![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-168_GCTACGCT-CTAAGCCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1140236 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
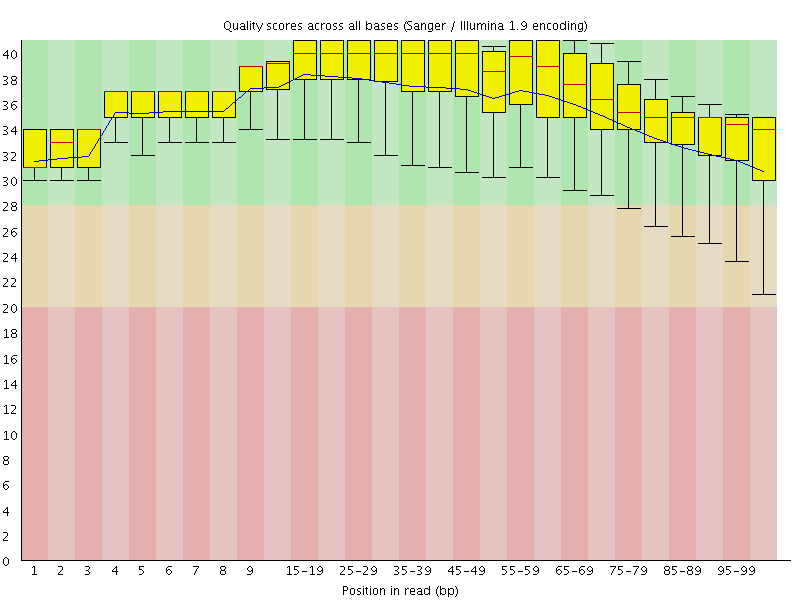
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
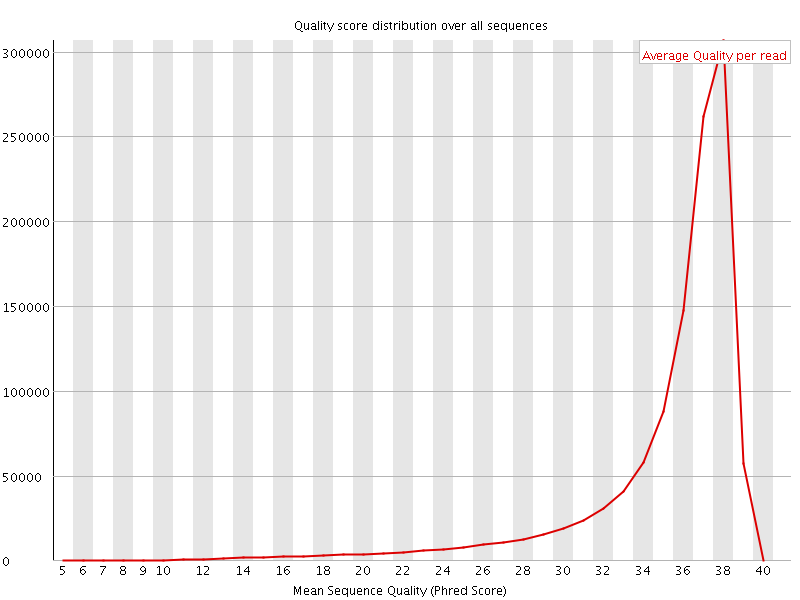
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
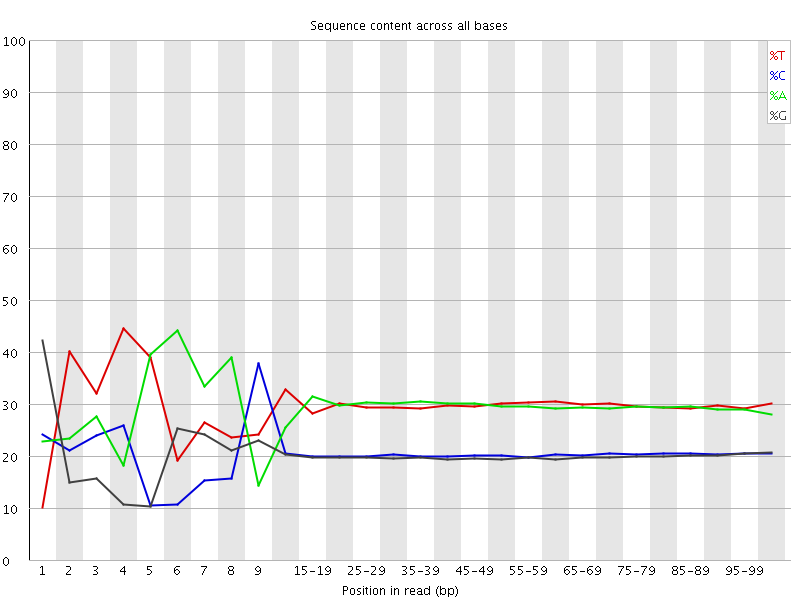
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
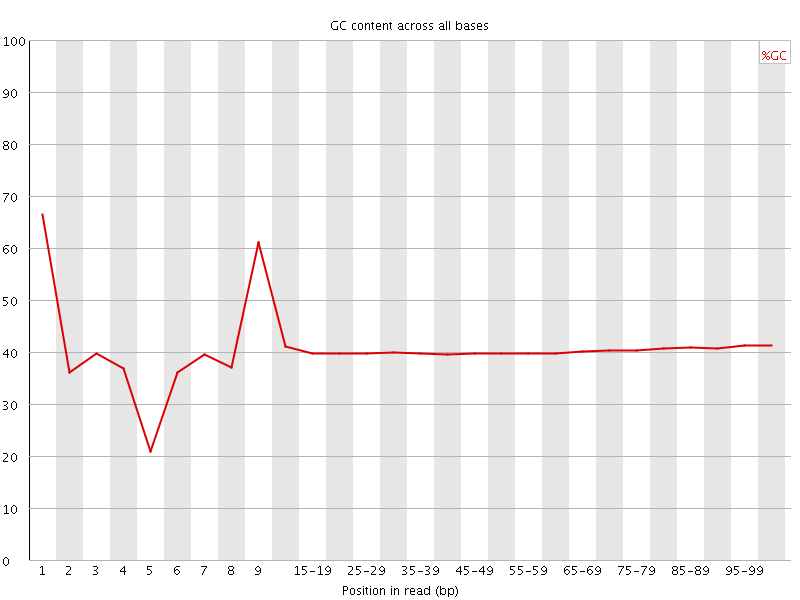
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
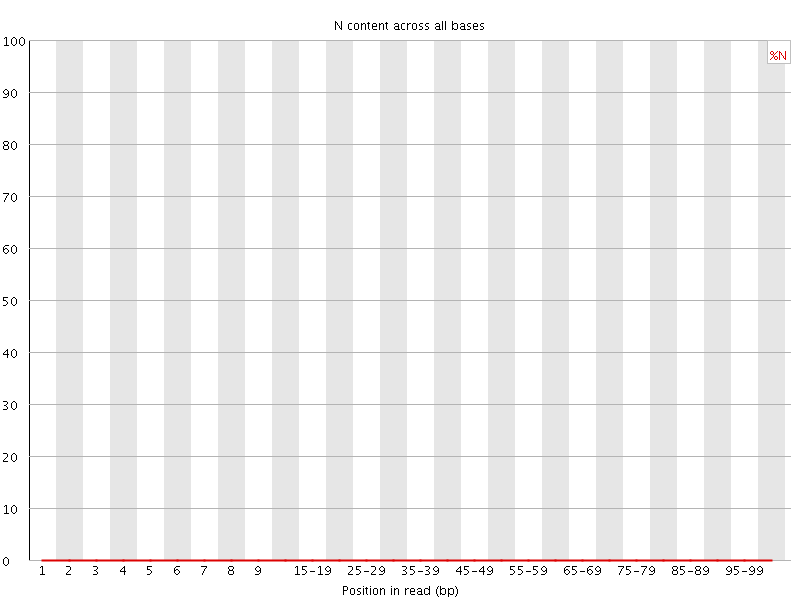
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
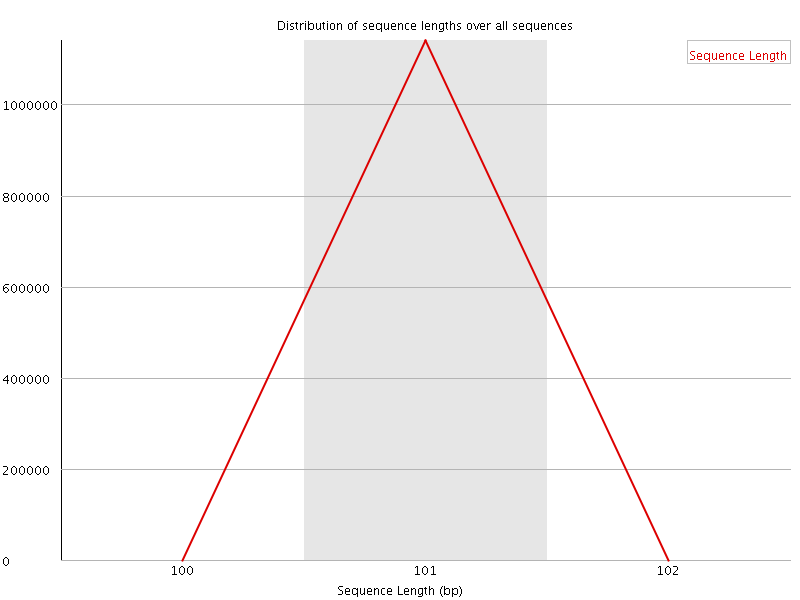
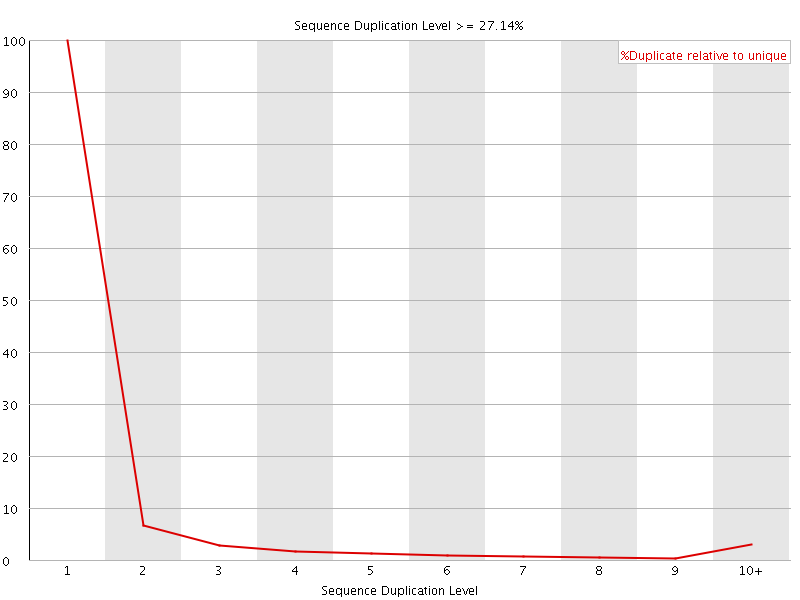
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CATATGGACTTTGGCTACACCATGAAAGCTTTGAGAAGCAAGAAGAAGGT | 1329 | 0.1165548184761751 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
