![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-168_GCTACGCT-CTAAGCCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1140236 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
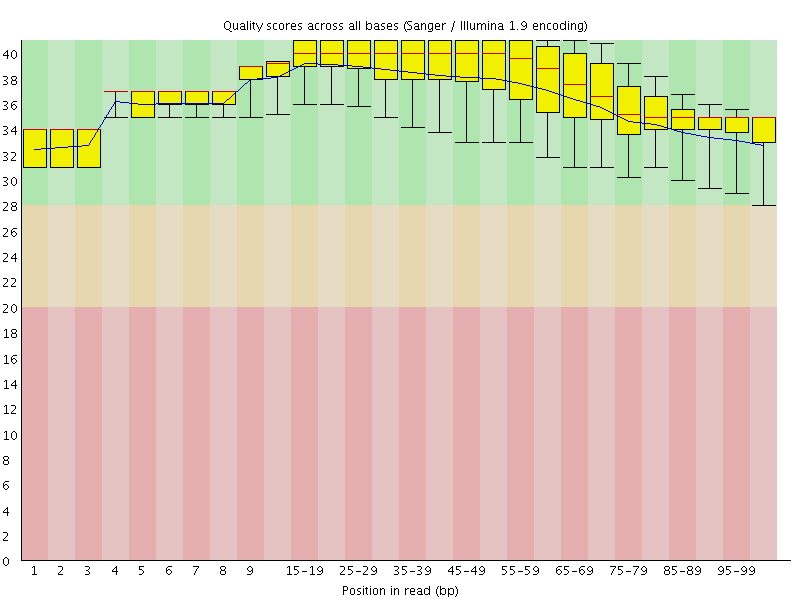
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
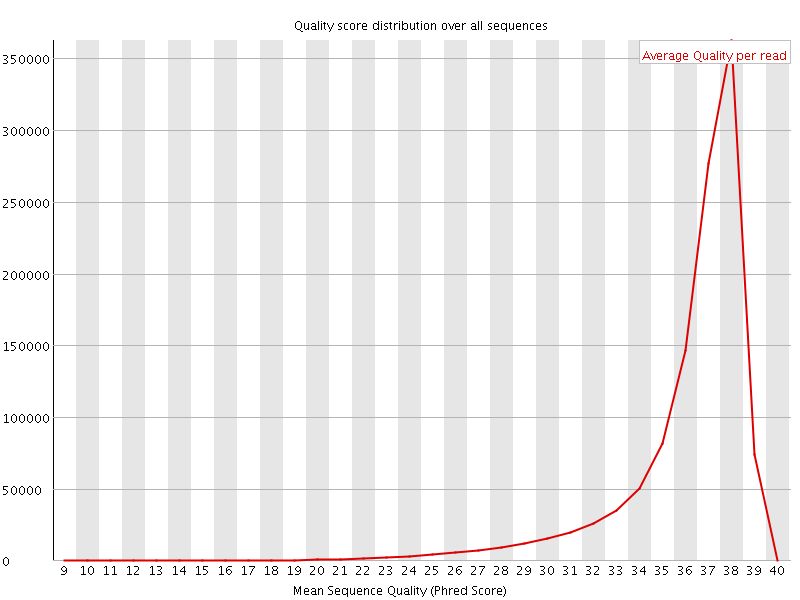
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
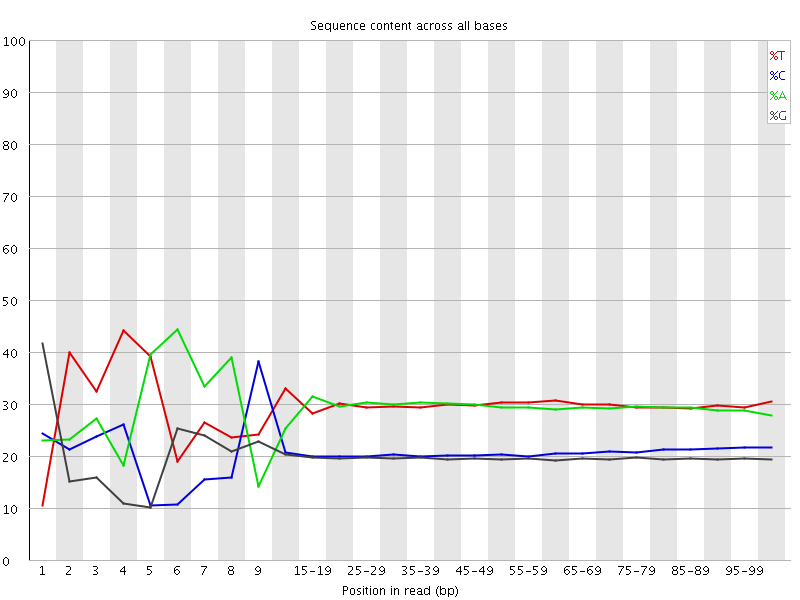
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
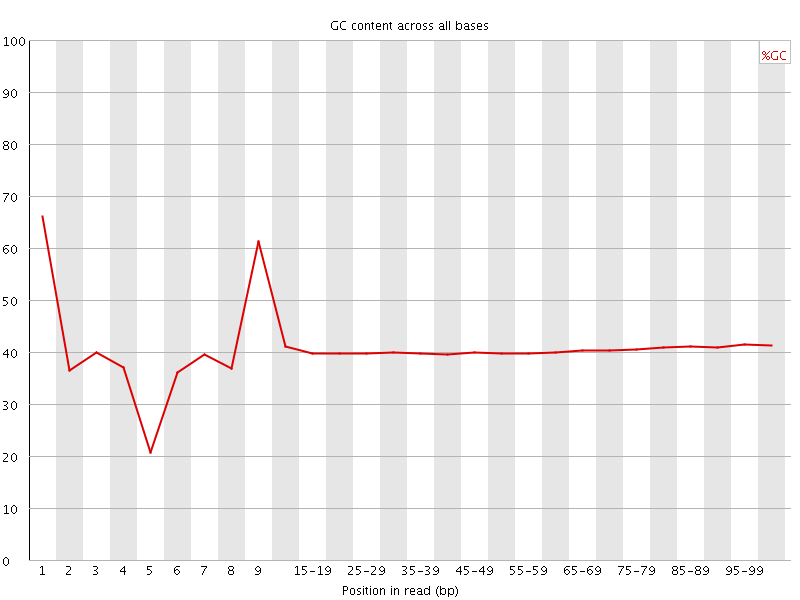
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
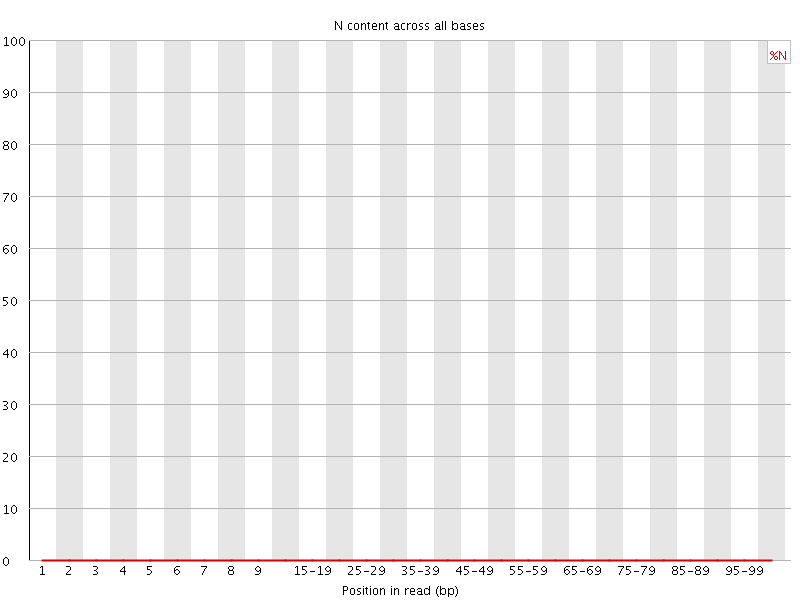
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
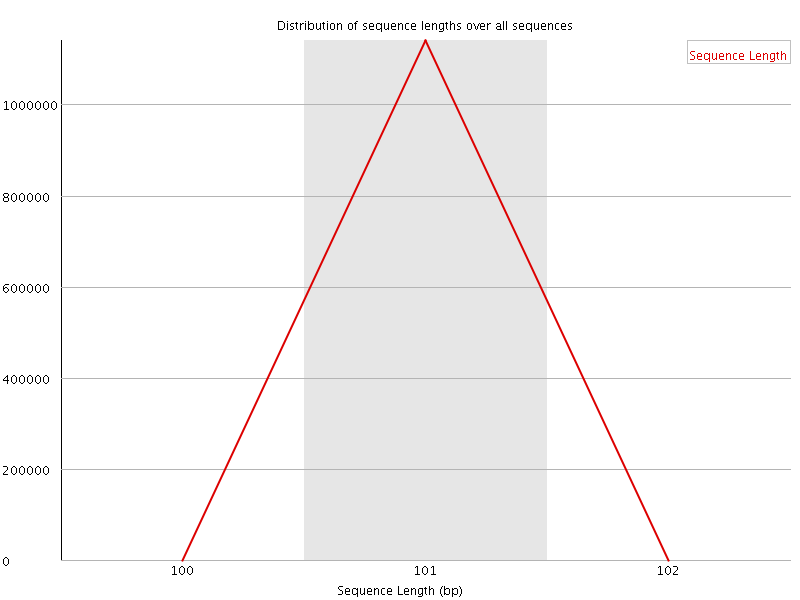
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
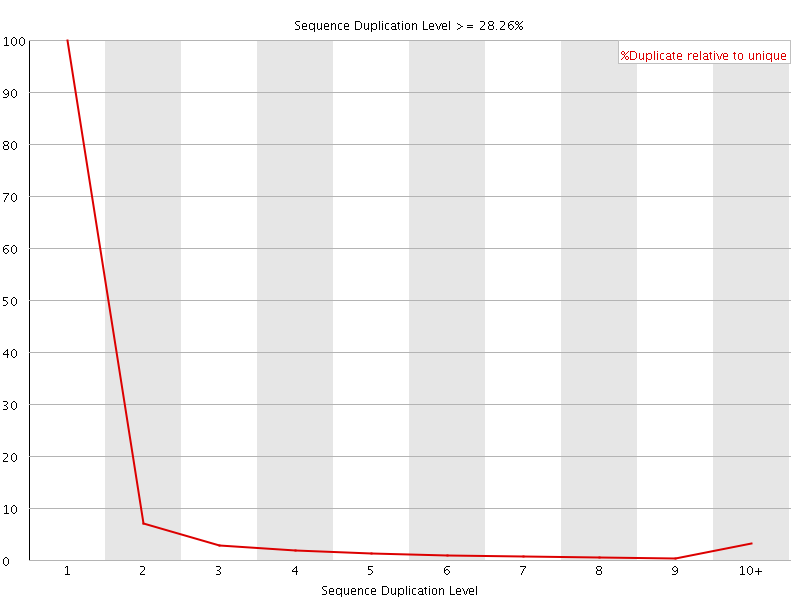
![[WARN]](Icons/warning.png) Overrepresented sequences
Overrepresented sequences
| Sequence | Count | Percentage | Possible Source |
|---|---|---|---|
