![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-171_CGAGGCTG-TATCCTCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1270236 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 39 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
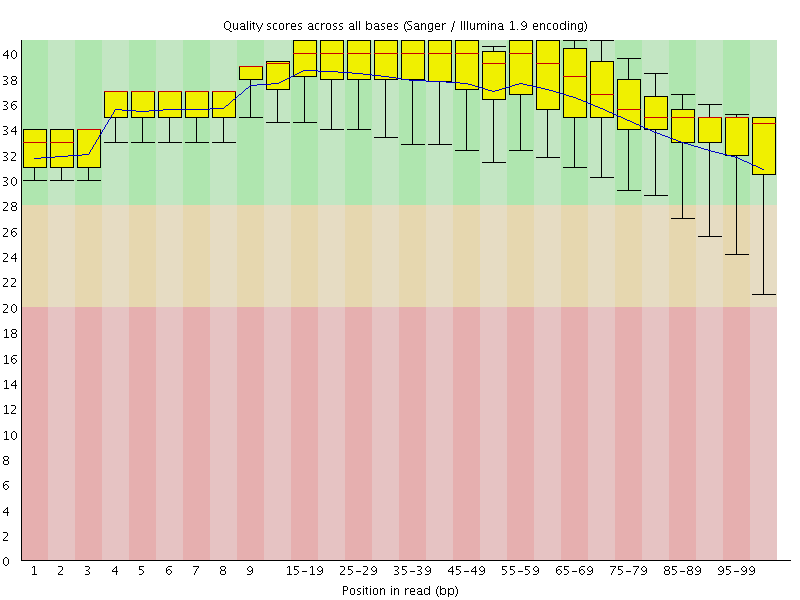
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
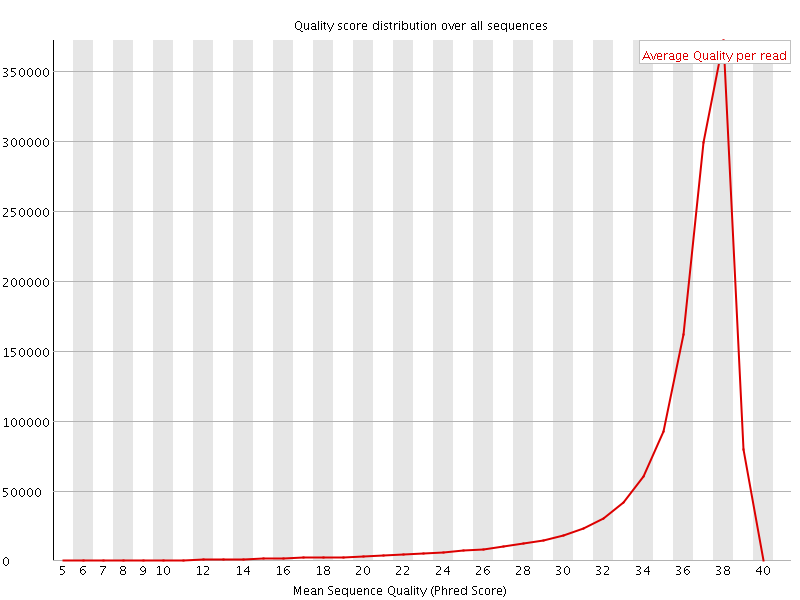
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
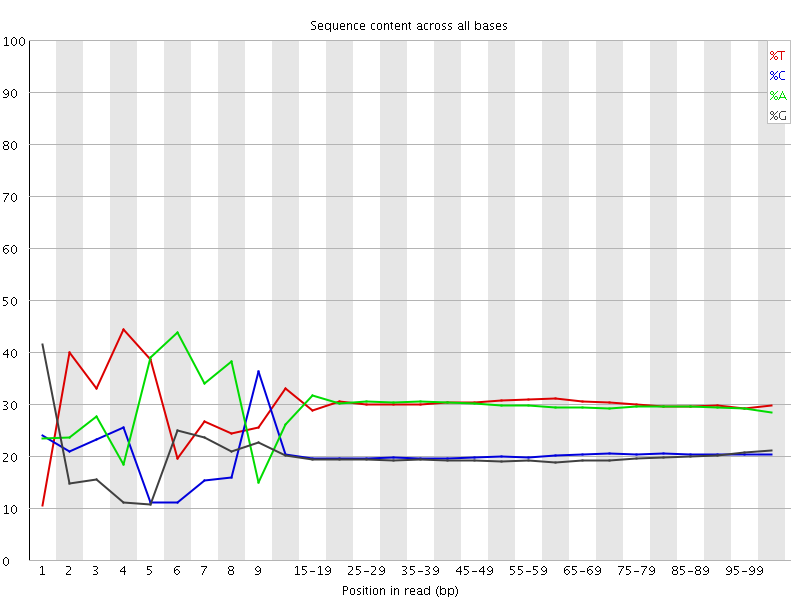
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
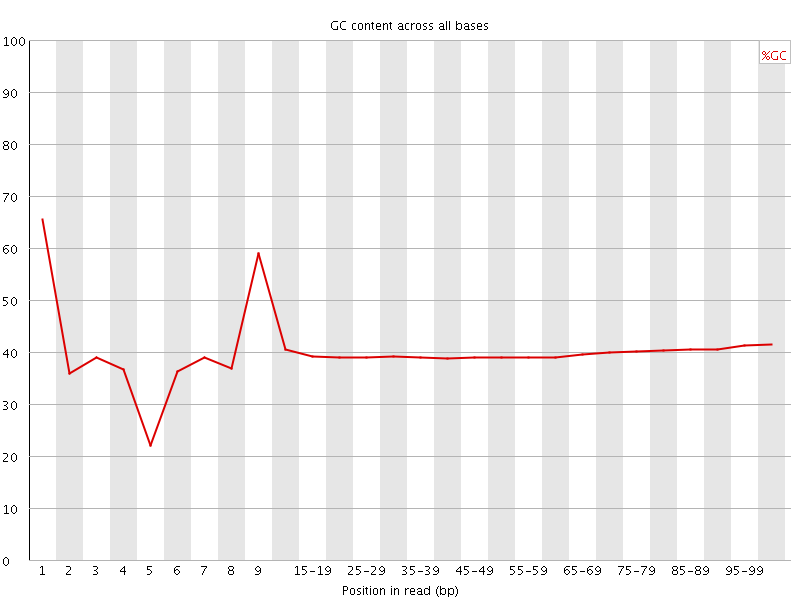
![[OK]](Icons/tick.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
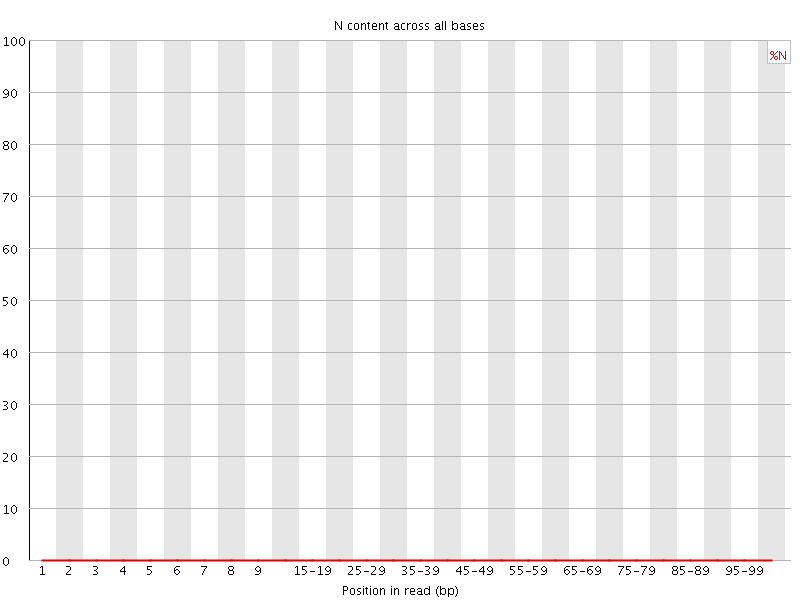
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
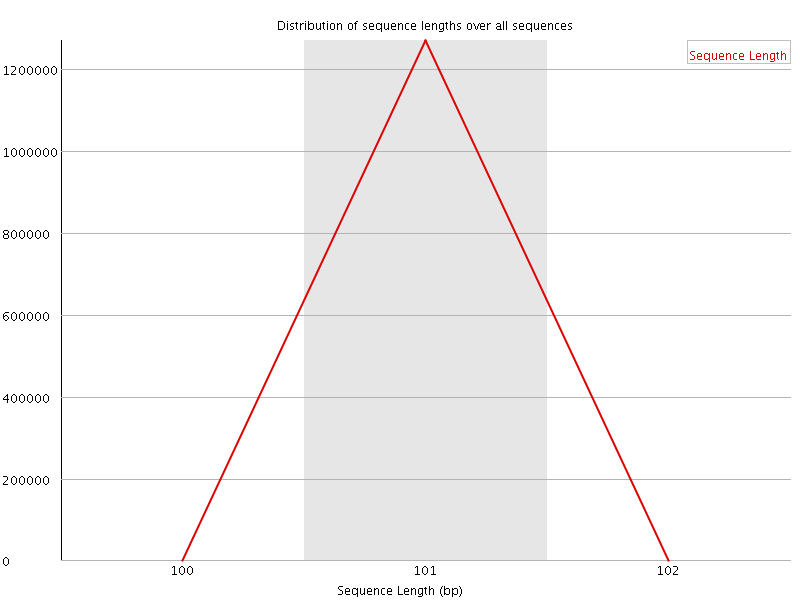
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
