![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-166_GCTACGCT-ACTGCATA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 3466647 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
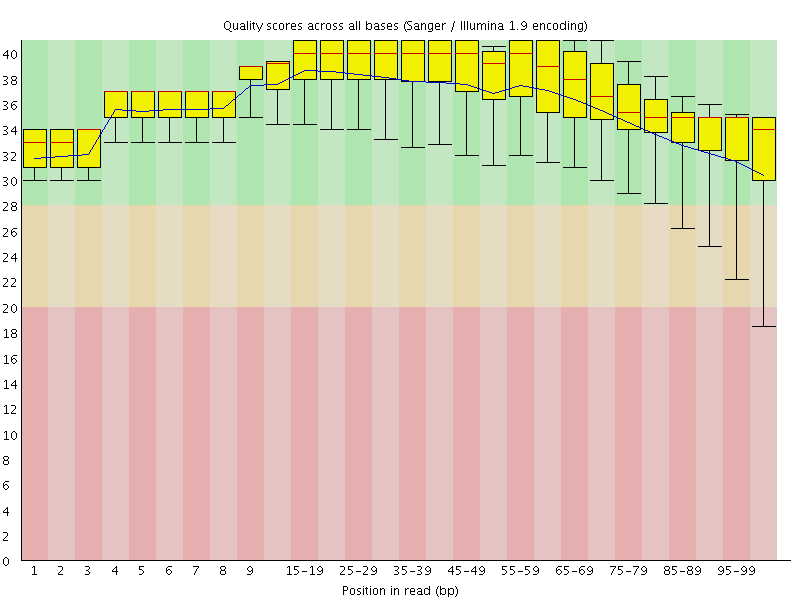
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
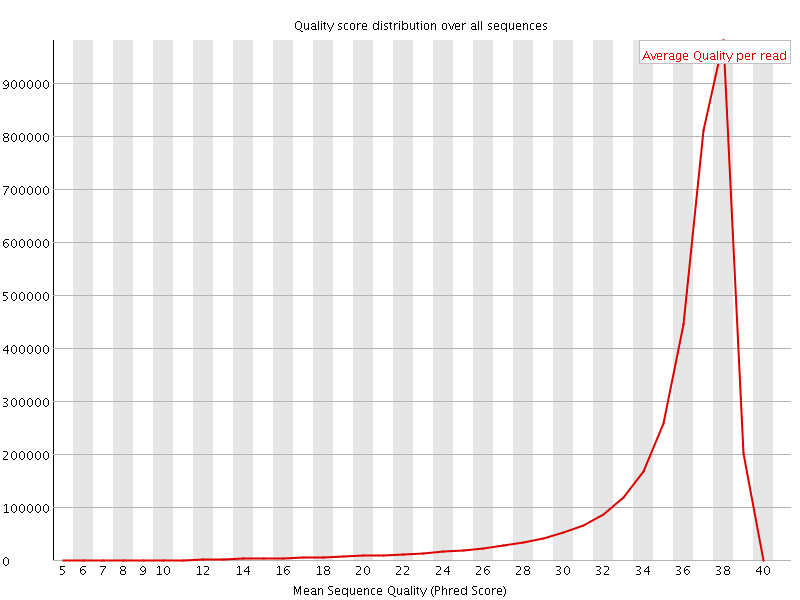
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
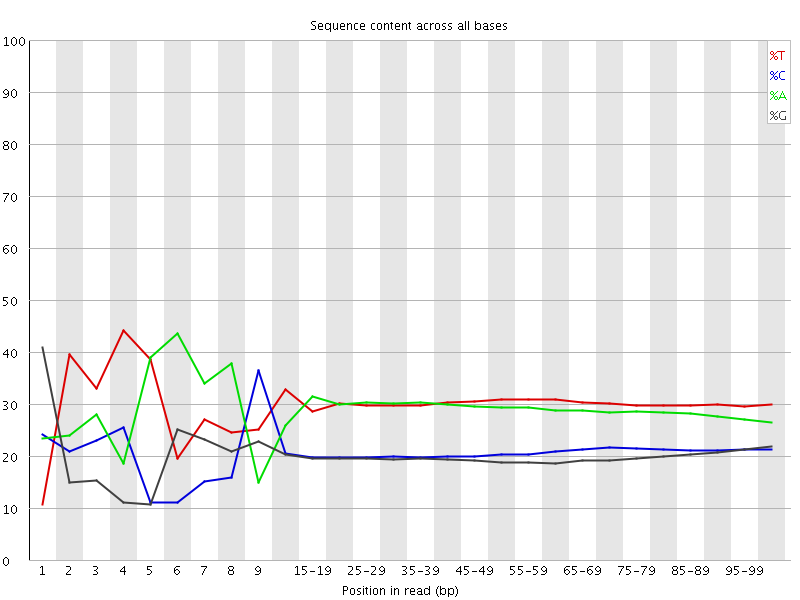
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
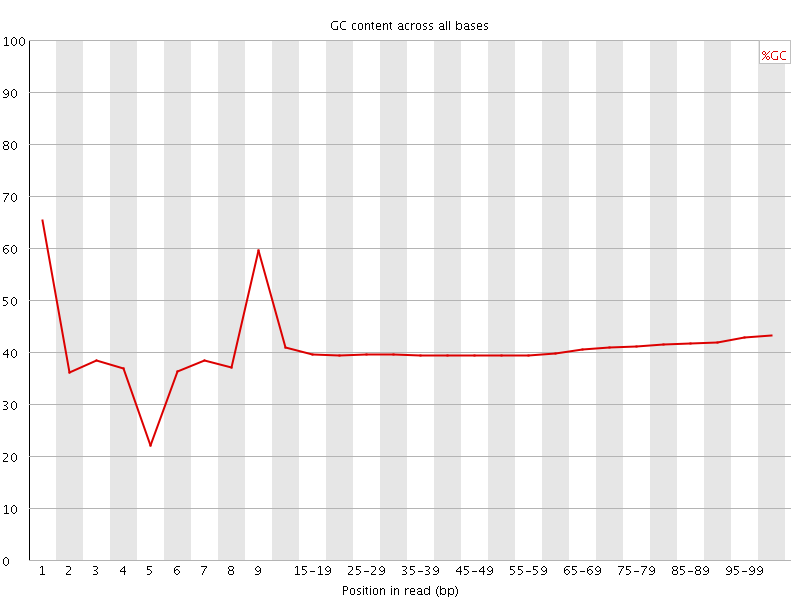
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
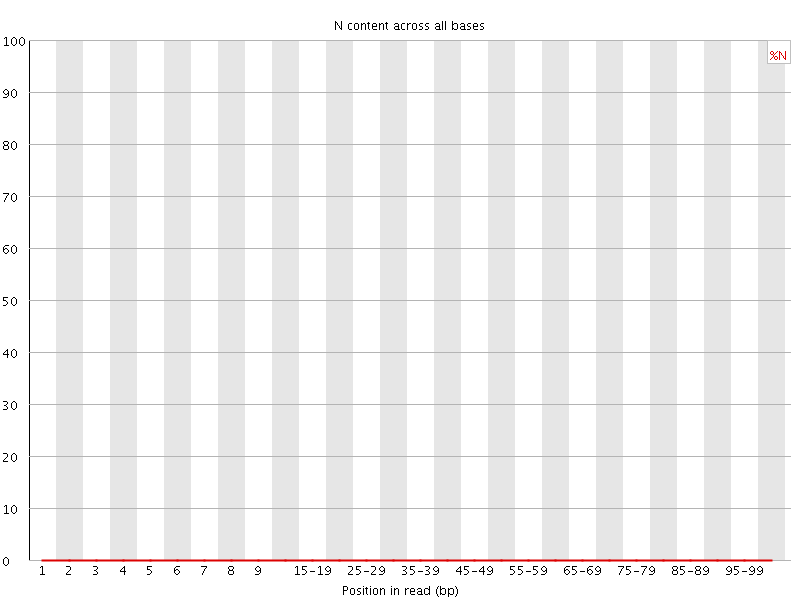
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
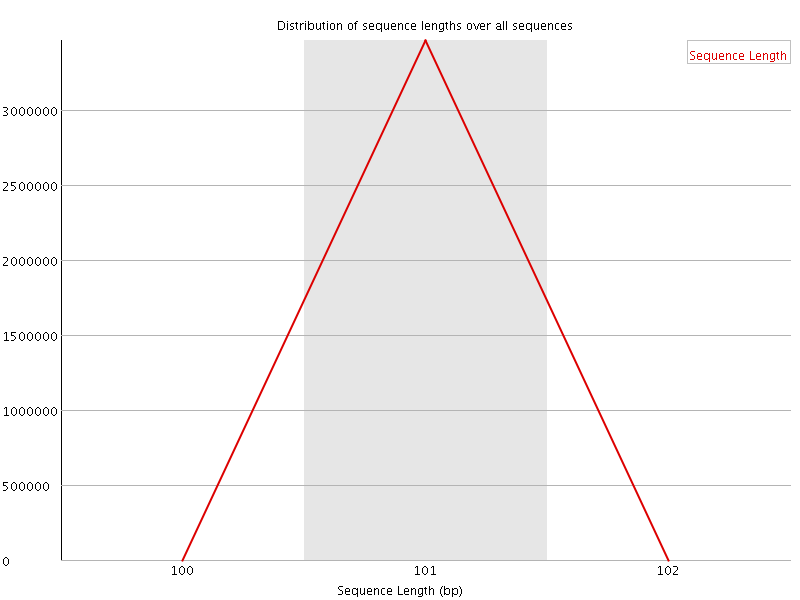
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
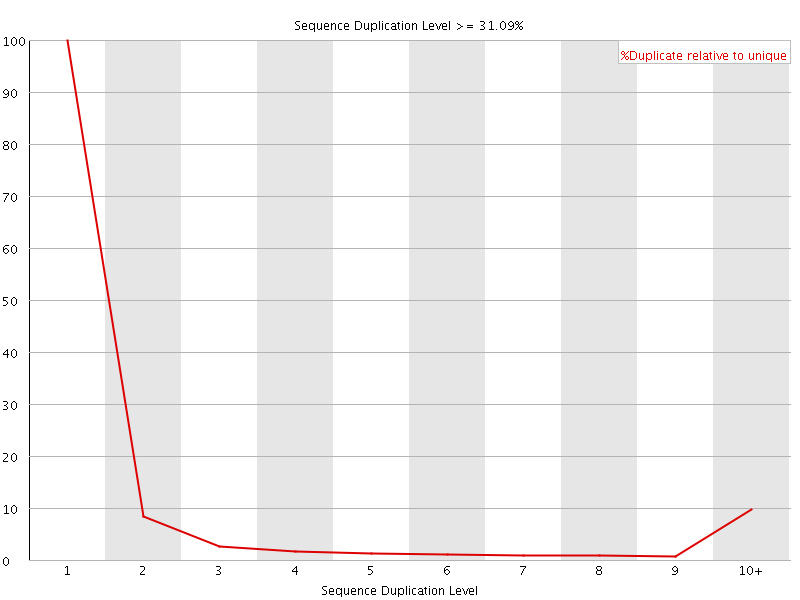
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
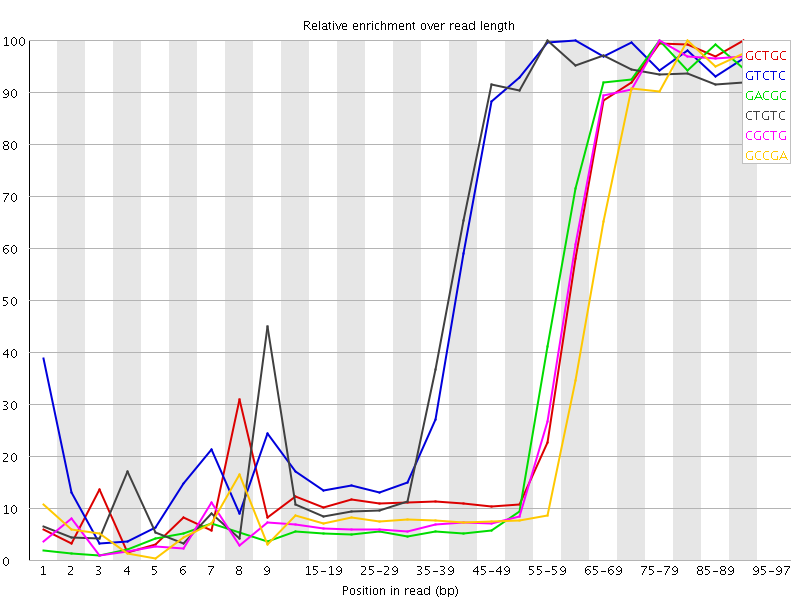
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 784010 | 4.5487175 | 10.626691 | 90-94 |
