![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-170_CGAGGCTG-CTCTCTAT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2431957 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
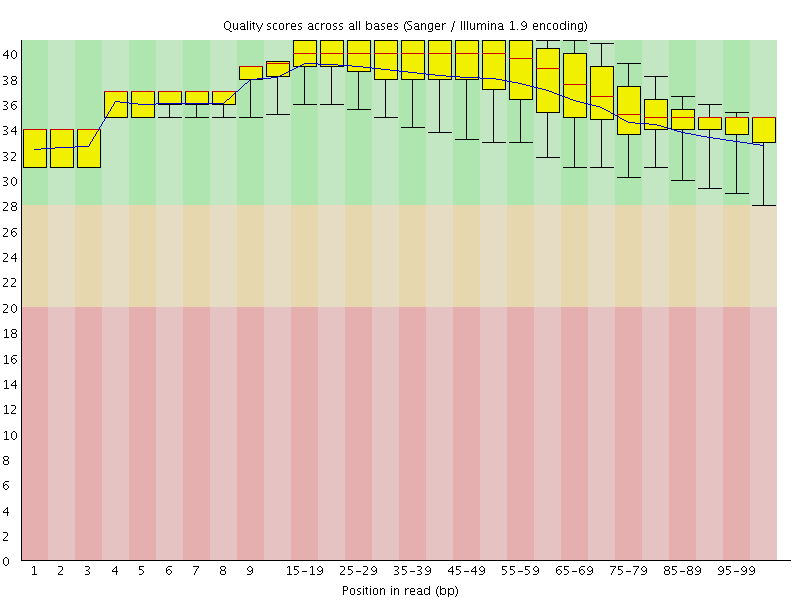
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
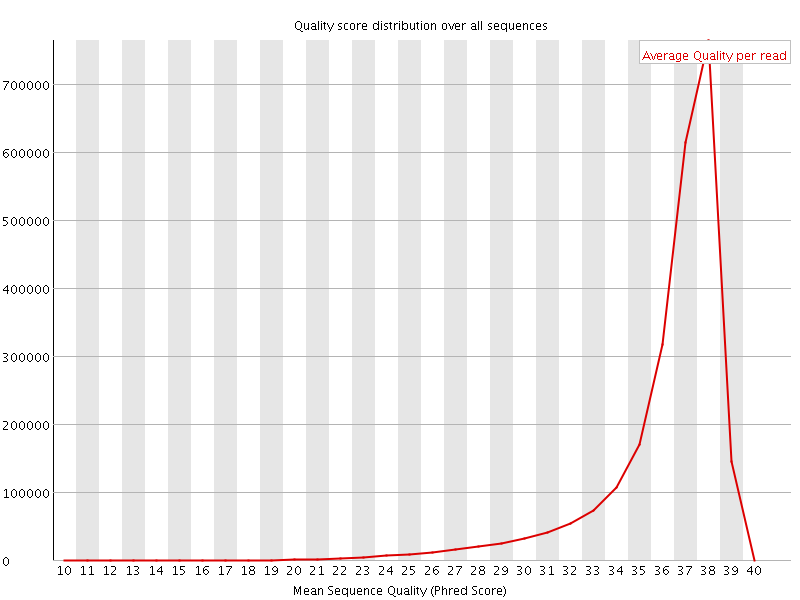
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
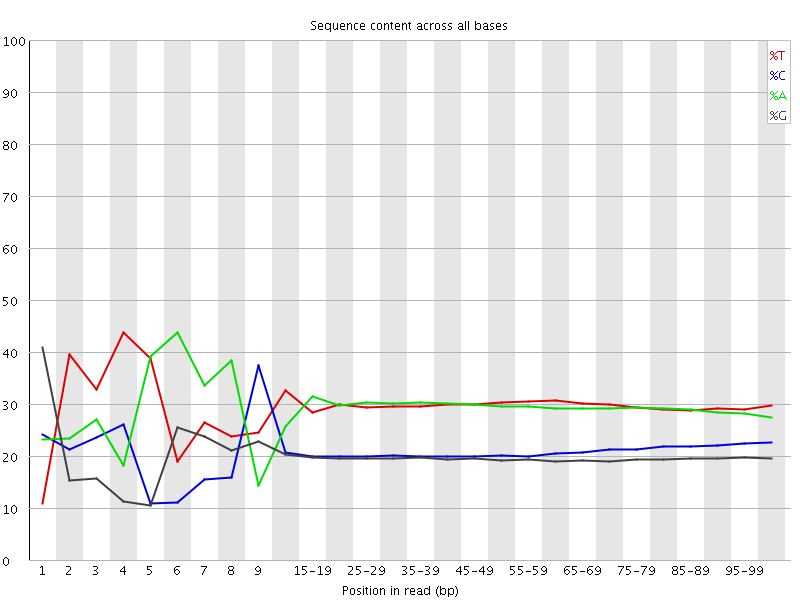
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
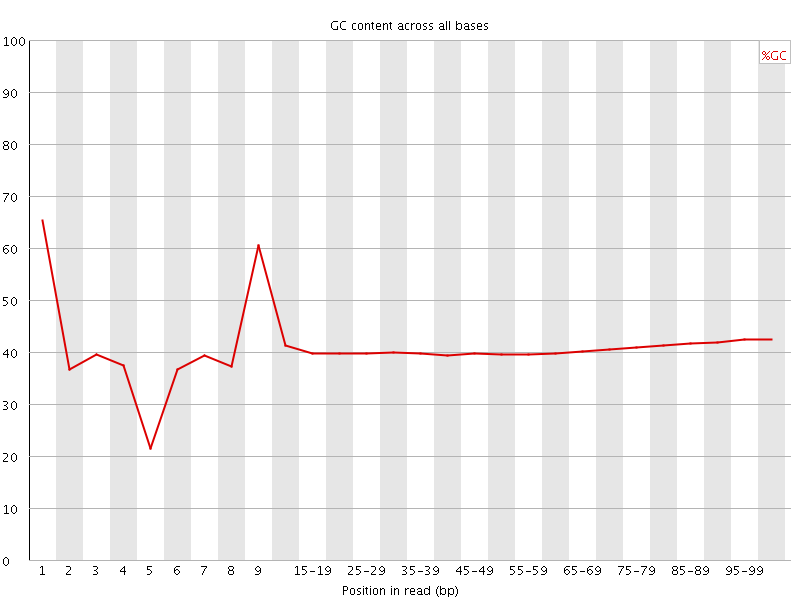
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
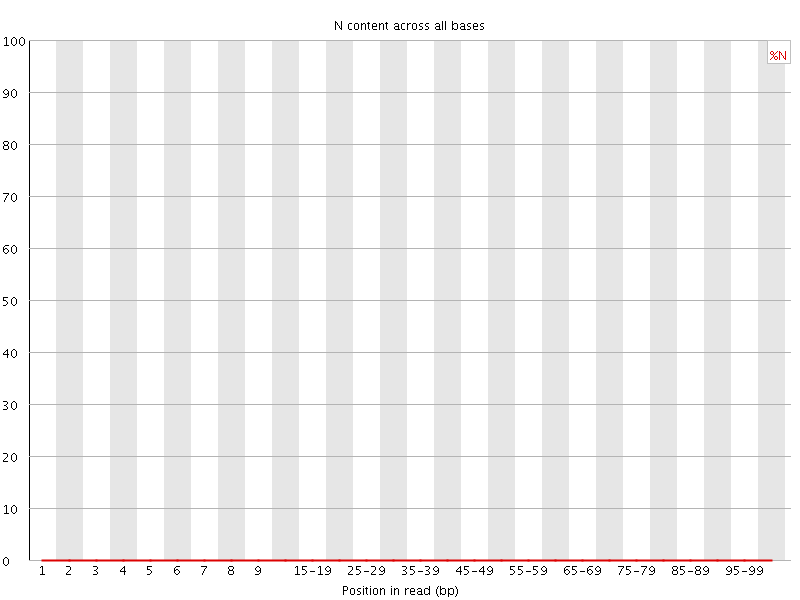
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
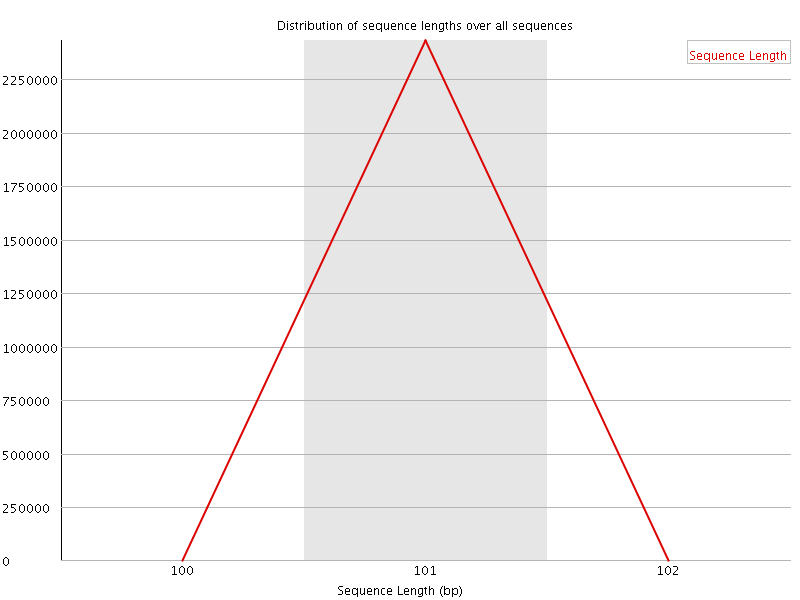
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
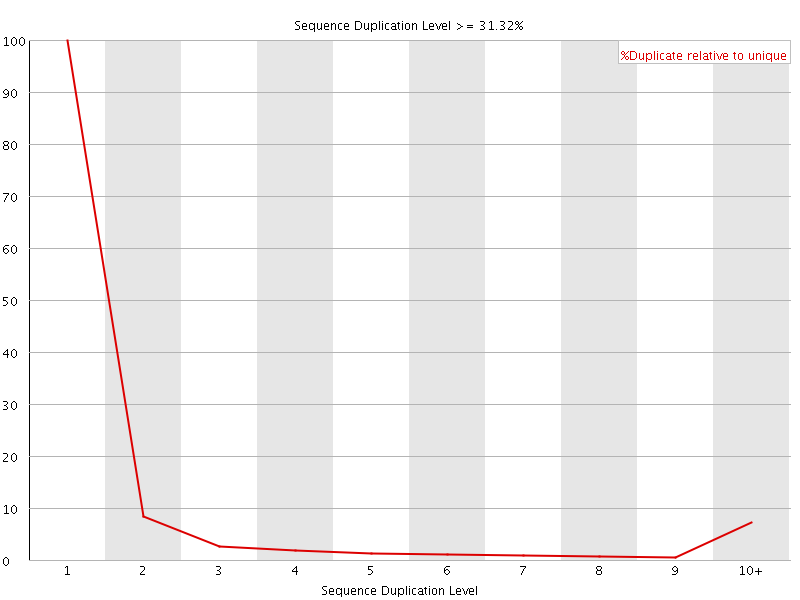
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CATATGGACTTTGGCTACACCATGAAAGCTTTGAGAAGCAAGAAGAAGGT | 2479 | 0.10193436808298832 | No Hit |
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
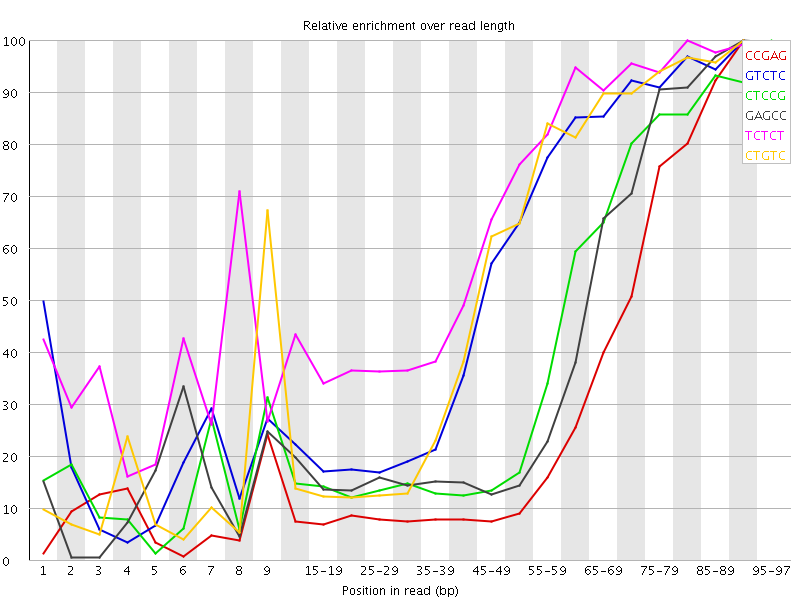