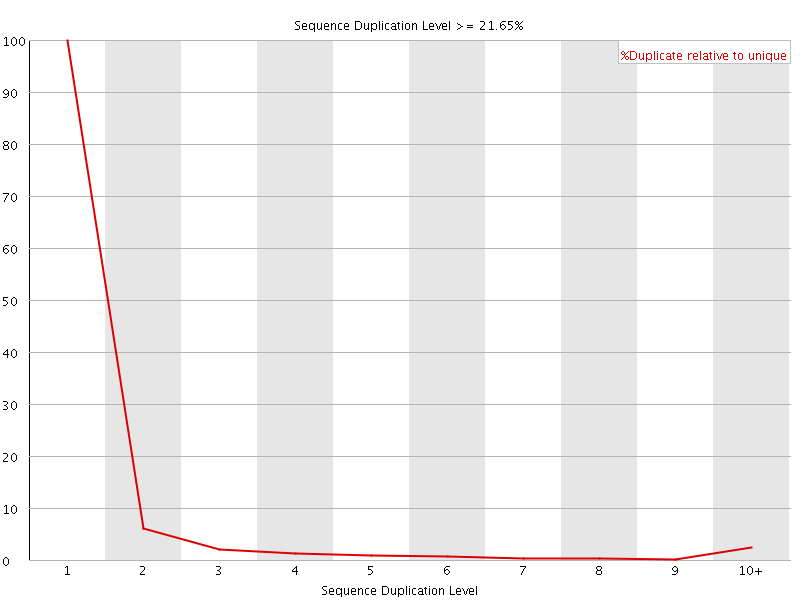
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
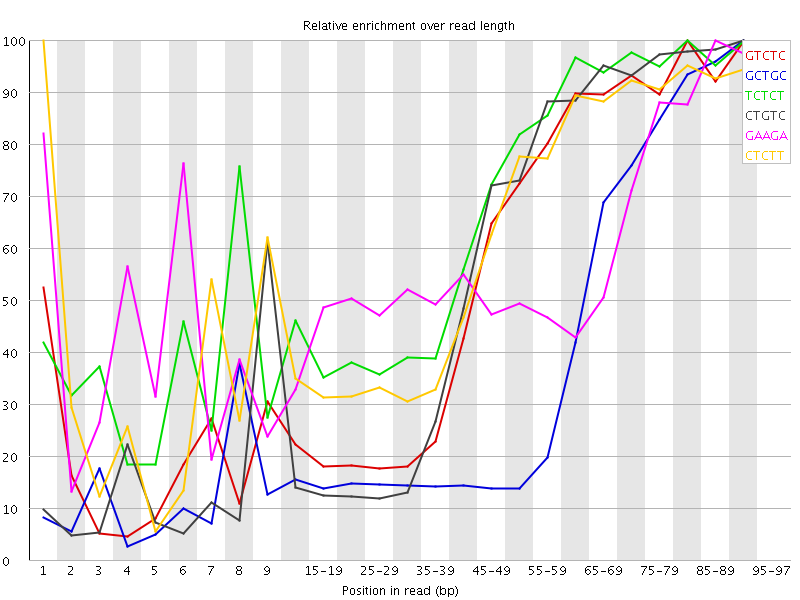
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 318950 | 3.5032566 | 6.030282 | 80-84 |
| GCTGC | 197220 | 3.32071 | 8.157749 | 90-94 |
| TCTCT | 462420 | 3.3132584 | 4.8290815 | 80-84 |
| CTGTC | 293390 | 3.222513 | 5.5335793 | 90-94 |
| GAAGA | 401100 | 3.1640124 | 5.334482 | 85-89 |
| CTCTT | 421170 | 3.0177004 | 4.7722197 | 1 |
| GACGC | 175615 | 3.0072927 | 7.9625144 | 95-97 |
| CGCTG | 173420 | 2.9199753 | 7.782117 | 90-94 |
| CTGCC | 172965 | 2.8467932 | 7.73954 | 90-94 |
| CGACG | 164660 | 2.8196952 | 8.297515 | 95-97 |
| GCCGA | 164545 | 2.817726 | 7.9893966 | 90-94 |
| ACACA | 364475 | 2.7471895 | 7.233596 | 6 |
| CCGAC | 157970 | 2.6442735 | 7.637233 | 95-97 |
| TGCCG | 155705 | 2.6216974 | 7.7148433 | 95-97 |
