![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-166_GCTACGCT-ACTGCATA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 3466647 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
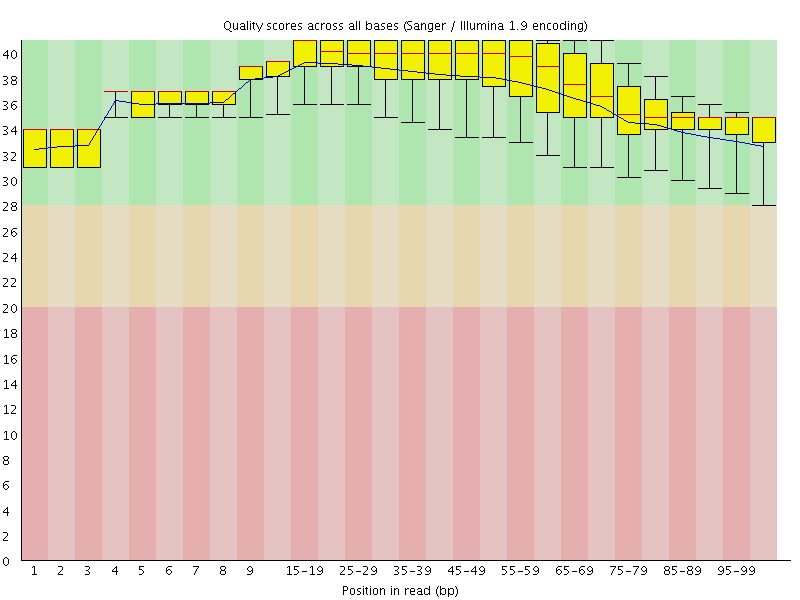
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
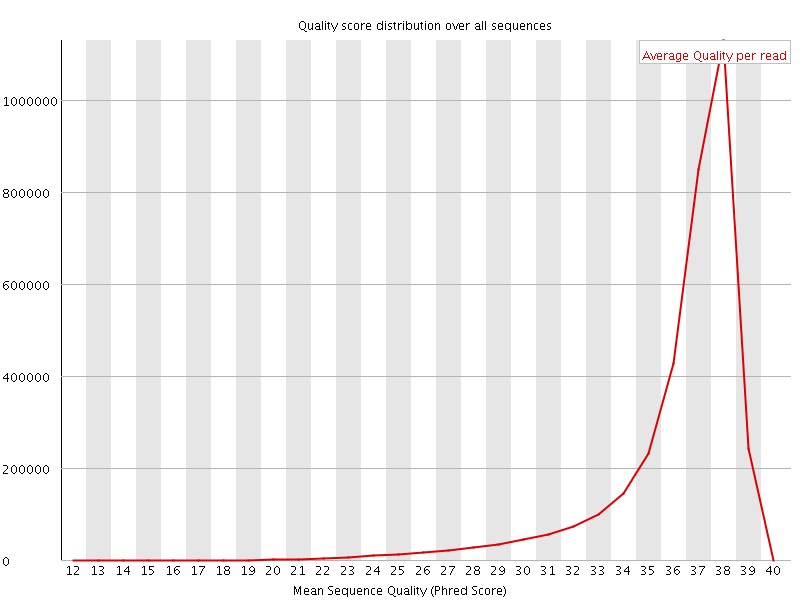
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
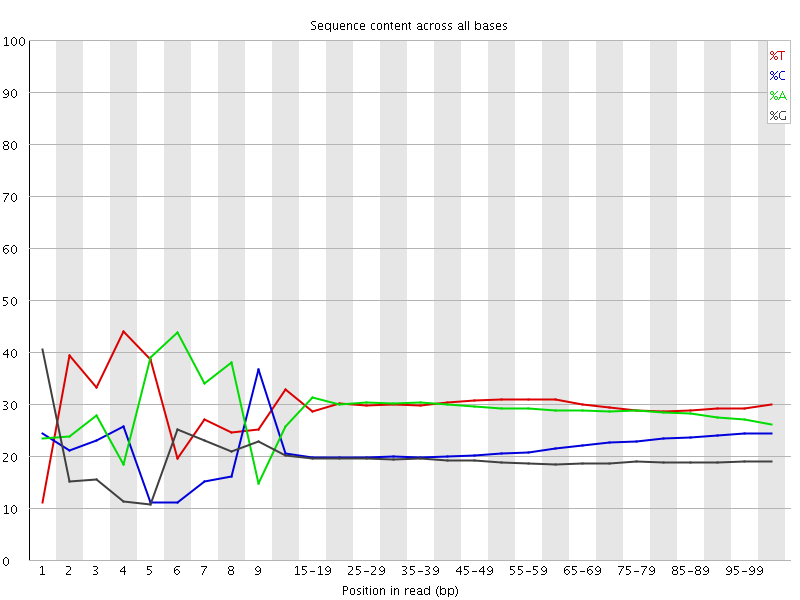
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
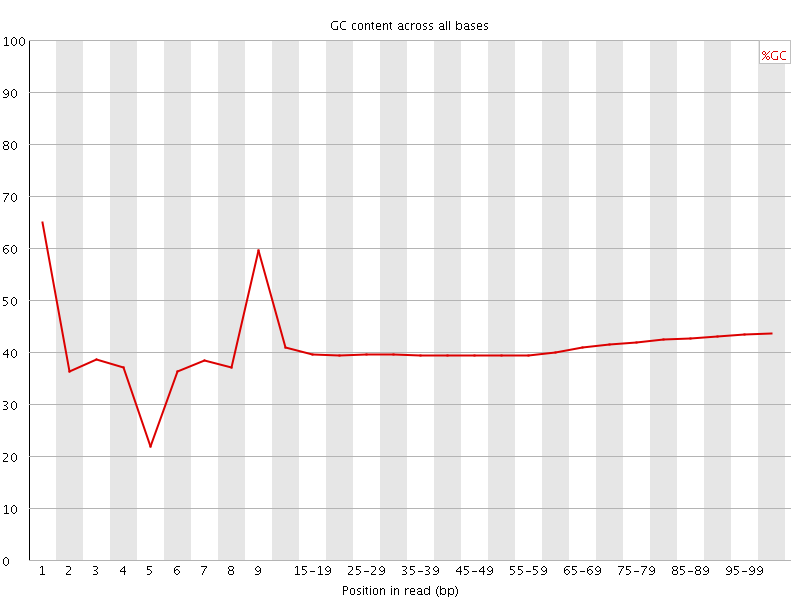
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
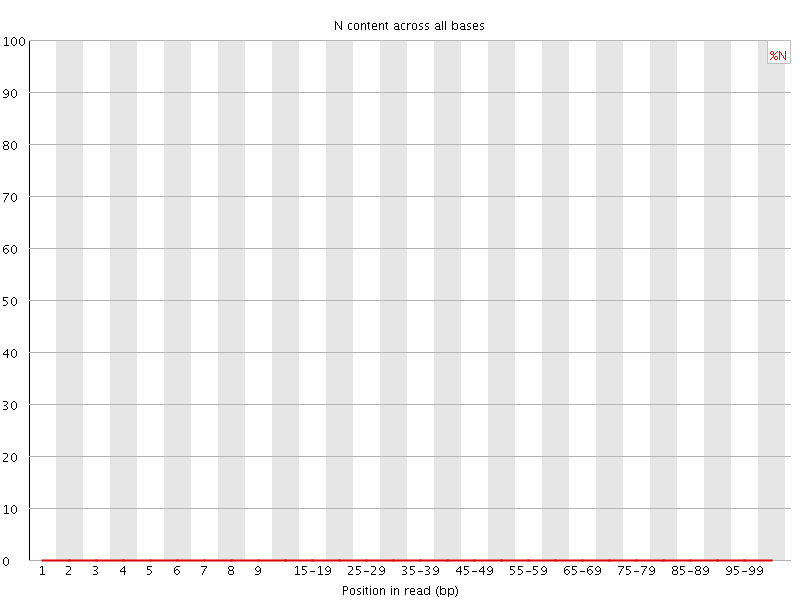
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
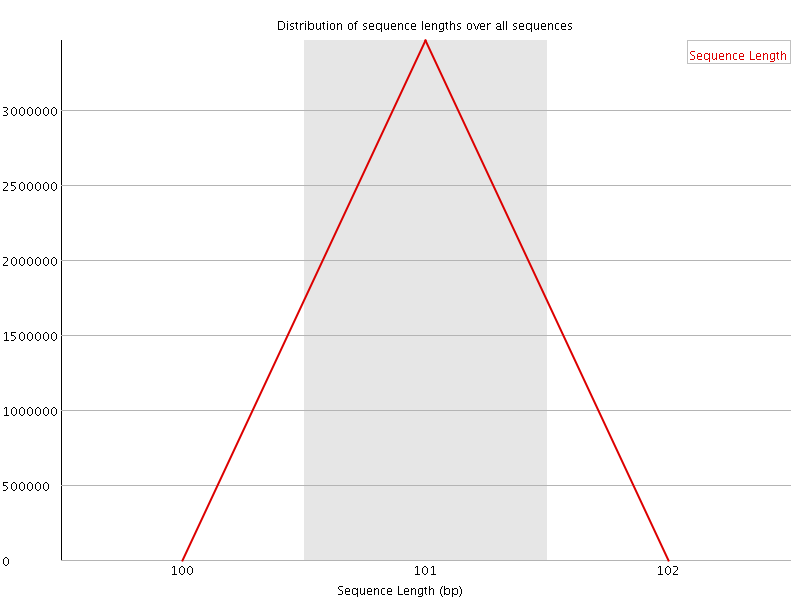
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
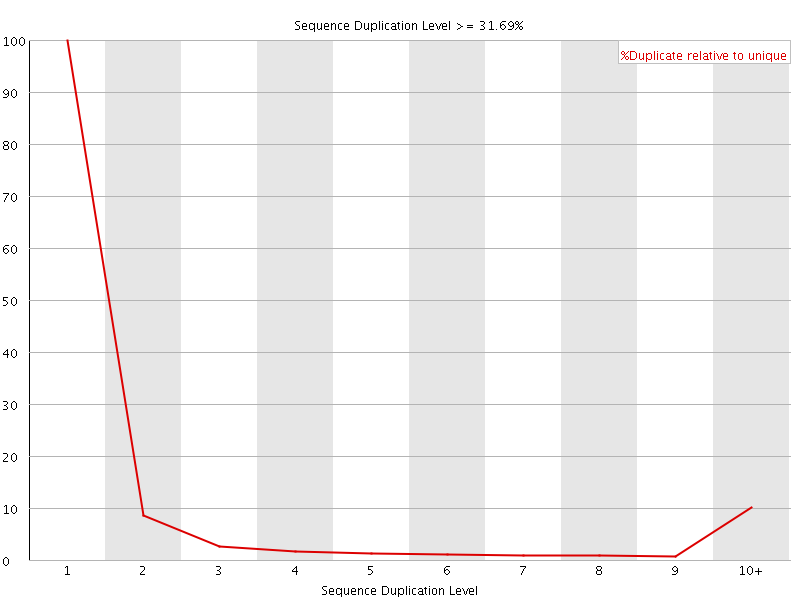
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
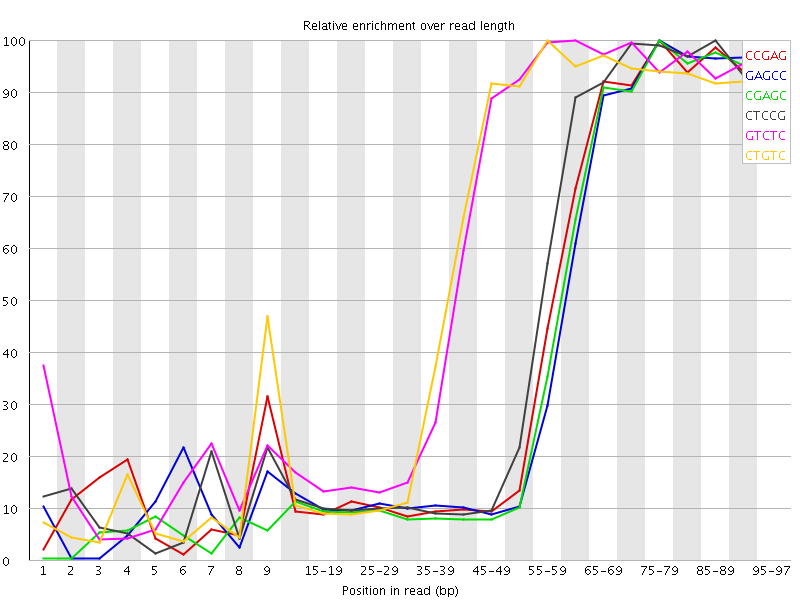
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 820195 | 4.8227 | 10.955868 | 75-79 |
