![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-167_GCTACGCT-AAGGAGTA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 3243511 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
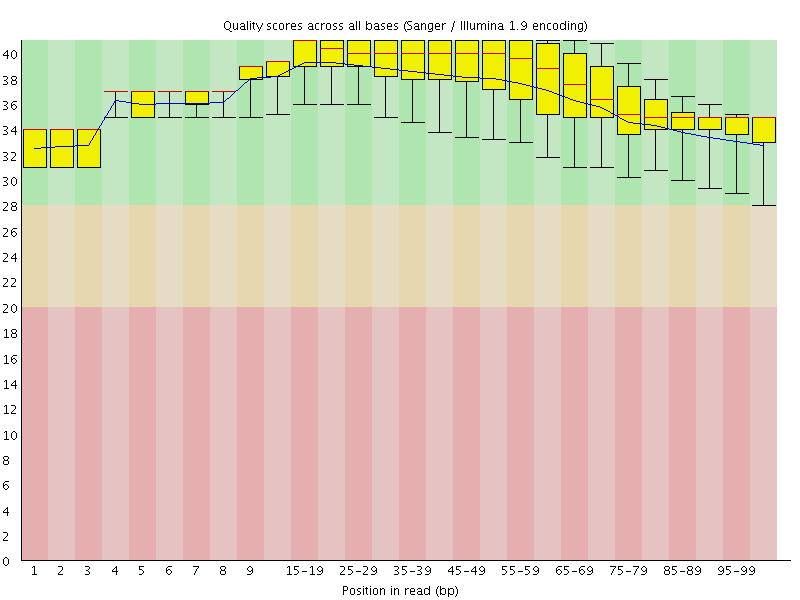
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
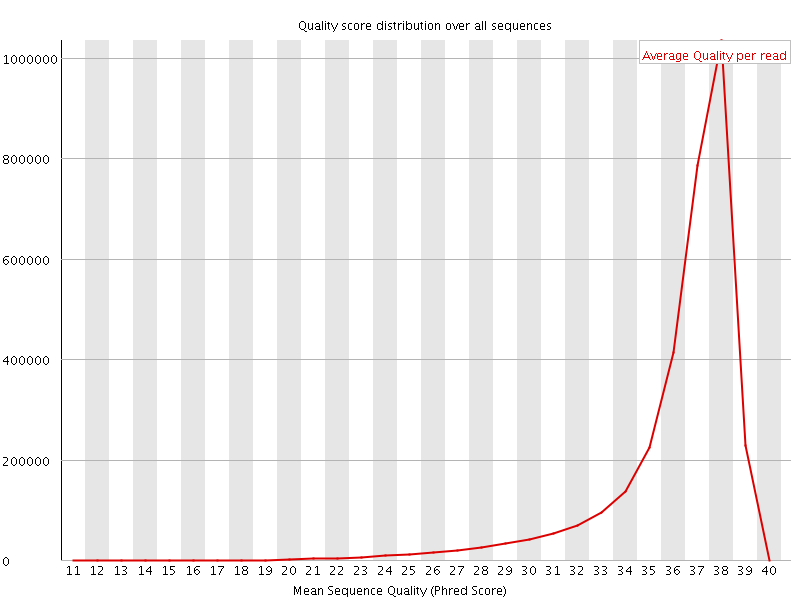
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
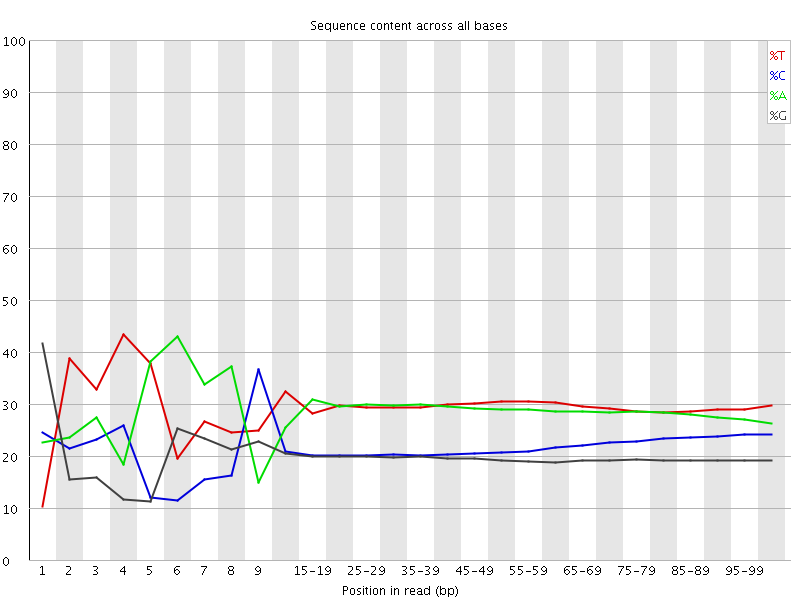
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
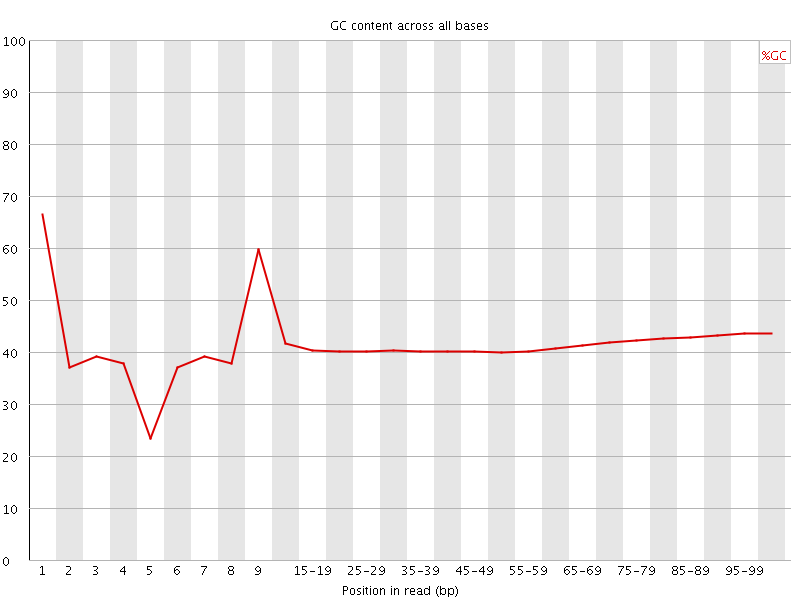
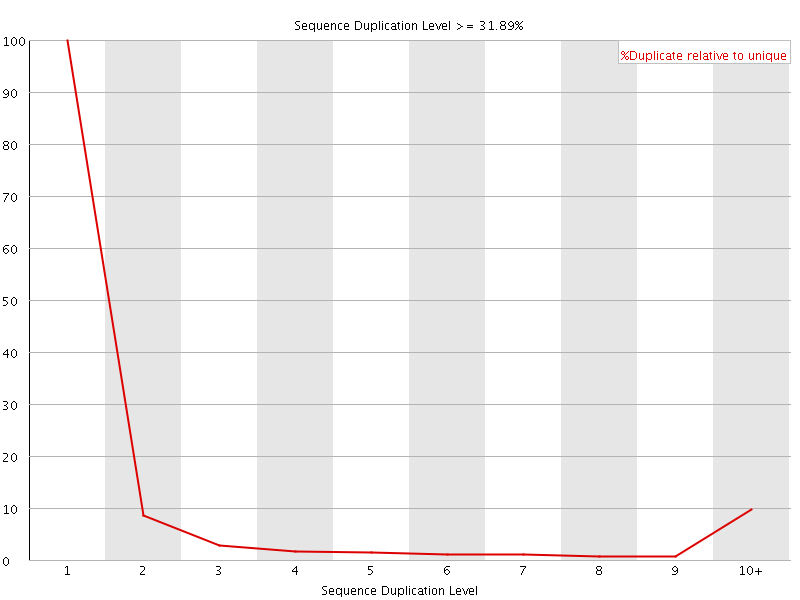
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
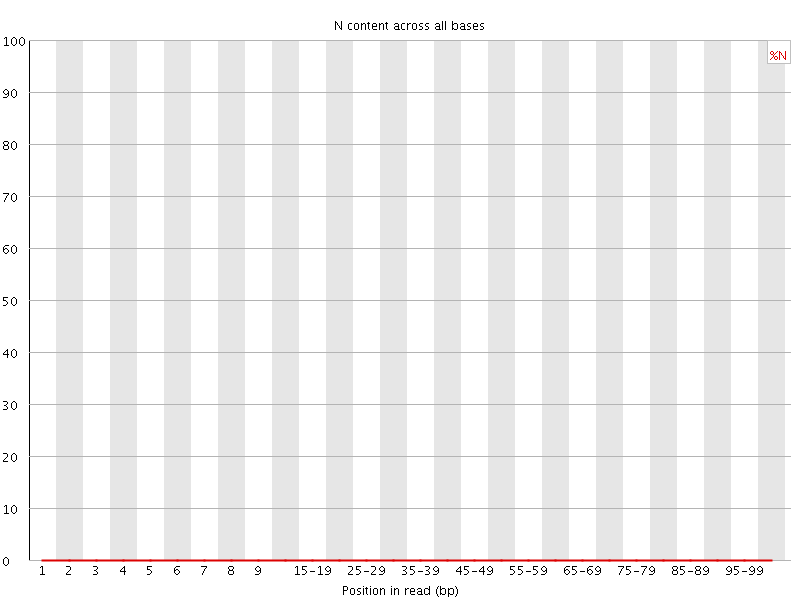
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
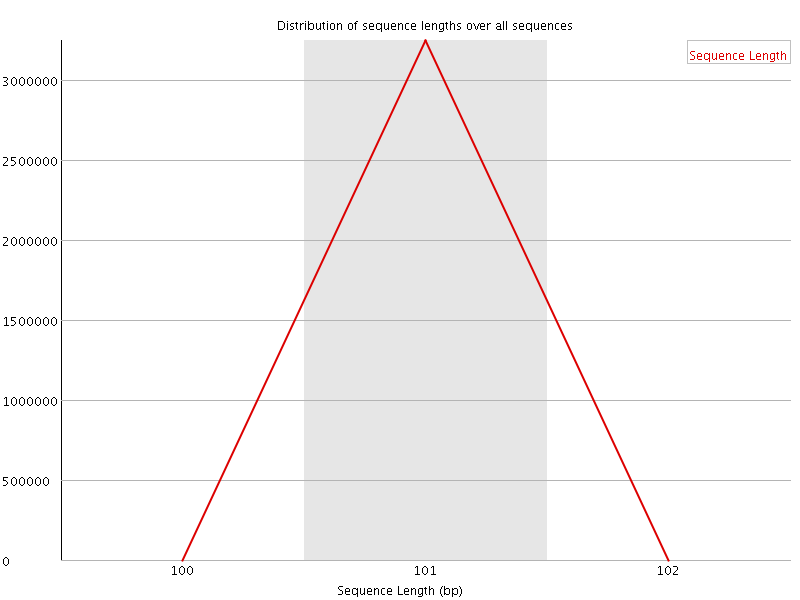
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
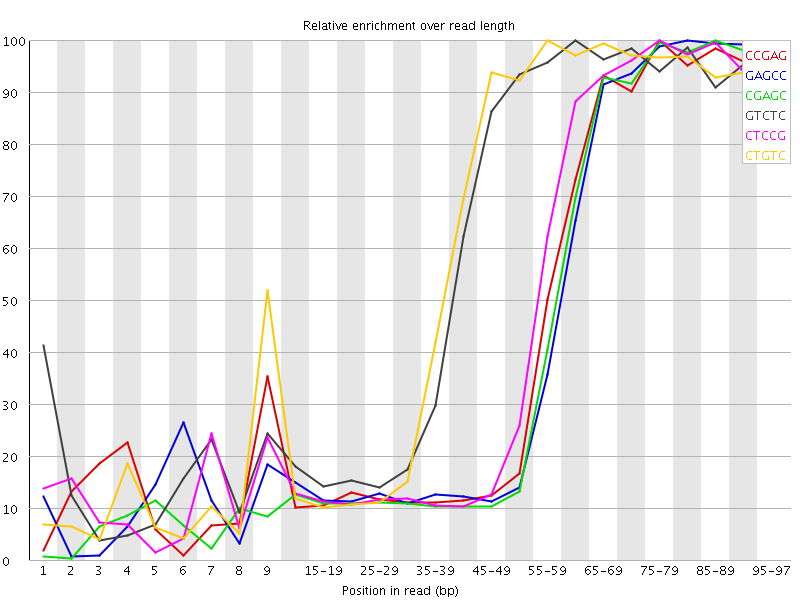
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 730470 | 4.3752728 | 9.579082 | 75-79 |
