![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-173_CGAGGCTG-GTAAGGAG_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2438877 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
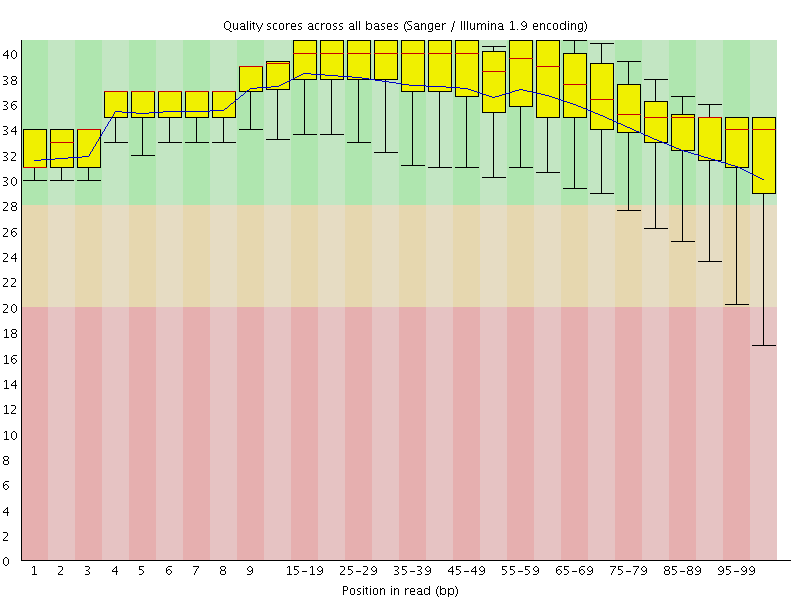
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
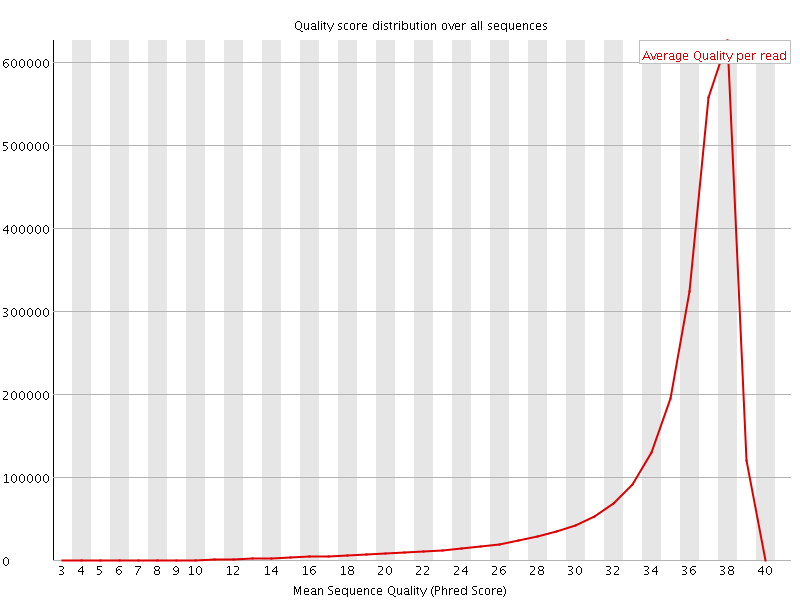
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
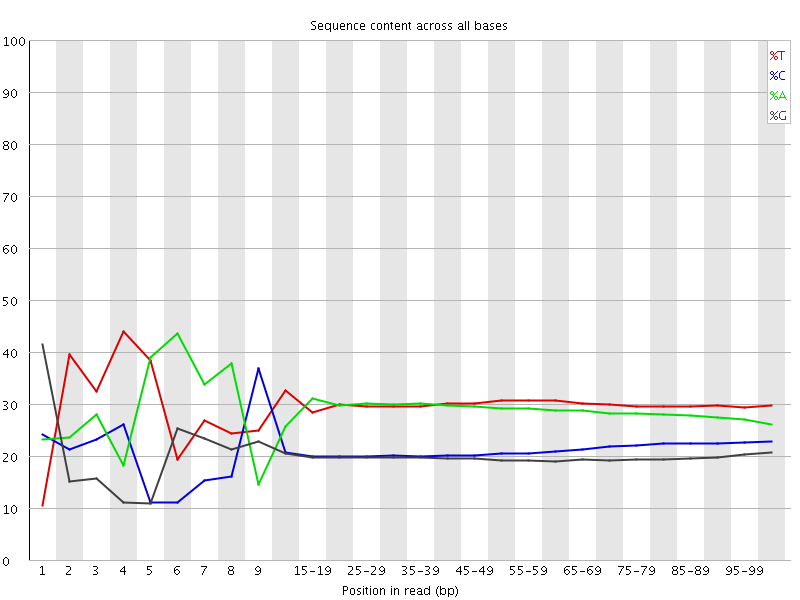
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
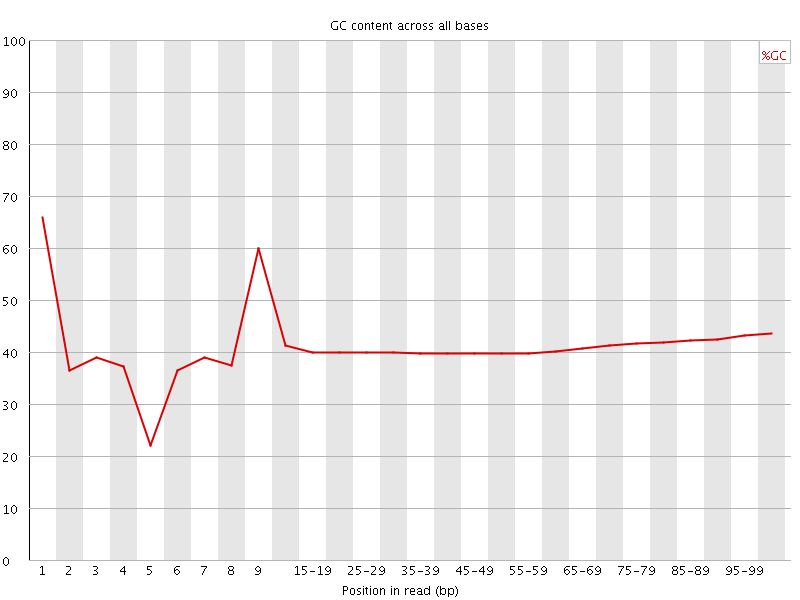
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
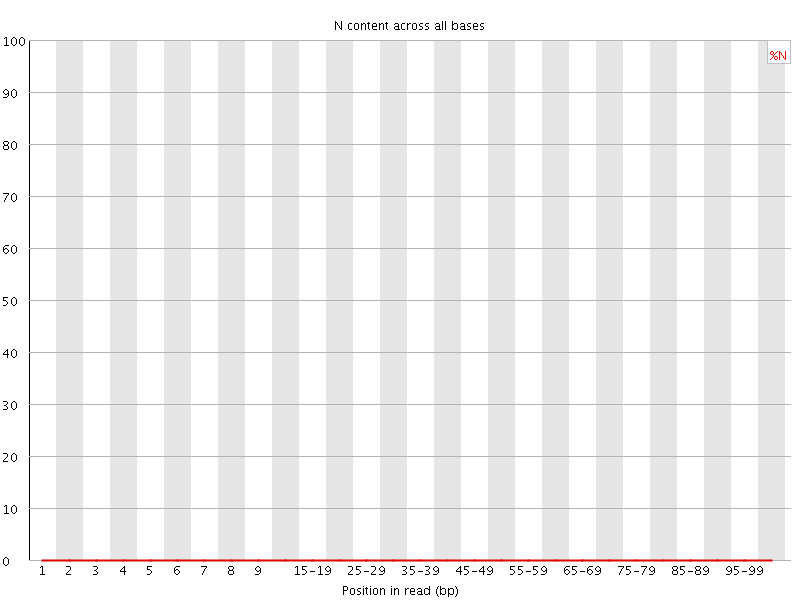
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
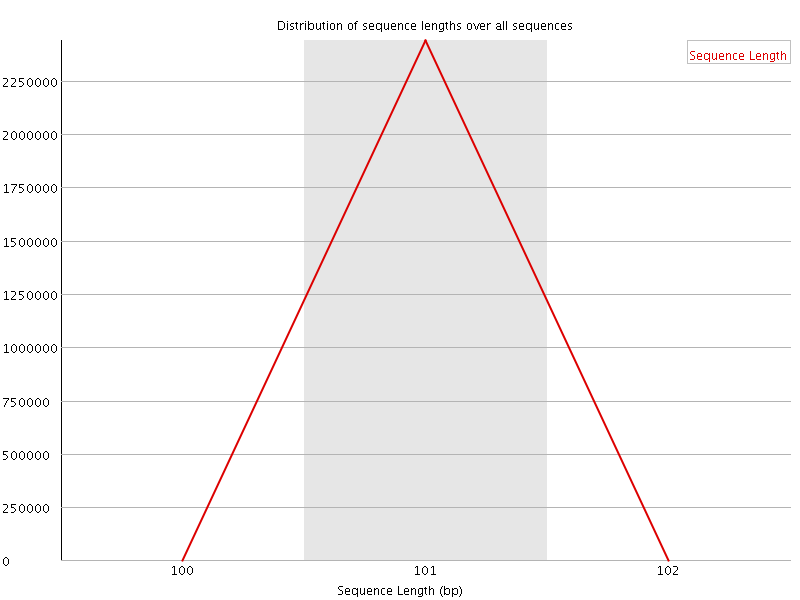
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
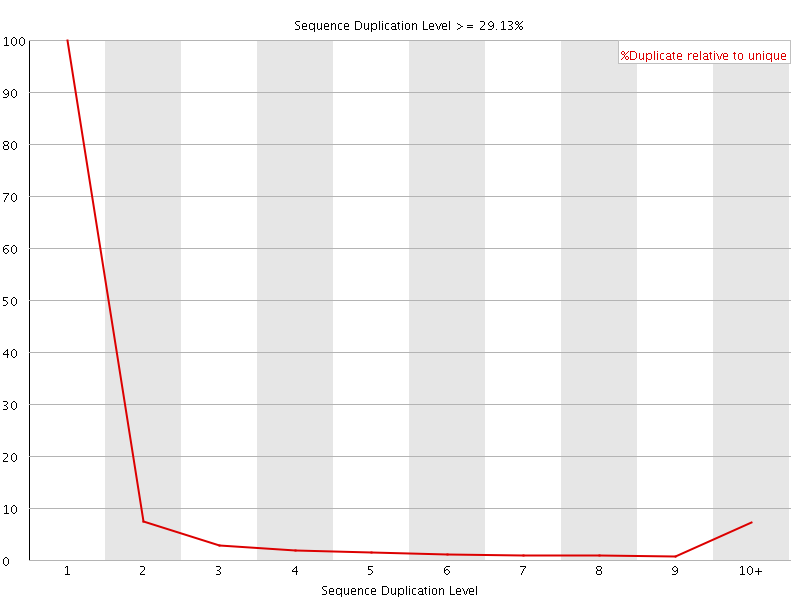