| GAGGA | 217820 | 2.5898812 | 6.037451 | 90-94 |
| TACAC | 326145 | 2.4171164 | 8.459174 | 5 |
| CTGAC | 208450 | 2.3285484 | 5.649336 | 95-97 |
| AGAGG | 192540 | 2.2893019 | 5.789495 | 90-94 |
| TGACG | 197745 | 2.2598064 | 5.600141 | 95-97 |
| CAAAG | 286070 | 2.2058477 | 5.2253914 | 4 |
| GACGA | 188990 | 2.1965375 | 5.9698615 | 95-97 |
| CTTTG | 283525 | 2.0782242 | 8.595218 | 9 |
| CGAAG | 174275 | 2.0255122 | 5.590286 | 95-97 |
| ACGCT | 180985 | 2.0217433 | 5.111999 | 85-89 |
| AGAGA | 237105 | 1.8703644 | 5.0573144 | 8 |
| GAGAG | 151530 | 1.8016928 | 6.1408653 | 7 |
| GCTTT | 235545 | 1.7265332 | 5.2376323 | 1 |
| TATAC | 335780 | 1.6607081 | 5.568431 | 5 |
| TTGAG | 193940 | 1.4790565 | 5.0002723 | 9 |
| GACTC | 125995 | 1.407462 | 5.4335313 | 7 |
| GAGTC | 122960 | 1.4051723 | 7.049404 | 9 |
| GTGTA | 179990 | 1.3726687 | 10.003576 | 1 |