| GAGCC | 702870 | 4.2099586 | 9.342167 | 80-84 |
| CGAGC | 689155 | 4.12781 | 9.299715 | 85-89 |
| GTCTC | 1027325 | 3.9762633 | 6.4025564 | 60-64 |
| CTCCG | 751850 | 3.9762404 | 8.278916 | 75-79 |
| CTGTC | 988340 | 3.8253717 | 6.1463146 | 55-59 |
| AGCCC | 673605 | 3.6669443 | 8.490706 | 80-84 |
| GCCCA | 670215 | 3.6484897 | 8.61529 | 80-84 |
| TCTCT | 1350055 | 3.4756885 | 5.154711 | 60-64 |
| TGTCT | 1163570 | 3.2959886 | 5.0323334 | 80-84 |
| ATCTC | 1225730 | 3.2481923 | 8.044391 | 95-97 |
| TCTCC | 923300 | 3.2479277 | 6.126683 | 95-97 |
| CTCTT | 1258015 | 3.2387333 | 4.9100223 | 60-64 |
| CCACG | 592605 | 3.2259994 | 8.280924 | 90-94 |
| CCCAC | 637350 | 3.1533573 | 7.7355604 | 80-84 |
| TCCGA | 787685 | 3.1381772 | 6.5102606 | 75-79 |
| ACGCT | 781960 | 3.1153686 | 11.578254 | 95-97 |
| ACACA | 1090185 | 3.0609932 | 5.4766445 | 6 |
| CGAGA | 653945 | 2.9507163 | 7.436745 | 85-89 |