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
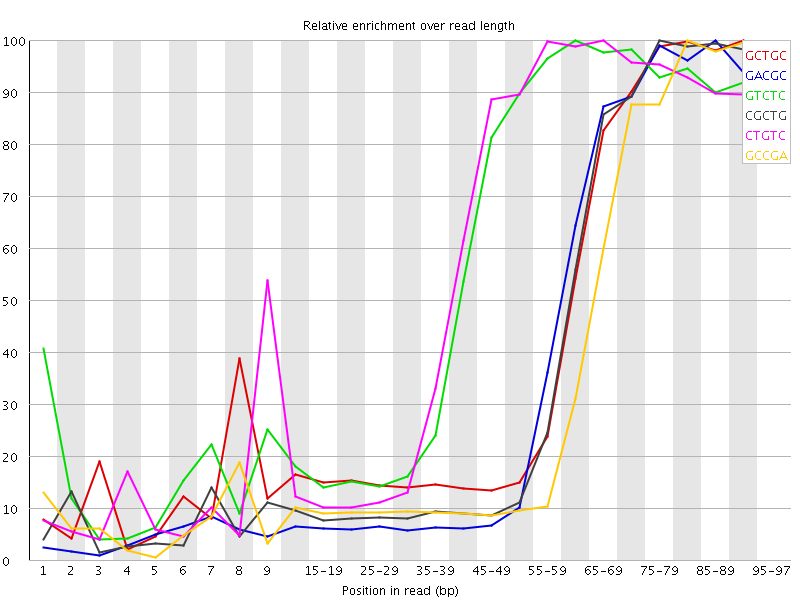
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 531550 | 4.237816 | 9.568855 | 90-94 |
| GACGC | 474545 | 3.930563 | 9.602187 | 85-89 |
| GTCTC | 745590 | 3.9270155 | 6.5098886 | 60-64 |
| CGCTG | 479765 | 3.8249574 | 9.262894 | 75-79 |
| CTGTC | 714675 | 3.7641866 | 6.257766 | 65-69 |
| GCCGA | 439560 | 3.640789 | 9.656538 | 80-84 |
| CTGCC | 474850 | 3.554038 | 8.95462 | 80-84 |
| CGACG | 424485 | 3.5159252 | 9.741897 | 95-97 |
| TCTCT | 992405 | 3.4531548 | 5.311902 | 60-64 |
| TGCCG | 431500 | 3.4401615 | 9.242522 | 90-94 |
| CTCTT | 928300 | 3.230096 | 4.985724 | 70-74 |
| CCGAC | 414155 | 3.2203858 | 8.871525 | 95-97 |
| TGTCT | 841880 | 3.1203961 | 4.850035 | 60-64 |
| ACACA | 789460 | 3.0803158 | 6.3934817 | 6 |
