![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-171_CGAGGCTG-TATCCTCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1270236 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
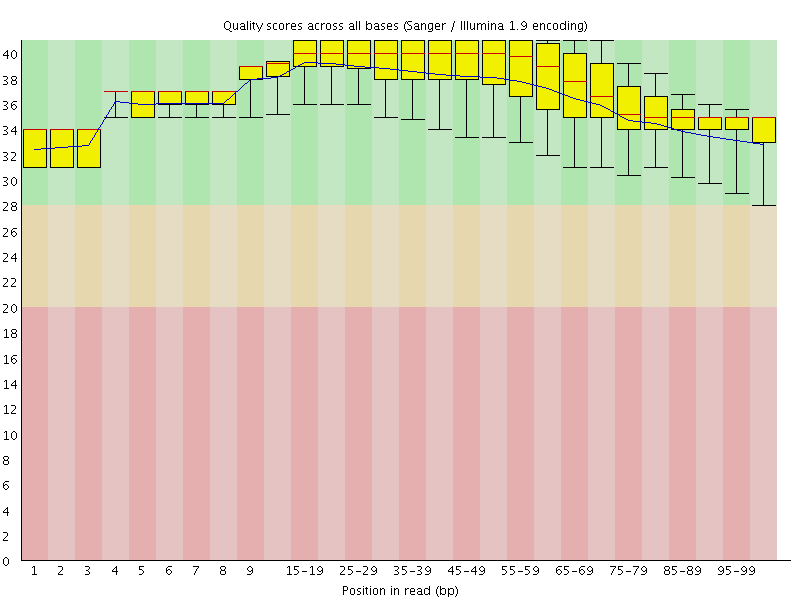
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
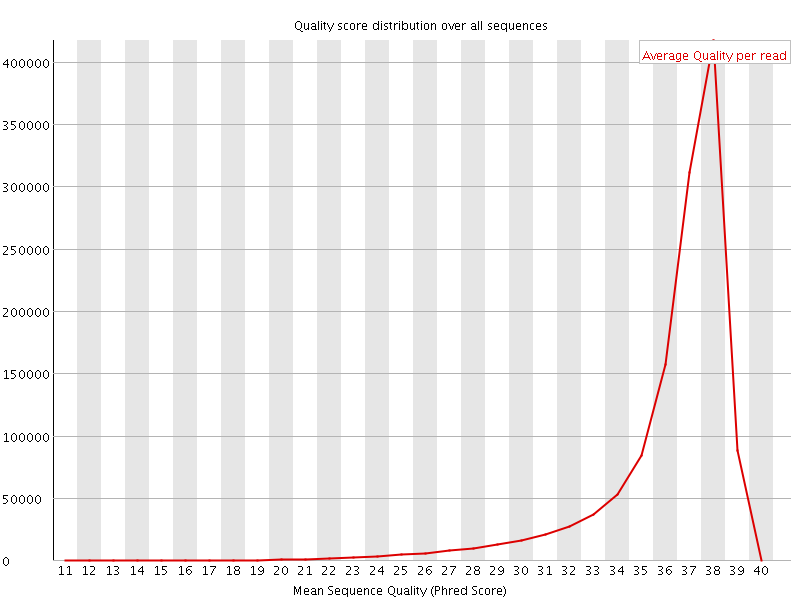
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
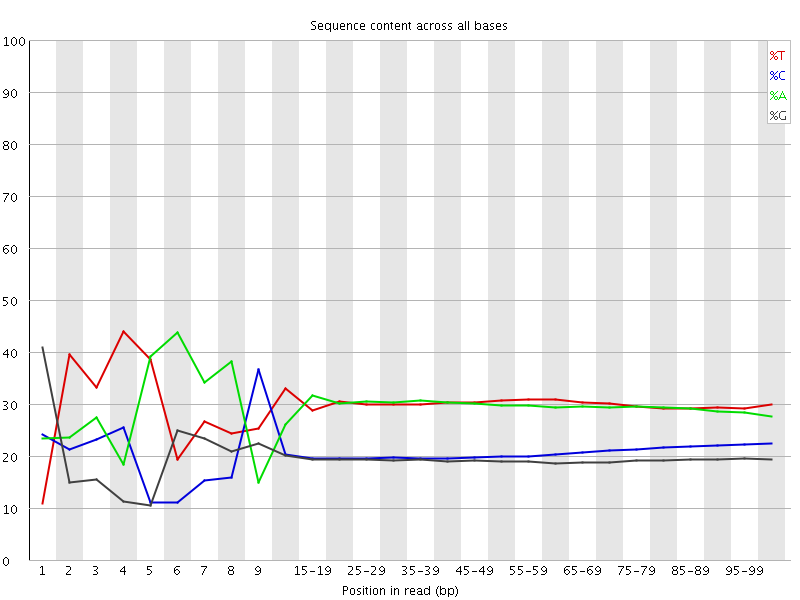
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
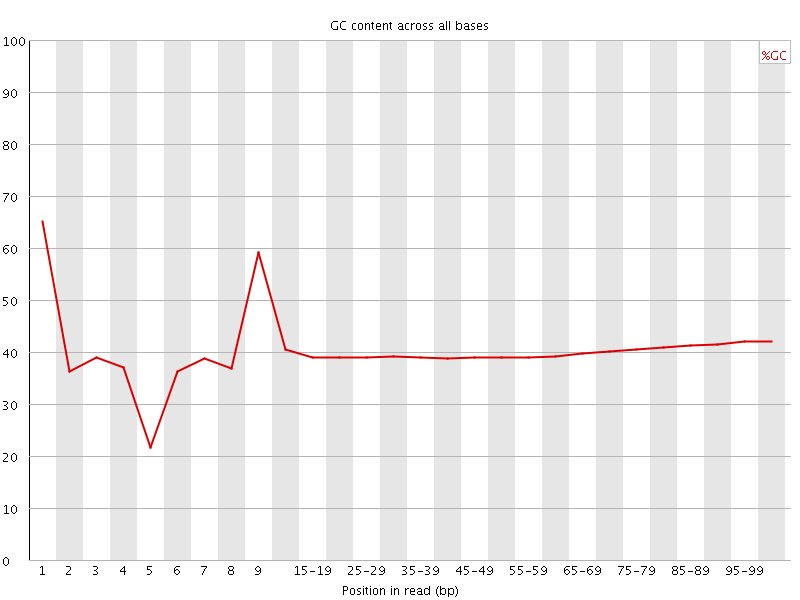
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
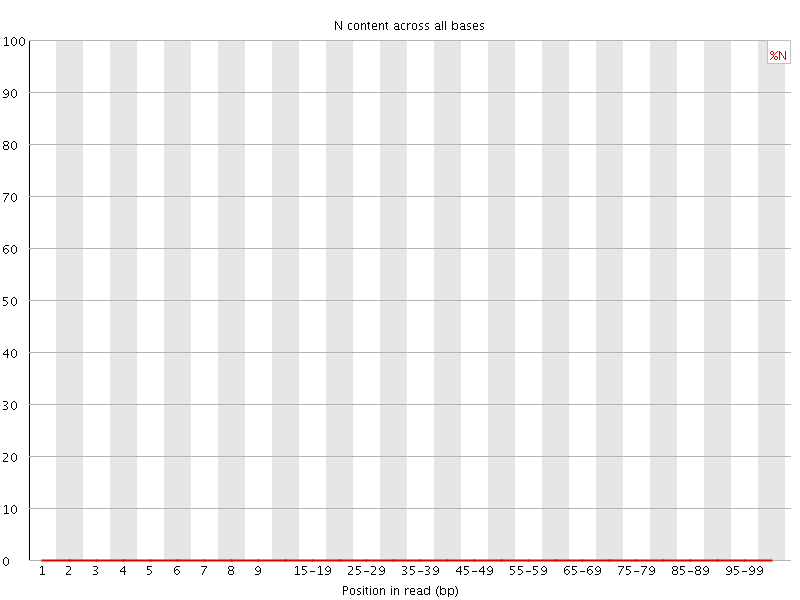
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
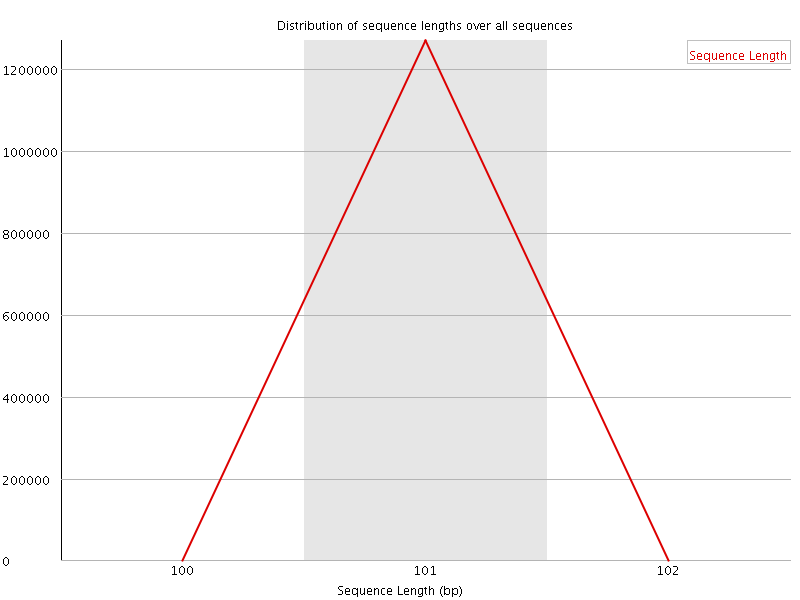
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
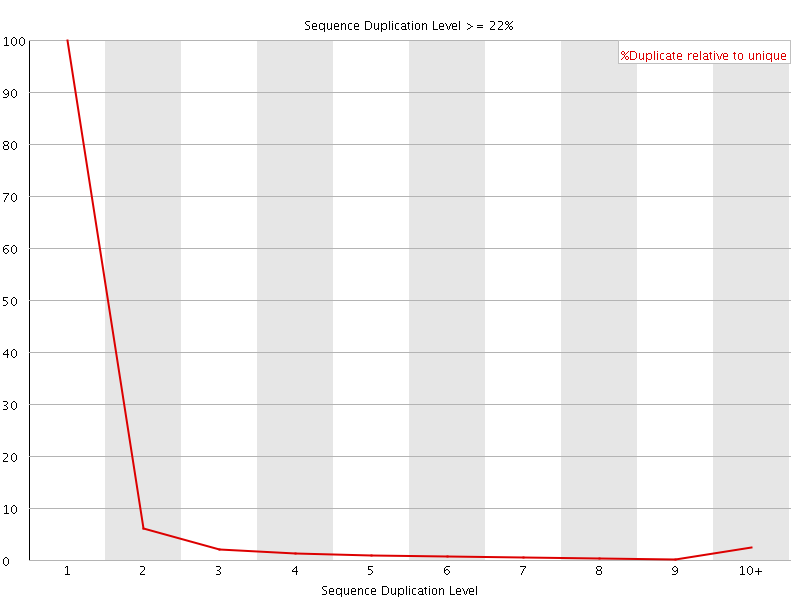
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
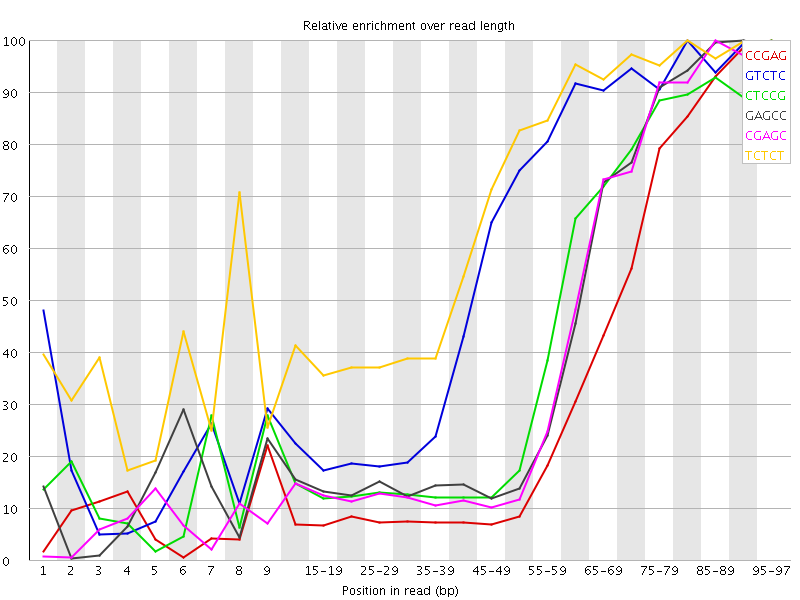
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 290755 | 4.898611 | 14.677623 | 95-97 |