| GTCTC | 1176740 | 4.483311 | 7.233358 | 60-64 |
| GACGC | 737505 | 4.441986 | 10.74403 | 75-79 |
| CTGTC | 1126625 | 4.292376 | 7.143454 | 55-59 |
| CGCTG | 734485 | 4.2613797 | 10.552174 | 75-79 |
| GCCGA | 681730 | 4.106054 | 11.0265255 | 80-84 |
| CGACG | 665640 | 4.009144 | 11.034677 | 95-97 |
| CTGCC | 718105 | 3.9980118 | 10.209826 | 80-84 |
| TGCCG | 669235 | 3.8828082 | 10.510804 | 80-84 |
| TCTCT | 1523645 | 3.8120024 | 5.803444 | 60-64 |
| CCGAC | 653890 | 3.7792506 | 10.440272 | 80-84 |
| CTCTT | 1429365 | 3.5761235 | 5.5333414 | 60-64 |
| TGTCT | 1325650 | 3.4562845 | 5.3107758 | 55-59 |
| ACACA | 1228540 | 3.4386928 | 6.2885995 | 6 |
| TGACG | 798955 | 3.2930381 | 7.483942 | 75-79 |
| CTGAC | 830690 | 3.285506 | 7.1107416 | 85-89 |
| TACAC | 1145885 | 3.0895855 | 7.4981475 | 5 |
| GACGA | 717385 | 3.069528 | 8.077292 | 85-89 |
| CACAT | 1122160 | 3.0256171 | 5.3700147 | 95-97 |
