![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-170_CGAGGCTG-CTCTCTAT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2431957 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
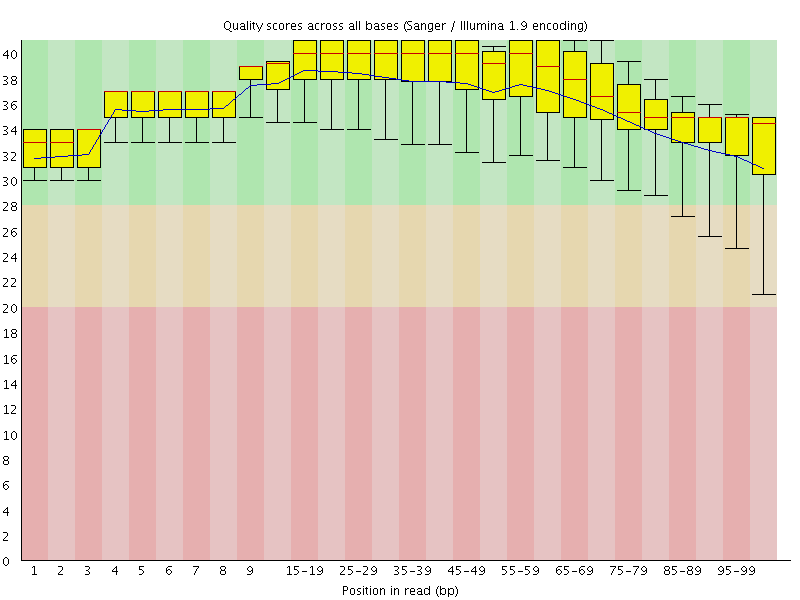
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
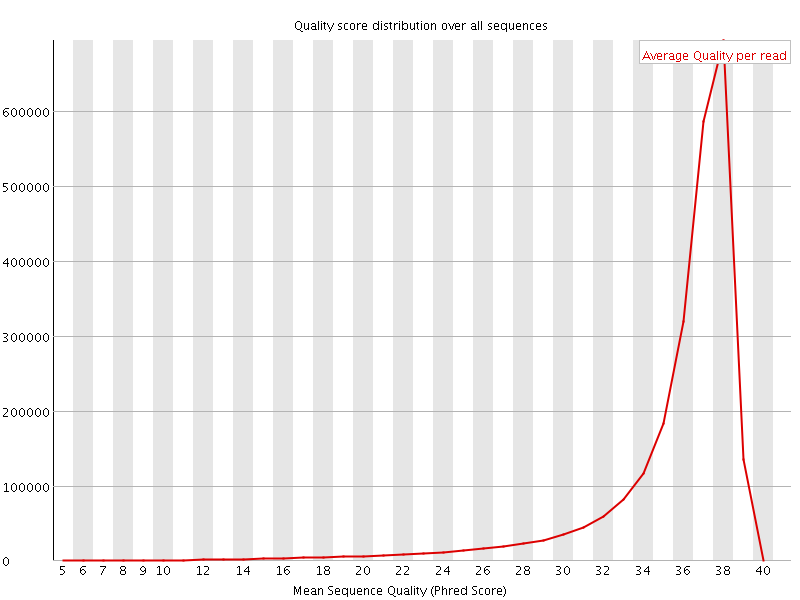
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
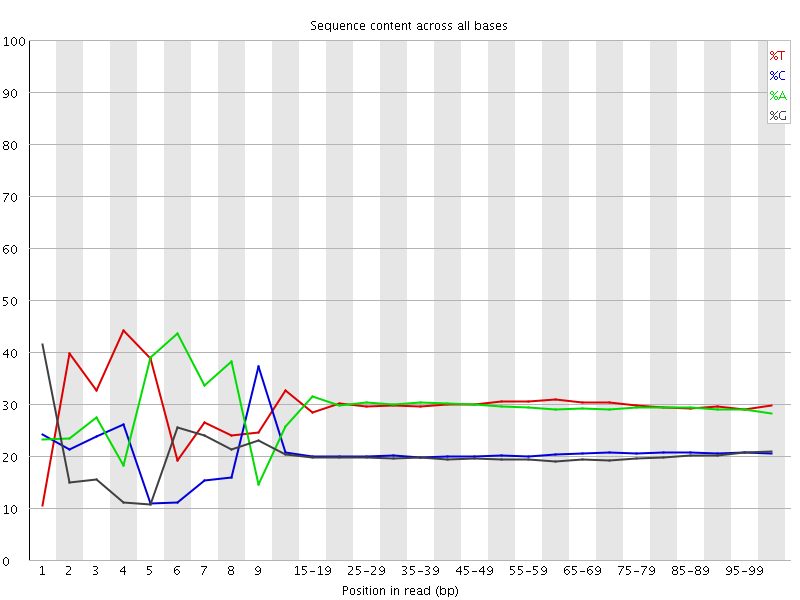
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
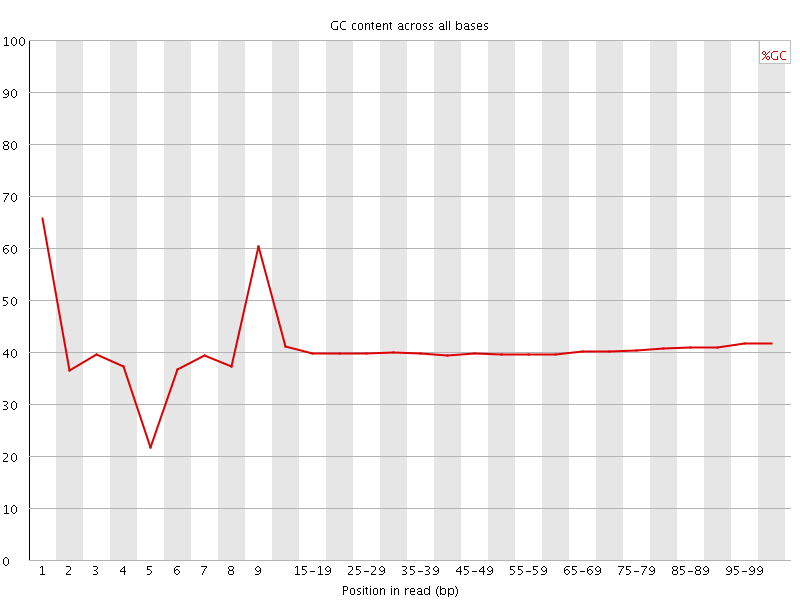
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
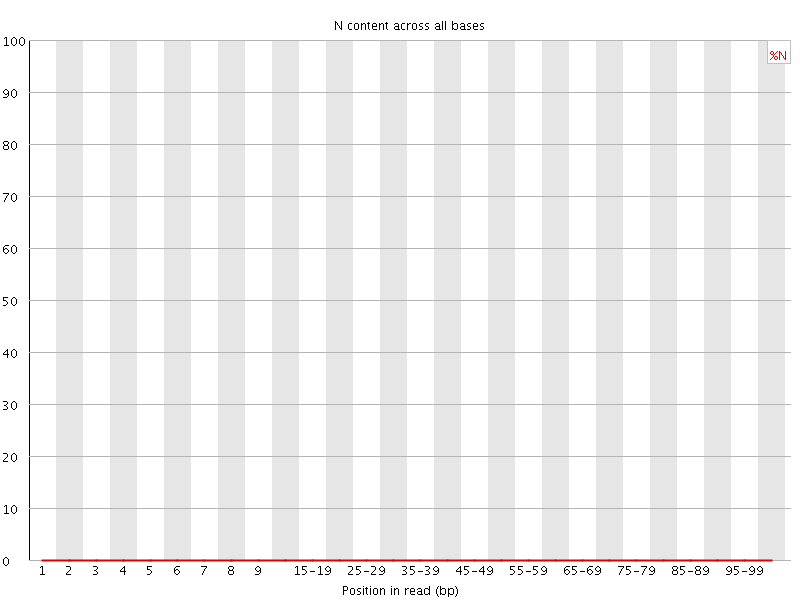
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
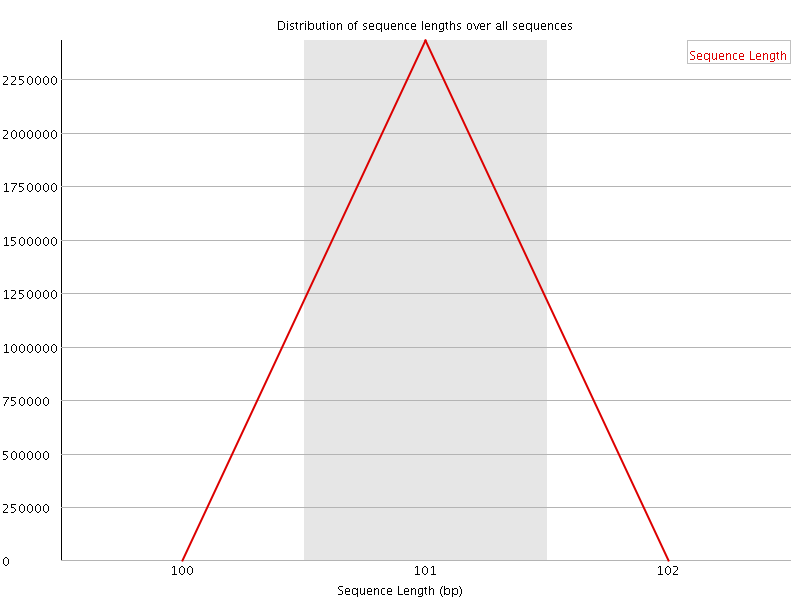
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
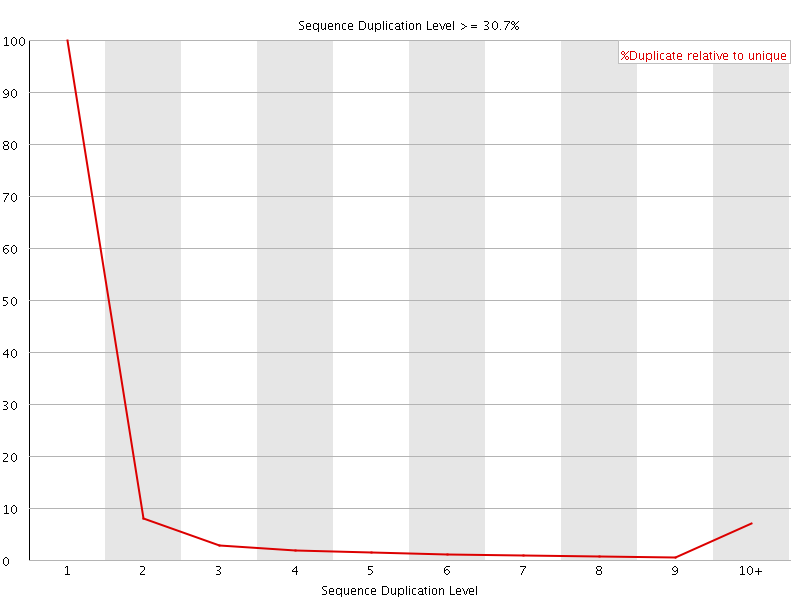
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
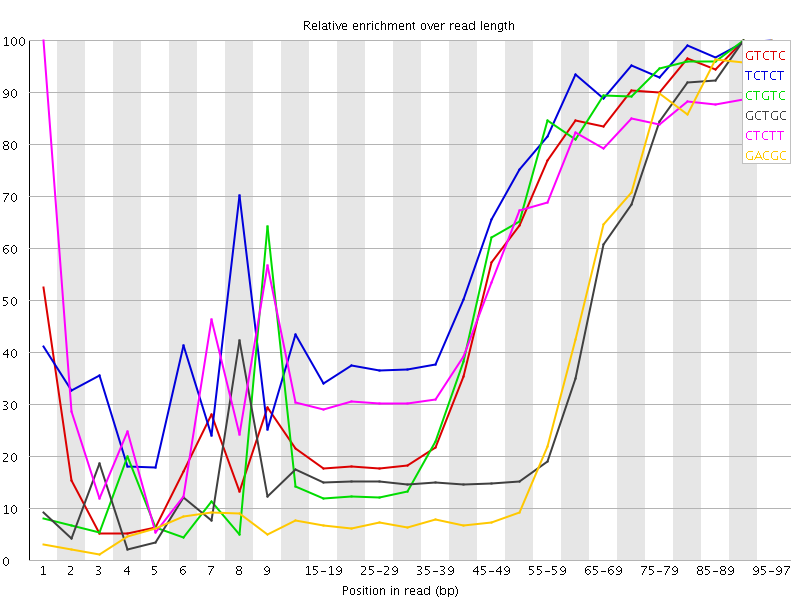