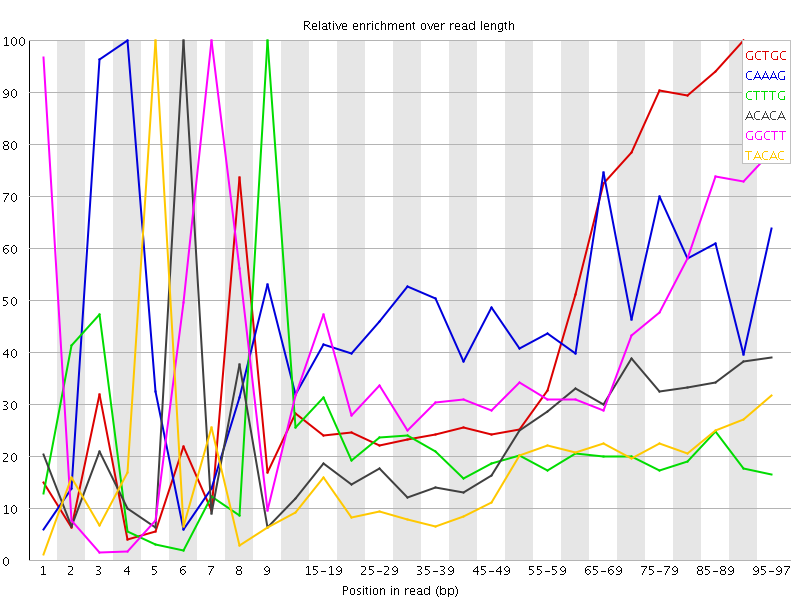
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 136435 | 2.457991 | 5.1710663 | 90-94 |
| CAAAG | 287895 | 2.44289 | 5.075388 | 4 |
| CTTTG | 279705 | 2.3127484 | 10.822188 | 9 |
| ACACA | 274210 | 2.2923272 | 9.243104 | 6 |
| GGCTT | 168980 | 2.077853 | 5.0925655 | 7 |
| TACAC | 242395 | 2.0089529 | 11.635392 | 5 |
| GCTTT | 236155 | 1.9526541 | 5.475189 | 1 |
| CACAT | 228725 | 1.895657 | 6.792537 | 7 |
| GCCAA | 144950 | 1.786552 | 6.0089946 | 1 |
| TTGGC | 138540 | 1.7035495 | 5.392429 | 5 |
| TGGCT | 138400 | 1.7018278 | 6.0226917 | 6 |
| CTCCA | 140770 | 1.6946652 | 5.1158314 | 1 |
| ATACA | 288435 | 1.6457592 | 5.8403277 | 4 |
| CACCA | 134750 | 1.6362504 | 5.040737 | 7 |
| GAGAG | 127895 | 1.6240672 | 5.5484858 | 7 |
| GACTC | 132340 | 1.6171169 | 6.553973 | 7 |