| GTCTC | 318835 | 3.4172206 | 5.83509 | 80-84 |
| CTCCG | 214435 | 3.352757 | 7.9452972 | 95-97 |
| GAGCC | 197530 | 3.327965 | 7.9653187 | 90-94 |
| CGAGC | 189525 | 3.1930976 | 7.9376736 | 85-89 |
| TCTCT | 458935 | 3.1792595 | 4.66837 | 80-84 |
| CTGTC | 294060 | 3.151686 | 5.4122443 | 90-94 |
| TCTCC | 291290 | 2.943742 | 5.9946175 | 95-97 |
| GCCCA | 184220 | 2.9265075 | 7.550581 | 90-94 |
| AGCCC | 183415 | 2.9137194 | 7.4165487 | 90-94 |
| AGAAG | 335935 | 2.7454605 | 5.226703 | 5 |
| CCCAC | 178400 | 2.6722364 | 7.0729837 | 90-94 |
| ACACA | 365290 | 2.6541903 | 6.9192476 | 6 |
| ATCTC | 371180 | 2.6125576 | 6.3764386 | 95-97 |
| CCACG | 162220 | 2.577017 | 7.2979207 | 90-94 |
| TCCGA | 229680 | 2.5011325 | 5.674528 | 95-97 |
| CGAGG | 138935 | 2.482505 | 7.8931384 | 90-94 |
| GGCTG | 136855 | 2.4067595 | 8.353969 | 95-97 |
| GAGGC | 134240 | 2.3986144 | 7.82729 | 90-94 |
| TACAC | 328050 | 2.3459992 | 8.571551 | 5 |
| GAGAC | 197830 | 2.321373 | 5.9799557 | 95-97 |