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 587885 | 3.2999358 | 5.9117684 | 90-94 |
| TCTCT | 845070 | 3.1547356 | 4.7541723 | 95-97 |
| CTGTC | 544400 | 3.0558445 | 5.5442567 | 90-94 |
| GCTGC | 359520 | 3.0344431 | 7.6283946 | 90-94 |
| CTCTT | 781125 | 2.9160223 | 5.036152 | 1 |
| GACGC | 315705 | 2.7116976 | 7.469733 | 95-97 |
| ACACA | 681170 | 2.680015 | 7.7102847 | 6 |
| AGAAG | 645870 | 2.6689363 | 5.0855246 | 5 |
| CGCTG | 312975 | 2.641591 | 7.227245 | 90-94 |
| CTGCC | 314695 | 2.591732 | 7.1703043 | 90-94 |
| GCCGA | 296850 | 2.5497456 | 7.4572353 | 95-97 |
| CGACG | 293840 | 2.5238917 | 7.6822023 | 95-97 |
| GAGAG | 404245 | 2.4681797 | 5.91906 | 7 |
| TACAC | 616525 | 2.383574 | 9.532795 | 5 |
| CCGAC | 284030 | 2.3805008 | 7.1626716 | 95-97 |
| TGCCG | 280685 | 2.369055 | 7.1191797 | 90-94 |
| CACAT | 591415 | 2.286495 | 5.32856 | 7 |