| GAGCC | 790980 | 4.6509175 | 10.90683 | 75-79 |
| CGAGC | 773375 | 4.5474005 | 10.794499 | 75-79 |
| CTCCG | 848535 | 4.3569307 | 9.324293 | 85-89 |
| GTCTC | 1179560 | 4.3438044 | 7.029602 | 60-64 |
| CTGTC | 1129430 | 4.1591973 | 6.908893 | 55-59 |
| AGCCC | 754315 | 3.993555 | 9.51093 | 90-94 |
| GCCCA | 746095 | 3.950036 | 9.736888 | 80-84 |
| TCTCT | 1534925 | 3.6501486 | 5.5450444 | 60-64 |
| CCACG | 668740 | 3.540497 | 9.548466 | 80-84 |
| TGTCT | 1331040 | 3.5154474 | 5.3860545 | 55-59 |
| TCTCC | 1047945 | 3.4747417 | 6.6623263 | 70-74 |
| ATCTC | 1397035 | 3.4255216 | 8.723257 | 95-97 |
| CCCAC | 718315 | 3.4241734 | 8.774457 | 80-84 |
| CTCTT | 1431660 | 3.404578 | 5.26456 | 60-64 |
| ACGCT | 889945 | 3.3791652 | 13.001716 | 90-94 |
| TCCGA | 884335 | 3.3578637 | 7.1684203 | 75-79 |
| ACACA | 1233330 | 3.2150724 | 6.119948 | 6 |
| CGAGA | 732980 | 3.1871357 | 8.336779 | 85-89 |
| GAGAC | 730555 | 3.1765919 | 8.421961 | 85-89 |
| CGCTA | 829860 | 3.1510196 | 12.812442 | 90-94 |