| CGAGA | 194035 | 2.2768419 | 6.0709925 | 95-97 |
| GACCG | 133985 | 2.2573657 | 7.218994 | 95-97 |
| ACGAG | 180625 | 2.1194866 | 5.7637444 | 95-97 |
| CTTTG | 283730 | 2.0845525 | 8.948502 | 9 |
| AGAGA | 236485 | 1.932696 | 5.1514134 | 6 |
| CACGA | 171875 | 1.9016591 | 5.227211 | 95-97 |
| GAGAG | 150155 | 1.8686391 | 6.4021244 | 7 |
| AGGCT | 161495 | 1.8651141 | 5.478676 | 95-97 |
| AGACC | 167915 | 1.8578452 | 5.3470273 | 95-97 |
| GCTTT | 236220 | 1.7354985 | 5.3334165 | 1 |
| TGGAG | 141260 | 1.7302082 | 5.0836544 | 5 |
| GCTGA | 146065 | 1.6869122 | 5.3069425 | 95-97 |
| TATAC | 337465 | 1.654311 | 5.6047378 | 5 |
| AAGAG | 197920 | 1.6175199 | 5.0523477 | 5 |
| TTGAG | 194015 | 1.5359682 | 5.454615 | 9 |
| GAGTC | 122690 | 1.4169533 | 7.4868793 | 9 |
| GTCTT | 188105 | 1.3819997 | 6.2419147 | 1 |
| GACTC | 125790 | 1.3698078 | 5.353953 | 7 |
| AAGAC | 176860 | 1.3628772 | 5.9520426 | 5 |
| CCATG | 121665 | 1.324888 | 5.533474 | 9 |