| CATATGGACTTTGGCTACACCATGAAAGCTTTGAGAAGCAAGAAGAAGGT | 1477 | 0.12953458757660696 | No Hit |
| ATCATACACATGACATCAAGTCATATTCGACTCCAAAACACTAACCAACC | 1188 | 0.10418895737373667 | No Hit |
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
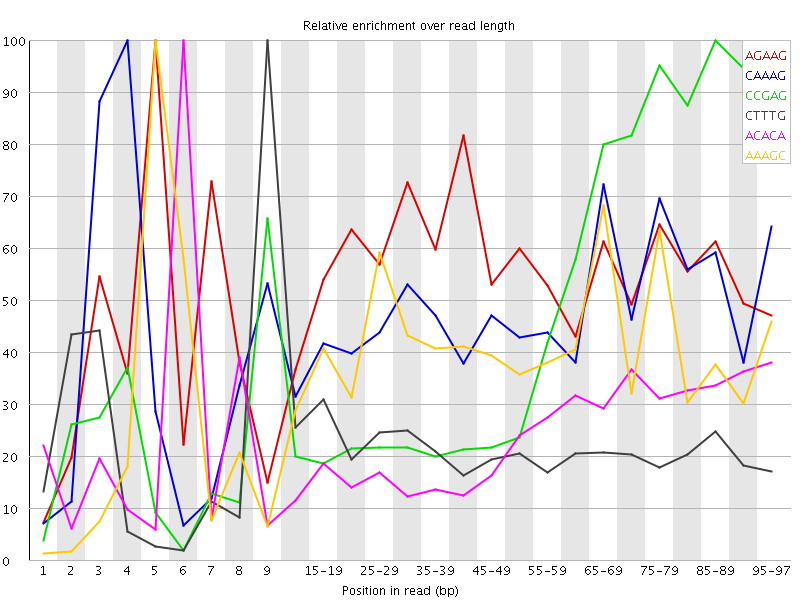
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| AGAAG | 315710 | 2.8338761 | 5.091824 | 5 |
| CAAAG | 289185 | 2.4792428 | 5.24563 | 4 |
| CCGAG | 133500 | 2.4189072 | 5.0485153 | 85-89 |
| CTTTG | 280985 | 2.3029613 | 10.609863 | 9 |
| ACACA | 275345 | 2.2546089 | 9.361228 | 6 |
| AAAGC | 236060 | 2.023791 | 5.066896 | 5 |
| TACAC | 244840 | 1.974982 | 11.717031 | 5 |
| GCTTT | 237035 | 1.9427459 | 5.2699537 | 1 |
| CACAT | 230205 | 1.8569299 | 7.137377 | 7 |
| GCCAA | 145460 | 1.7717791 | 5.422111 | 1 |
| AGAGA | 194385 | 1.7448386 | 5.0091357 | 8 |
| TGGCT | 139205 | 1.7228347 | 6.14446 | 6 |
| TTGGC | 138575 | 1.7150377 | 5.4724097 | 5 |
| TGGAG | 130275 | 1.7136114 | 5.076441 | 5 |
| GAGAG | 123385 | 1.6475055 | 5.865266 | 7 |