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 520290 | 4.390524 | 13.585603 | 90-94 |
| GTCTC | 585585 | 3.2220805 | 5.740527 | 90-94 |
| CTCCG | 396040 | 3.1158435 | 7.492203 | 95-97 |
| GAGCC | 365120 | 3.081105 | 7.513368 | 90-94 |
| TCTCT | 843415 | 3.0694442 | 4.607011 | 80-84 |
| CTGTC | 545520 | 3.0016298 | 5.4342375 | 90-94 |
| CGAGC | 346600 | 2.9248219 | 7.38413 | 95-97 |
| TCTCC | 533410 | 2.7756824 | 5.70228 | 95-97 |
| GCCCA | 345070 | 2.7538497 | 7.1032147 | 90-94 |
| AGCCC | 344695 | 2.750857 | 7.0165334 | 90-94 |
| AGAAG | 647170 | 2.748538 | 5.275586 | 5 |
| ACACA | 683965 | 2.5980153 | 7.5085626 | 6 |
| ATCTC | 672645 | 2.4831378 | 6.146169 | 95-97 |
| CCCAC | 325180 | 2.4542515 | 6.6576276 | 90-94 |
| TCCGA | 423775 | 2.3652563 | 5.463098 | 95-97 |
| CCACG | 292395 | 2.3334742 | 6.811762 | 95-97 |
| TACAC | 621570 | 2.327563 | 9.624484 | 5 |
| CGAGG | 255340 | 2.2783873 | 7.366566 | 90-94 |
| CAAAG | 564460 | 2.2671423 | 5.333913 | 4 |
| CACAT | 592160 | 2.2174327 | 5.366907 | 7 |
| GGCTG | 251785 | 2.2148387 | 7.71298 | 95-97 |