| GAGAC | 644960 | 2.9101744 | 7.4126782 | 85-89 |
| CGCTA | 725585 | 2.8907676 | 11.348175 | 90-94 |
| GACGC | 477695 | 2.861235 | 8.951612 | 90-94 |
| ACGAG | 625850 | 2.8239467 | 7.1271925 | 95-97 |
| TACAC | 1018300 | 2.7776675 | 6.6616054 | 5 |
| AGAAG | 779745 | 2.6504664 | 5.1254554 | 5 |
| CACGA | 612510 | 2.5118613 | 6.4278984 | 95-97 |
| AGACG | 543655 | 2.4530683 | 7.0075917 | 95-97 |
| GCTAC | 547420 | 2.1809492 | 6.6724424 | 90-94 |
| TATAC | 1047300 | 2.090747 | 5.341869 | 5 |
| CAAAG | 676265 | 2.0892134 | 5.046256 | 4 |
| CTTTG | 669640 | 1.8968569 | 8.395566 | 9 |
| AGAGA | 537320 | 1.8264287 | 5.0628295 | 8 |
| GAGAG | 352460 | 1.749846 | 5.9864335 | 7 |
| CTACG | 434870 | 1.7325442 | 6.00838 | 95-97 |
| TCTCG | 436125 | 1.6880227 | 5.7860003 | 90-94 |
| TACGC | 413785 | 1.6485406 | 6.1783743 | 95-97 |
| GCTTT | 573545 | 1.6246531 | 5.2581434 | 1 |
| CTCGT | 377775 | 1.462179 | 5.6504993 | 90-94 |
