![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-165_GCTACGCT-GTAAGGAG_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2604350 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
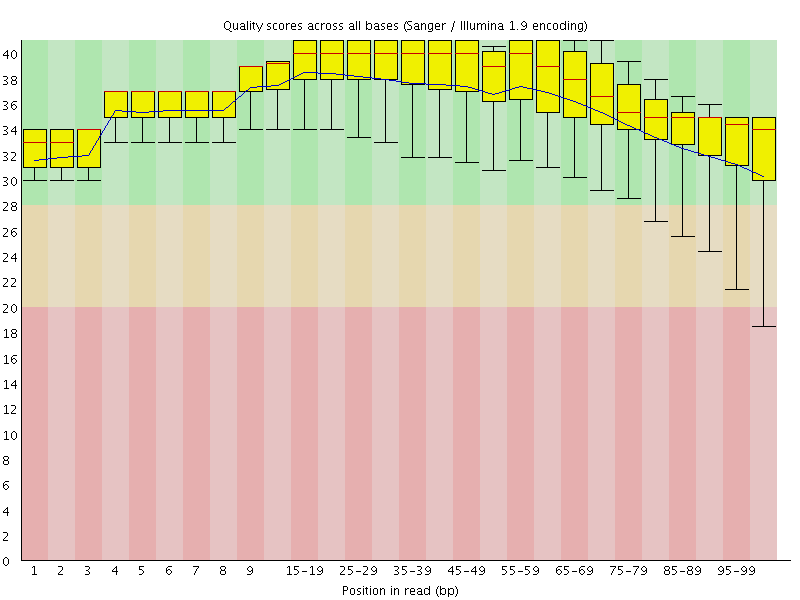
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
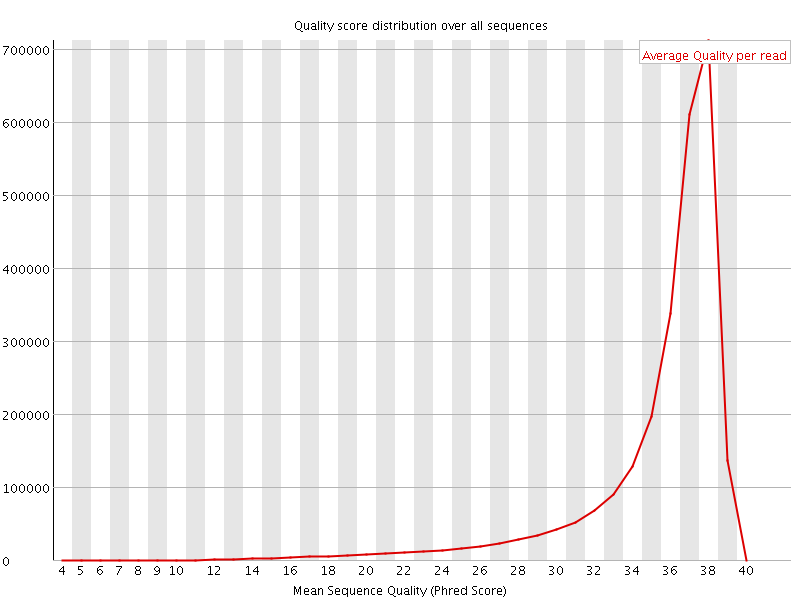
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
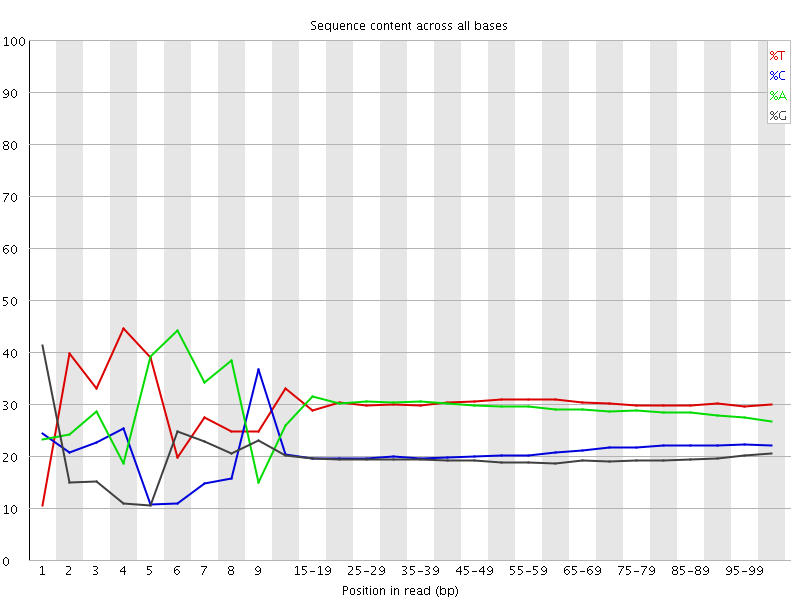
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
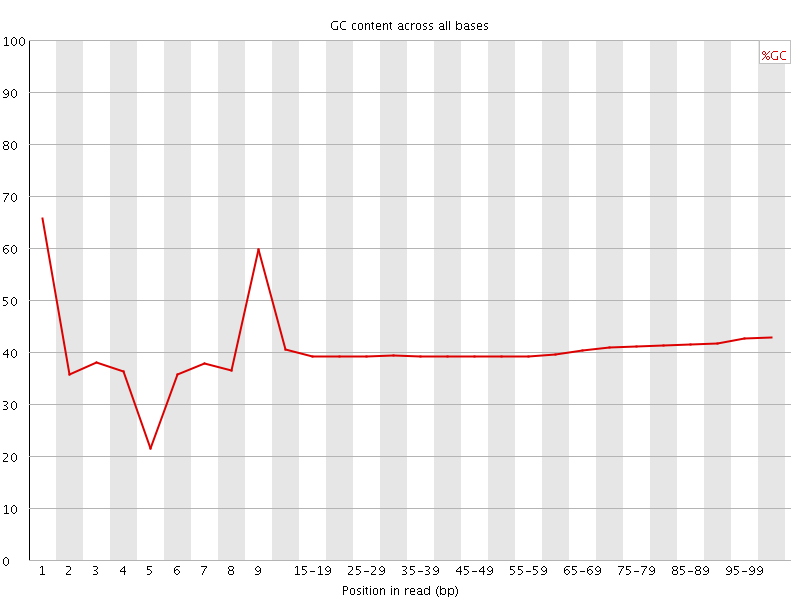
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
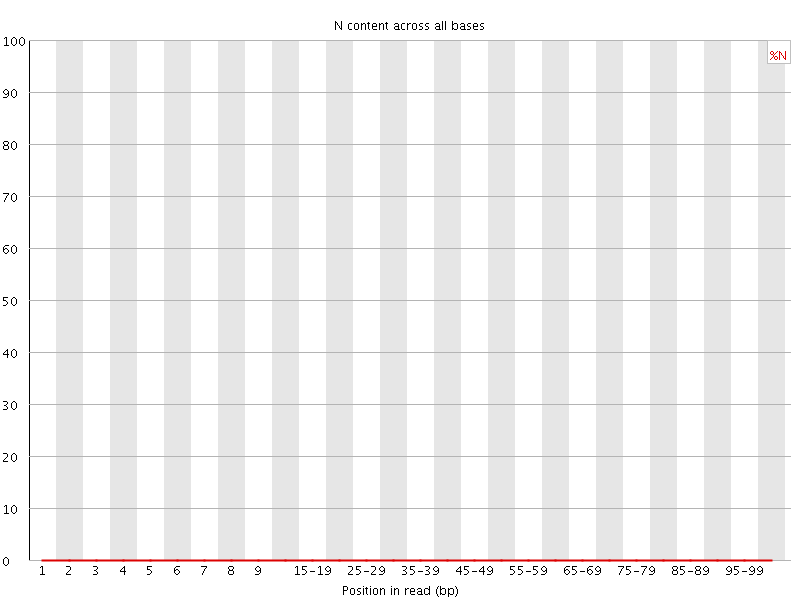
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
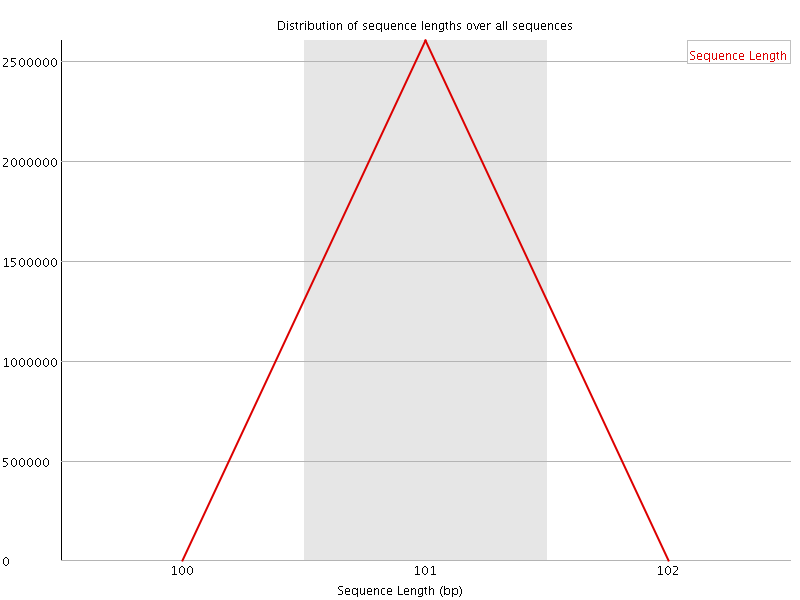
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
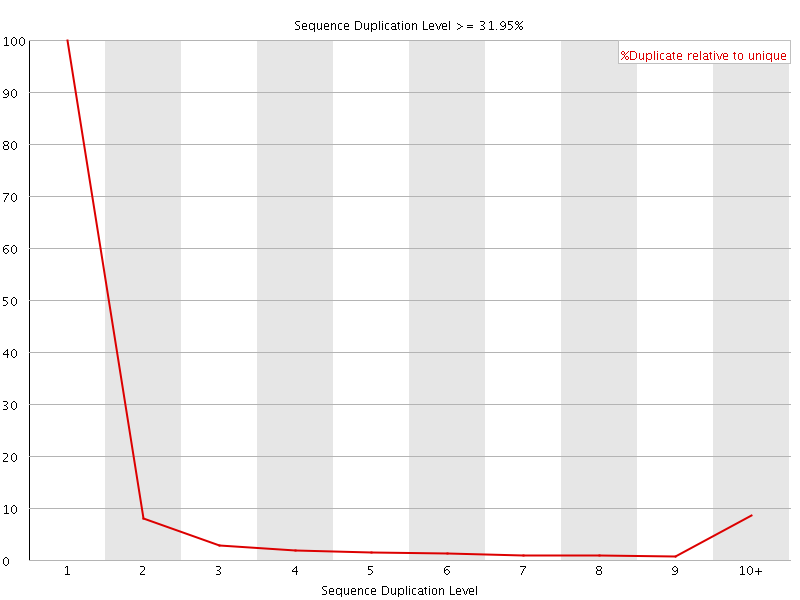
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
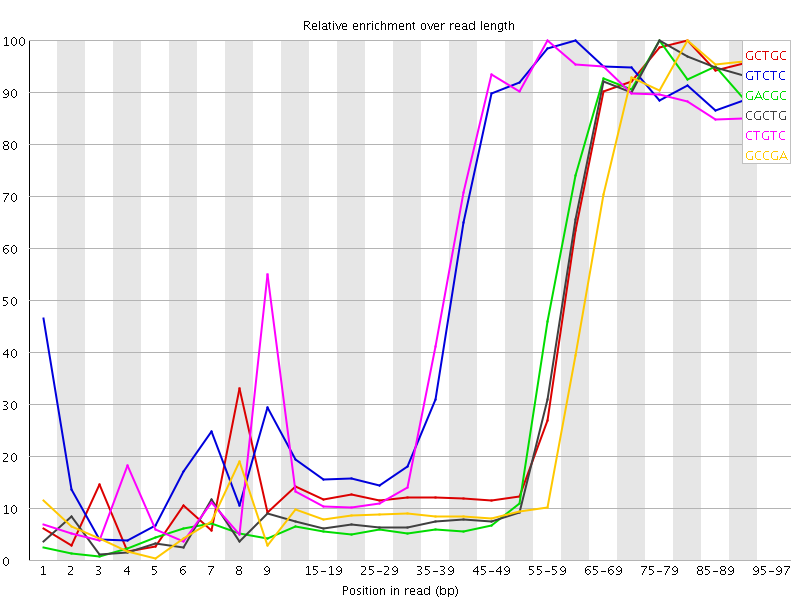