| GAGGC | 242690 | 2.1655118 | 7.3129153 | 90-94 |
| GAGAC | 358350 | 2.1452827 | 5.7271123 | 95-97 |
| CTTTG | 552650 | 2.1267009 | 9.064044 | 9 |
| CGAGA | 353950 | 2.1189418 | 5.5742345 | 95-97 |
| GACCG | 242485 | 2.0462363 | 6.689907 | 95-97 |
| ACGAG | 329405 | 1.9720019 | 5.552948 | 95-97 |
| GAGAG | 285525 | 1.80742 | 5.809969 | 7 |
| CACGA | 317925 | 1.799965 | 5.0255094 | 95-97 |
| GCTTT | 458410 | 1.7640479 | 5.344403 | 1 |
| AGACC | 307020 | 1.738225 | 5.058451 | 95-97 |
| AGGCT | 294225 | 1.736442 | 5.095594 | 95-97 |
| TATAC | 626725 | 1.6413387 | 6.0886087 | 5 |
| TGGCT | 278640 | 1.6211667 | 5.2638574 | 6 |
| GCTGA | 263695 | 1.5562615 | 5.116579 | 95-97 |
| TTGAG | 370990 | 1.5312734 | 5.8776364 | 9 |
| GAGTC | 247090 | 1.4582629 | 7.345389 | 9 |
| GACTC | 257180 | 1.4354236 | 5.3879356 | 7 |
| AAGAC | 352780 | 1.4169338 | 6.1615543 | 5 |
| GTCTT | 363115 | 1.3973348 | 6.3222194 | 1 |
| CATGA | 345610 | 1.3684705 | 5.0414114 | 9 |
| CCATG | 244290 | 1.3634794 | 5.7776203 | 9 |