| ATCTG | 1106985 | 2.9961755 | 5.7651486 | 70-74 |
| ACGCT | 743310 | 2.9399045 | 7.21341 | 75-79 |
| CATCT | 1128335 | 2.9305716 | 5.4125443 | 70-74 |
| ACATC | 1078065 | 2.9067266 | 5.3907666 | 70-74 |
| TCTGA | 1038085 | 2.8096905 | 5.404926 | 70-74 |
| AGAAG | 833660 | 2.534052 | 5.2299867 | 5 |
| TGCAG | 609375 | 2.51165 | 7.715187 | 90-94 |
| TATAC | 1189435 | 2.1946342 | 5.4363713 | 5 |
| GCAGT | 508000 | 2.0938144 | 7.222243 | 90-94 |
| GTGTG | 496155 | 2.0528548 | 7.40039 | 95-97 |
| CAAAG | 702230 | 2.0483053 | 5.172903 | 4 |
| ATGCA | 724405 | 2.0354097 | 5.483775 | 90-94 |
| CAGTG | 492260 | 2.028939 | 7.184799 | 95-97 |
| ACGAT | 691100 | 1.9418304 | 5.279907 | 85-89 |
| GGTGG | 303445 | 1.9119202 | 8.358132 | 95-97 |
| CTTTG | 695895 | 1.8143637 | 7.7721543 | 9 |
| AGAGA | 595550 | 1.8102759 | 5.1358194 | 8 |
| AGTGT | 625865 | 1.7652956 | 5.1784883 | 95-97 |
| CGATA | 622935 | 1.7503026 | 5.1602626 | 85-89 |
| GAGAG | 380440 | 1.6963552 | 5.66541 | 7 |