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 506150 | 3.984031 | 9.169775 | 80-84 |
| GTCTC | 751905 | 3.821265 | 6.2383623 | 60-64 |
| GACGC | 468020 | 3.812302 | 9.249924 | 75-79 |
| CGCTG | 463230 | 3.6461976 | 8.933798 | 75-79 |
| CTGTC | 705690 | 3.5863953 | 6.0241995 | 55-59 |
| GCCGA | 436935 | 3.5590963 | 9.297354 | 80-84 |
| CGACG | 422880 | 3.4446099 | 9.168203 | 85-89 |
| CTGCC | 461955 | 3.4210925 | 8.536162 | 80-84 |
| TCTCT | 1026590 | 3.3685412 | 5.0141807 | 60-64 |
| TGCCG | 421795 | 3.3200524 | 8.854393 | 80-84 |
| CCGAC | 421795 | 3.2325552 | 8.603198 | 80-84 |
| CTCTT | 955745 | 3.136078 | 4.769105 | 60-64 |
| ACACA | 834075 | 3.0331101 | 6.46671 | 6 |
| TGTCT | 867310 | 3.024806 | 4.581444 | 60-64 |
| TGACG | 511115 | 2.8570678 | 6.44531 | 75-79 |
| CTGAC | 536360 | 2.8208497 | 6.03858 | 75-79 |
| GACTC | 519605 | 2.732731 | 6.909142 | 85-89 |
| GACGA | 466885 | 2.7007928 | 6.8144765 | 85-89 |
| TACAC | 766355 | 2.6929834 | 7.592607 | 5 |