| TTGAG | 186665 | 1.5802097 | 6.8557076 | 9 |
| GAGTC | 124450 | 1.5435535 | 8.817485 | 9 |
| AAGAC | 175255 | 1.4871 | 6.6519275 | 5 |
| ATGGA | 172920 | 1.4765366 | 5.3144956 | 4 |
| CCATG | 118565 | 1.4487945 | 6.198205 | 9 |
| CATGA | 172010 | 1.4470257 | 6.31914 | 9 |
| GTGTA | 169505 | 1.434942 | 12.870313 | 1 |
| GACTT | 170050 | 1.4182475 | 8.10536 | 7 |
| GTCTT | 170075 | 1.4062698 | 7.3604193 | 1 |
| TATGA | 243965 | 1.4007968 | 7.9828906 | 4 |
| TCCAT | 169890 | 1.3959398 | 5.3769007 | 2 |
| TCATA | 246715 | 1.3956184 | 7.9604945 | 2 |
| ACTCA | 164970 | 1.3672597 | 5.9686685 | 8 |
| TGAGT | 159590 | 1.3510066 | 6.236121 | 8 |
| TATAC | 235850 | 1.3341572 | 6.289587 | 5 |
| ATGAG | 154450 | 1.3188242 | 5.896689 | 7 |
| AACAC | 156195 | 1.3057512 | 5.446352 | 5 |
| GTGTT | 154670 | 1.2981076 | 6.388014 | 1 |
| GATTG | 151980 | 1.2865845 | 7.664705 | 7 |