| ATACA | 288955 | 1.6405454 | 6.1274276 | 4 |
| TTGAG | 186225 | 1.6221985 | 6.7109637 | 9 |
| GACTC | 133440 | 1.601175 | 6.2888823 | 7 |
| GAGTC | 124640 | 1.5658835 | 8.692025 | 9 |
| ATGGA | 172430 | 1.524727 | 5.5133457 | 4 |
| CATGA | 175250 | 1.4800898 | 6.8177214 | 9 |
| AAGAC | 172575 | 1.4795212 | 6.4302607 | 5 |
| ACGCT | 122055 | 1.4645641 | 5.1737375 | 90-94 |
| GTCTT | 177575 | 1.45541 | 7.0782714 | 1 |
| CCATG | 119805 | 1.4375657 | 6.3063354 | 9 |
| TATGA | 244790 | 1.4334669 | 7.929758 | 4 |
| GACTT | 168425 | 1.4012748 | 8.035282 | 7 |
| TCATA | 249960 | 1.3980262 | 8.347822 | 2 |
| TGAGT | 159760 | 1.3916628 | 6.064785 | 8 |
| ATGAG | 154975 | 1.3703796 | 5.830599 | 7 |
| GTGTT | 156495 | 1.3429295 | 6.2458735 | 1 |
| ACTCA | 165285 | 1.3332582 | 5.846783 | 8 |
| TATAC | 236455 | 1.3224927 | 6.567709 | 5 |
| CGCTA | 109075 | 1.308814 | 5.118096 | 95-97 |
| GATTG | 149950 | 1.3062086 | 7.344473 | 7 |
| GTGTA | 149235 | 1.2999802 | 12.955064 | 1 |
| AACAC | 157230 | 1.2874472 | 5.3555117 | 5 |