| AGAAG | 648285 | 2.6632123 | 5.207344 | 5 |
| ACGCT | 473430 | 2.4898853 | 6.016134 | 75-79 |
| CTCCT | 495980 | 2.3715372 | 5.762571 | 90-94 |
| ACTCC | 475765 | 2.3541694 | 5.708563 | 85-89 |
| CGACT | 428335 | 2.2527196 | 5.842469 | 85-89 |
| ATACA | 882920 | 2.2033496 | 5.1057673 | 6 |
| ACGAC | 396500 | 2.1579738 | 6.131628 | 85-89 |
| CAAAG | 555295 | 2.146274 | 5.028718 | 4 |
| GGTGG | 216820 | 1.9279668 | 7.3863387 | 95-97 |
| CTTTG | 548460 | 1.912794 | 8.483632 | 9 |
| TATAC | 790345 | 1.9058969 | 5.8666563 | 5 |
| AGAGA | 458050 | 1.88171 | 5.1579404 | 8 |
| GAGAG | 291460 | 1.7920027 | 5.838014 | 7 |
| GTGTA | 428145 | 1.6423732 | 9.379007 | 1 |
| GCTTT | 462395 | 1.6126359 | 5.0756283 | 1 |
| CTCGG | 195770 | 1.540954 | 7.05762 | 95-97 |
| ACGTG | 270930 | 1.5144643 | 5.8489156 | 95-97 |
| CGTGT | 257345 | 1.3900751 | 5.459795 | 95-97 |
| AAGAC | 350650 | 1.3552995 | 5.9155874 | 5 |
| GAGTC | 239880 | 1.3408988 | 7.11323 | 9 |