| TGACG | 510400 | 2.9749863 | 6.852186 | 85-89 |
| CTGAC | 533305 | 2.9182167 | 6.345707 | 85-89 |
| GACGA | 455885 | 2.7606342 | 7.3239684 | 95-97 |
| ATCTG | 713830 | 2.7487397 | 5.2432957 | 70-74 |
| TACAC | 728880 | 2.7374222 | 7.471295 | 5 |
| ACGCT | 472975 | 2.5880942 | 6.2767096 | 85-89 |
| GACTC | 472530 | 2.585659 | 6.9900436 | 85-89 |
| AGAAG | 577255 | 2.5556276 | 5.3904366 | 5 |
| CTCCT | 466555 | 2.3069227 | 6.002713 | 90-94 |
| ACTCC | 436805 | 2.243867 | 5.820745 | 90-94 |
| ACGAC | 386400 | 2.1966374 | 6.4829097 | 85-89 |
| CGACT | 395740 | 2.1654682 | 6.0670605 | 85-89 |
| TATAC | 761125 | 2.0115862 | 5.552534 | 5 |
| CAAAG | 483765 | 2.0106292 | 5.090721 | 4 |
| GGTGG | 208235 | 1.8837203 | 6.7538257 | 95-97 |
| AGAGA | 411570 | 1.8221056 | 5.062675 | 8 |
| CTTTG | 483570 | 1.7923335 | 7.75611 | 9 |