| TATGC | 623650 | 1.6879766 | 5.0566626 | 90-94 |
| GTGTA | 593650 | 1.674431 | 8.82911 | 1 |
| CTCGG | 282835 | 1.6409693 | 8.10911 | 95-97 |
| CGGTG | 227750 | 1.3770096 | 7.706658 | 95-97 |
| TCTCG | 356960 | 1.3599969 | 5.4039 | 95-97 |
| AAGAC | 445180 | 1.2985269 | 5.740281 | 5 |
| GAGTC | 305610 | 1.2596272 | 6.232298 | 9 |
| CGCCG | 147070 | 1.2468929 | 5.3814116 | 95-97 |
| TATGA | 611425 | 1.175644 | 5.3640018 | 4 |
| GTCTT | 439040 | 1.1446816 | 5.930337 | 1 |
| GACTT | 417740 | 1.1306589 | 5.5285773 | 7 |
| GATTG | 400155 | 1.1286649 | 5.163607 | 7 |
| TCATA | 609025 | 1.123716 | 5.3396626 | 2 |
| GTGTT | 399975 | 1.0867378 | 5.4182353 | 1 |
| TGGAC | 248025 | 1.0222802 | 6.6218247 | 5 |
| TCGGT | 254880 | 1.0119648 | 5.15321 | 95-97 |
| GTCCA | 245080 | 0.9693288 | 6.8666267 | 1 |
| GGACT | 226815 | 0.9348592 | 6.080247 | 6 |
| AGACT | 327660 | 0.9206485 | 5.0414996 | 6 |
| GTATA | 359885 | 0.6919846 | 6.931973 | 1 |