| TATGA | 478095 | 1.3239548 | 6.5575695 | 4 |
| GACTT | 334710 | 1.3065361 | 6.947768 | 7 |
| TGAGT | 313495 | 1.2939608 | 5.389334 | 8 |
| ATGAG | 308650 | 1.29227 | 5.0729604 | 7 |
| TCATA | 483930 | 1.2673709 | 6.4882407 | 2 |
| GTGTT | 308025 | 1.2533722 | 5.891878 | 1 |
| GATTG | 300115 | 1.2387342 | 6.5200334 | 7 |
| ACTCA | 320395 | 1.1997675 | 5.1671906 | 8 |
| GTGTA | 288445 | 1.1905662 | 10.864262 | 1 |
| GTTCT | 307305 | 1.1825672 | 5.0831532 | 1 |
| GTATG | 285315 | 1.1776468 | 5.687518 | 3 |
| ACATG | 284175 | 1.1252136 | 5.6999044 | 8 |
| TGGAC | 188465 | 1.1122729 | 7.6487045 | 5 |
| ACACC | 206885 | 1.1077214 | 5.2439814 | 6 |
| GTCCA | 191445 | 1.0685306 | 7.5295653 | 1 |
| GGACT | 176050 | 1.0390028 | 7.322498 | 6 |
| AGACT | 257305 | 1.0188198 | 5.6883855 | 6 |
| GAGTA | 239840 | 1.0041732 | 5.6035843 | 1 |
| ACACT | 265355 | 0.9936619 | 5.1599283 | 6 |
| TGTAT | 362335 | 0.98917425 | 5.083509 | 2 |
| CATAC | 235525 | 0.881959 | 5.2162123 | 3 |
| GTATA | 278820 | 0.7721166 | 7.133241 | 1 |
| ATACT | 288235 | 0.7548626 | 5.0448475 | 6 |