| GAGAG | 265780 | 1.7143847 | 5.8777018 | 7 |
| GTGTA | 369245 | 1.5145577 | 8.900247 | 1 |
| ACGTG | 252835 | 1.4737083 | 6.001762 | 95-97 |
| CTCGG | 180570 | 1.439606 | 6.5026803 | 95-97 |
| CGTGT | 242015 | 1.3578044 | 5.607343 | 95-97 |
| CGGTG | 151025 | 1.2825644 | 6.0493402 | 95-97 |
| AAGAC | 304570 | 1.265857 | 5.6814456 | 5 |
| GAGTC | 207355 | 1.2086172 | 6.1452208 | 9 |
| TATGA | 417745 | 1.1760516 | 5.528101 | 4 |
| GATTG | 277815 | 1.1395328 | 5.4940124 | 7 |
| GTCTT | 302460 | 1.1210563 | 6.0071416 | 1 |
| GTGTT | 282225 | 1.114262 | 5.7993546 | 1 |
| TCATA | 419295 | 1.1081598 | 5.320506 | 2 |
| GACTT | 285900 | 1.1009129 | 5.3351164 | 7 |
| TGGAC | 174055 | 1.0145204 | 6.6162286 | 5 |
| GTCCA | 172510 | 0.9439657 | 6.811308 | 1 |
| GGACT | 156450 | 0.9119055 | 5.921645 | 6 |
| GTATA | 248560 | 0.69975555 | 7.0892143 | 1 |