| GACGC | 535200 | 3.1469457 | 10.435359 | 85-89 |
| ACGAG | 704065 | 3.0614083 | 8.042069 | 95-97 |
| TACAC | 1151710 | 2.9117804 | 7.299762 | 5 |
| CACAT | 1126090 | 2.8470073 | 5.022934 | 95-97 |
| CATCT | 1135670 | 2.784656 | 5.1184163 | 70-74 |
| ACATC | 1081850 | 2.7351587 | 5.0397153 | 70-74 |
| CACGA | 688885 | 2.69705 | 7.161276 | 80-84 |
| AGACG | 615325 | 2.6755497 | 8.012079 | 85-89 |
| AGAAG | 825690 | 2.6549776 | 5.4675283 | 5 |
| GCTAC | 612385 | 2.3252559 | 7.3960414 | 90-94 |
| TATAC | 1193725 | 2.1645074 | 5.4305506 | 5 |
| AGAGA | 587740 | 1.8898574 | 5.3069477 | 8 |
| CTACG | 493720 | 1.8746793 | 6.8444233 | 90-94 |
| CTTTG | 696895 | 1.8405893 | 8.177677 | 9 |
| GAGAG | 378955 | 1.8300472 | 6.2844696 | 7 |
| TACGC | 468065 | 1.7772661 | 6.7328477 | 90-94 |
| TCTCG | 475180 | 1.7498804 | 6.337348 | 90-94 |
| GCTTT | 590835 | 1.560471 | 5.092114 | 1 |
| CTCGT | 411805 | 1.516498 | 6.2352076 | 90-94 |
| AGGAG | 301210 | 1.4546015 | 5.003693 | 7 |
| GTATG | 480860 | 1.454355 | 5.3899417 | 90-94 |