| CAAAG | 560910 | 2.2616773 | 5.310427 | 4 |
| CTGAC | 379045 | 2.1652484 | 5.4785023 | 95-97 |
| CTTTG | 550625 | 2.106599 | 8.939167 | 9 |
| TGACG | 357735 | 2.0942767 | 5.4498935 | 95-97 |
| GACGA | 338045 | 2.0139606 | 5.552896 | 95-97 |
| AGAGG | 324170 | 1.9792693 | 5.5328827 | 95-97 |
| ATACA | 715660 | 1.9191164 | 5.187504 | 6 |
| GCTTT | 463760 | 1.7742682 | 5.1742363 | 1 |
| GCCAA | 286995 | 1.6683805 | 5.3664503 | 1 |
| TATAC | 627175 | 1.6526445 | 5.9936743 | 5 |
| TGGCT | 279765 | 1.6093932 | 5.361941 | 6 |
| TTGAG | 373615 | 1.4907669 | 5.5845222 | 9 |
| GACTC | 257265 | 1.469595 | 5.7705083 | 7 |
| GAGTC | 246660 | 1.4440137 | 7.3588824 | 9 |
| AAGAC | 350745 | 1.4142588 | 5.879611 | 5 |
| GTGTA | 348655 | 1.3911736 | 10.744451 | 1 |
| CCATG | 243380 | 1.3902786 | 5.850798 | 9 |
| GTCTT | 348095 | 1.3317531 | 6.707206 | 1 |
| GACTT | 333995 | 1.3003783 | 6.740713 | 7 |