| TGAGT | 161605 | 1.2793864 | 5.1743994 | 8 |
| GTGTT | 163820 | 1.2764604 | 6.1362433 | 1 |
| ATGAG | 156335 | 1.2575049 | 5.0350385 | 7 |
| TATGA | 241720 | 1.2567064 | 5.7525444 | 4 |
| GACTT | 168295 | 1.2562768 | 6.32672 | 7 |
| GATTG | 156695 | 1.2405151 | 6.3029404 | 7 |
| GTATG | 154090 | 1.219892 | 5.1820765 | 3 |
| GTTCT | 163140 | 1.1985831 | 5.0519614 | 1 |
| TCATA | 243790 | 1.1951001 | 5.926343 | 2 |
| AACAC | 163835 | 1.190422 | 5.1295443 | 5 |
| GTGTA | 142715 | 1.129839 | 10.104362 | 1 |
| ACATG | 146305 | 1.1096342 | 5.0380716 | 8 |
| TGGAC | 94305 | 1.0891334 | 7.587675 | 5 |
| GTCCA | 95160 | 1.0362581 | 7.324299 | 1 |
| GGACT | 87745 | 1.0133716 | 7.106096 | 6 |
| ACACT | 139325 | 0.9963613 | 5.2323904 | 6 |
| AGACT | 131140 | 0.99461687 | 5.6485243 | 6 |
| GCCGT | 58320 | 0.9670673 | 5.1378627 | 95-97 |
| GAGTA | 120030 | 0.96547997 | 5.0551558 | 1 |
| TGCCG | 53210 | 0.8823329 | 5.2772317 | 95-97 |
| GTATA | 142015 | 0.73833835 | 6.8096046 | 1 |