| AAGAC | 177875 | 1.3715703 | 5.9659977 | 5 |
| CCATG | 118795 | 1.3270326 | 5.3360415 | 9 |
| GTCTT | 178110 | 1.3055375 | 6.4892592 | 1 |
| GACTT | 167650 | 1.2497945 | 6.2615347 | 7 |
| GTGTT | 165650 | 1.2421522 | 6.0384107 | 1 |
| TGAGT | 160490 | 1.2239547 | 5.033597 | 8 |
| TATGA | 241725 | 1.2230449 | 5.6131444 | 4 |
| GATTG | 158305 | 1.2072911 | 5.706649 | 7 |
| TCATA | 241690 | 1.1953558 | 5.7003846 | 2 |
| GTTCT | 162395 | 1.1903473 | 5.226965 | 1 |
| ACATG | 145305 | 1.1016654 | 5.0887156 | 8 |
| TGGAC | 94740 | 1.0826775 | 7.117869 | 5 |
| GTCCA | 94340 | 1.0538512 | 7.505237 | 1 |
| ACACT | 138625 | 1.0273737 | 5.035152 | 6 |
| GGACT | 87595 | 1.0010253 | 6.9827075 | 6 |
| AGACT | 131445 | 0.99658245 | 5.5370913 | 6 |
| GTATG | 122285 | 0.9325896 | 5.156966 | 3 |
| GTATA | 142650 | 0.7217596 | 7.031964 | 1 |