| TATGA | 476535 | 1.2868894 | 6.407171 | 4 |
| TCATA | 480735 | 1.2667663 | 6.616097 | 2 |
| TGAGT | 313320 | 1.2501829 | 5.3369694 | 8 |
| ACTCA | 318800 | 1.2325264 | 5.098027 | 8 |
| GTGTT | 309480 | 1.2134285 | 5.8809657 | 1 |
| GATTG | 303630 | 1.2115186 | 6.2521625 | 7 |
| AACAC | 307395 | 1.2094238 | 5.1798086 | 5 |
| GTTCT | 308665 | 1.1809006 | 5.1129913 | 1 |
| ACACC | 203920 | 1.1567112 | 5.22384 | 6 |
| TGGAC | 190555 | 1.11556 | 7.6193066 | 5 |
| ACATG | 280390 | 1.1109542 | 5.5820637 | 8 |
| GTCCA | 189350 | 1.0816388 | 8.01665 | 1 |
| GGACT | 175985 | 1.0302633 | 7.2253723 | 6 |
| ACACT | 263710 | 1.0195405 | 5.2890882 | 6 |
| AGACT | 257265 | 1.0193288 | 5.772094 | 6 |
| GAGTA | 240490 | 0.9765315 | 5.505523 | 1 |
| GTATG | 237095 | 0.94603634 | 5.585901 | 3 |
| CATAC | 232700 | 0.89965135 | 5.1816754 | 3 |
| GTATA | 279415 | 0.7545641 | 7.3111544 | 1 |