| GTATG | 440450 | 1.4130293 | 5.0803814 | 95-97 |
| GTCTT | 484890 | 1.3735244 | 5.951812 | 1 |
| GAGTC | 295410 | 1.2949526 | 6.7798758 | 9 |
| AAGAC | 418615 | 1.2932445 | 5.3353405 | 5 |
| TGCCG | 219615 | 1.2779304 | 8.036401 | 95-97 |
| GCCGT | 218320 | 1.2703949 | 7.805043 | 95-97 |
| ATGCC | 309310 | 1.2323068 | 5.8087664 | 95-97 |
| GTGTT | 387395 | 1.2073994 | 6.155718 | 1 |
| TGAGT | 375850 | 1.2057829 | 5.7800636 | 8 |
| CCGTC | 223100 | 1.1798886 | 6.5175695 | 95-97 |
| GATTG | 365305 | 1.171953 | 5.000782 | 7 |
| GACTT | 399050 | 1.1635313 | 6.061888 | 7 |
| GTGTA | 333910 | 1.071233 | 8.462931 | 1 |
| GTTCT | 377925 | 1.0705299 | 5.0713344 | 1 |
| TGGAC | 232665 | 1.0199051 | 6.495079 | 5 |
| GGACT | 215590 | 0.94505554 | 6.588595 | 6 |
| GTCCA | 236855 | 0.9436423 | 6.5782337 | 1 |
| AGACT | 309625 | 0.9292752 | 5.0101156 | 6 |
| GTATA | 333025 | 0.7314945 | 6.9275885 | 1 |