| CGGTG | 156205 | 1.3068231 | 6.6356626 | 95-97 |
| CCATG | 242455 | 1.2751307 | 5.134149 | 9 |
| ATGAG | 313920 | 1.2461759 | 5.0456853 | 7 |
| TATGA | 480010 | 1.2303009 | 5.7841268 | 4 |
| GTCTT | 344805 | 1.2025324 | 6.278553 | 1 |
| GATTG | 311540 | 1.195074 | 5.5100913 | 7 |
| TGAGT | 311390 | 1.1944985 | 5.4135027 | 8 |
| GACTT | 327620 | 1.1824235 | 6.1923175 | 7 |
| TCATA | 483180 | 1.1651763 | 5.4033766 | 2 |
| GTGTT | 307980 | 1.1416265 | 5.718394 | 1 |
| CGCCG | 98655 | 1.1315756 | 5.110678 | 95-97 |
| GTTCT | 321025 | 1.1195978 | 5.1348443 | 1 |
| TGGAC | 193205 | 1.0799915 | 7.1100335 | 5 |
| GTCCA | 191820 | 1.0088289 | 7.284086 | 1 |
| GGACT | 176515 | 0.9866964 | 6.863649 | 6 |
| AGACT | 257520 | 0.96181816 | 5.3539696 | 6 |
| GAGTA | 238190 | 0.94554865 | 5.1995945 | 1 |
| GAATA | 345515 | 0.91644704 | 5.0426764 | 1 |
| GGATA | 221845 | 0.8806635 | 5.1918917 | 1 |
| GTATA | 281160 | 0.7206337 | 7.242731 | 1 |