| ACACC | 101565 | 1.2332896 | 5.4881587 | 6 |
| GTTCT | 148290 | 1.2261399 | 5.282655 | 1 |
| ATGAT | 212965 | 1.2228011 | 5.038747 | 5 |
| ACCAT | 146130 | 1.2111152 | 5.245194 | 8 |
| TGGAC | 95775 | 1.1878972 | 8.808545 | 5 |
| GTCCA | 95475 | 1.1666483 | 9.093142 | 1 |
| ACATG | 138070 | 1.161507 | 6.9558764 | 8 |
| ATTGA | 199335 | 1.1445404 | 5.4743943 | 8 |
| GGACT | 91265 | 1.1319598 | 8.228157 | 6 |
| ACACT | 135965 | 1.1268685 | 5.7916164 | 6 |
| AGACT | 125655 | 1.0570664 | 6.4212117 | 6 |
| TGTAT | 183180 | 1.0427458 | 6.1246758 | 2 |
| GTATG | 121790 | 1.0310113 | 7.1452975 | 3 |
| GAGTA | 119060 | 1.0166347 | 5.6624694 | 1 |
| CATAC | 120340 | 0.9973695 | 6.838639 | 3 |
| GAATA | 155145 | 0.89853 | 5.0908623 | 1 |
| TATAA | 217370 | 0.8465313 | 5.278262 | 2 |
| ATACT | 142480 | 0.80598146 | 5.310877 | 6 |
| GTATA | 139395 | 0.80037737 | 8.007995 | 1 |