| GCTAT | 515845 | 1.4047686 | 5.0582533 | 95-97 |
| GTCTT | 511440 | 1.3507787 | 5.82212 | 1 |
| AAGAC | 444520 | 1.2869716 | 5.686583 | 5 |
| TGCCG | 225585 | 1.2864335 | 9.355405 | 95-97 |
| GAGTC | 304685 | 1.2848828 | 6.4611673 | 9 |
| GCCGT | 223035 | 1.2718916 | 8.814358 | 95-97 |
| ATGCC | 328270 | 1.2464576 | 6.4507437 | 95-97 |
| TATGA | 606385 | 1.2211511 | 5.5919027 | 4 |
| GATTG | 398250 | 1.2045019 | 5.7205677 | 7 |
| CCATG | 315320 | 1.1972857 | 5.0314984 | 9 |
| CCGTC | 228810 | 1.1748594 | 7.318966 | 95-97 |
| GTGTT | 394770 | 1.1579767 | 5.793384 | 1 |
| GACTT | 416335 | 1.1337793 | 5.853219 | 7 |
| TCATA | 611860 | 1.1094477 | 5.323915 | 2 |
| GTGTA | 351005 | 1.0616101 | 9.097367 | 1 |
| TGGAC | 244650 | 1.0317099 | 7.039809 | 5 |
| GGACT | 226220 | 0.953989 | 6.522507 | 6 |
| GTCCA | 249310 | 0.9466424 | 6.512437 | 1 |
| GAGTA | 300095 | 0.9358507 | 5.2745633 | 1 |
| AGACT | 327695 | 0.92013466 | 5.162516 | 6 |
| GTATA | 360590 | 0.72616386 | 7.016329 | 1 |