8.323068 | 70-74 |
| GAGGC | 617120 | 3.9907954 | 14.787439 | 90-94 |
| TCCGA | 1003710 | 3.963852 | 9.121266 | 75-79 |
| CCCAC | 833535 | 3.8693082 | 10.570307 | 80-84 |
| GACCG | 659820 | 3.820515 | 12.959202 | 85-89 |
| ACACA | 1312375 | 3.7539628 | 7.0148892 | 70-74 |
| CGAGA | 823765 | 3.7442172 | 10.595063 | 85-89 |
| GAGAC | 812050 | 3.6909692 | 10.710111 | 85-89 |
| ACGAG | 800745 | 3.6395853 | 10.274601 | 85-89 |
| TACAC | 1238280 | 3.437129 | 6.6006413 | 65-69 |
| CACAT | 1234205 | 3.425818 | 6.536895 | 70-74 |
| CATCT | 1257150 | 3.3861725 | 6.72025 | 70-74 |
| ACATC | 1207810 | 3.3525527 | 6.7167106 | 70-74 |
| CACGA | 795150 | 3.2360353 | 9.239835 | 80-84 |
| TCTTA | 1495425 | 3.0682647 | 5.221449 | 60-64 |
| AGACC | 745010 | 3.03198 | 9.416336 | 85-89 |
| ATACA | 1380350 | 3.007651 | 5.3560123 | 65-69 |
| CTTAT | 1416265 | 2.9058468 | 5.080038 | 60-64 |
| AGGCT | 648010 | 2.8581457 | 10.253368 | 90-94 |
| TATAC | 1313030 | 2.7762454 | 5.7184396 | 5 |
| ACCGA | 670275 | 2.7278295 | 9.055471 | 90-94 |
| GCTGA | 596580 | 2.6313057 | 10.099435 | 95-97 |
| AGAAG | 654840 | 2.336437 | 5.5029855 | 5 |
| GATCT | 675740 | 2.0328019 | 6.9252186 | 95-97 |
| TGATC | 673745 | 2.0268006 | 7.09102 | 95-97 |
| CTGAT | 636765 | 1.9155549 | 7.084212 | 95-97 |
| TCTCG | 488675 | 1.8727262 | 7.6917014 | 95-97 |
| AGAGA | 523390 | 1.8674299 | 5.3524795 | 8 |
| GAGAG | 341100 | 1.731541 | 6.2172775 | 7 |
| CTCGT | 426445 | 1.6342452 | 7.392081 | 90-94 |
| AAGAG | 446110 | 1.5916986 | 5.0410876 | 5 |
| CTTTG | 532435 | 1.554272 | 6.605627 | 9 |
| GTATG | 443350 | 1.4895517 | 6.7944164 | 95-97 |
| TGCCG | 258280 | 1.4512163 | 10.646253 | 95-97 |
| GCCGT | 243265 | 1.3668505 | 9.926088 | 95-97 |
| ATGCC | 339125 | 1.3392726 | 7.7164245 | 95-97 |
| GTCTT | 421715 | 1.2310607 | 5.3637557 | 1 |
| CCGTC | 238580 | 1.2002784 | 8.022425 | 95-97 |
| TCGTA | 390740 | 1.1754477 | 5.926525 | 95-97 |
| TATGC | 379725 | 1.1423116 | 5.950835 | 95-97 |
| AAGAC | 353500 | 1.1293145 | 5.1843824 | 5 |
| CGTAT | 355555 | 1.069602 | 5.9357624 | 95-97 |
| GTGTT | 325255 | 1.0604198 | 5.709069 | 1 |
| TGGAC | 206455 | 0.91060084 | 5.8916926 | 5 |
| CGTCT | 237320 | 0.90947026 | 5.494692 | 95-97 |
| GTGTA | 261135 | 0.8773522 | 8.231069 | 1 |
| GTCCA | 207760 | 0.82048583 | 5.6399913 | 1 |
| GGACT | 182220 | 0.8037087 | 5.143201 | 6 |
| GTATA | 293820 | 0.6938379 | 7.150288 | 1 |