| GTTCT | 149505 | 1.2253474 | 5.262005 | 1 |
| GTATG | 139440 | 1.2146562 | 7.4373865 | 3 |
| TGGAC | 96675 | 1.2145522 | 8.685934 | 5 |
| ACATG | 139845 | 1.1810737 | 7.456499 | 8 |
| ACACC | 101065 | 1.1757569 | 5.7081094 | 6 |
| ACCAT | 144375 | 1.1645892 | 5.283614 | 8 |
| GTCCA | 95980 | 1.1516846 | 8.477584 | 1 |
| ATTGA | 194955 | 1.1416378 | 5.3688407 | 8 |
| GGACT | 89715 | 1.1271119 | 8.271737 | 6 |
| ACACT | 138700 | 1.1188123 | 5.4948025 | 6 |
| TATGG | 127210 | 1.1081212 | 5.1187463 | 3 |
| TGTAT | 184585 | 1.0648223 | 6.030999 | 2 |
| AGACT | 124755 | 1.0536296 | 6.309975 | 6 |
| GAGTA | 116385 | 1.0291443 | 5.561334 | 1 |
| TGTAG | 114655 | 0.9987551 | 5.0096855 | 2 |
| CATAC | 120425 | 0.97139853 | 7.2586155 | 3 |
| GAATA | 152210 | 0.9047956 | 5.1164103 | 1 |
| GGATA | 99650 | 0.8811636 | 5.13255 | 1 |
| TATAA | 213410 | 0.8401112 | 5.0222993 | 2 |
| ATACT | 143500 | 0.80259544 | 5.4667635 | 6 |
| GTATA | 136830 | 0.8012635 | 7.6185875 | 1 |