![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-163_GCTACGCT-TATCCTCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1538541 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
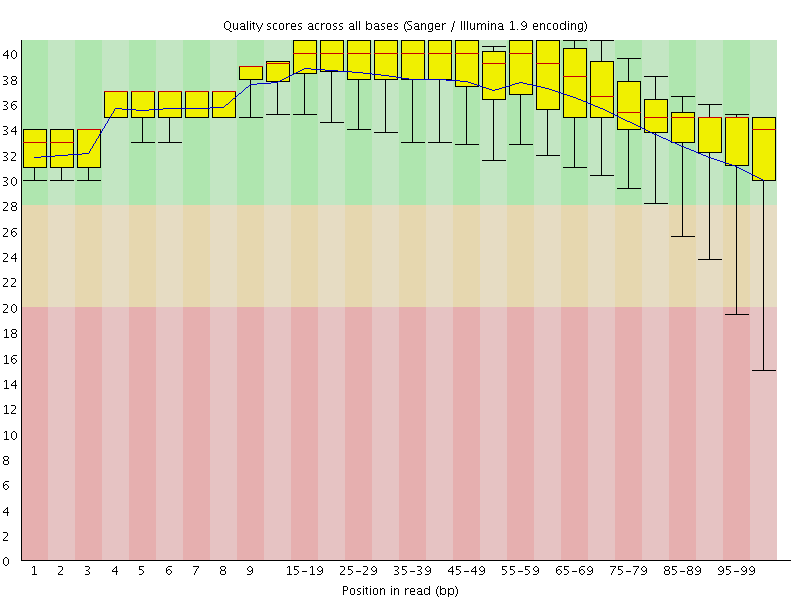
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
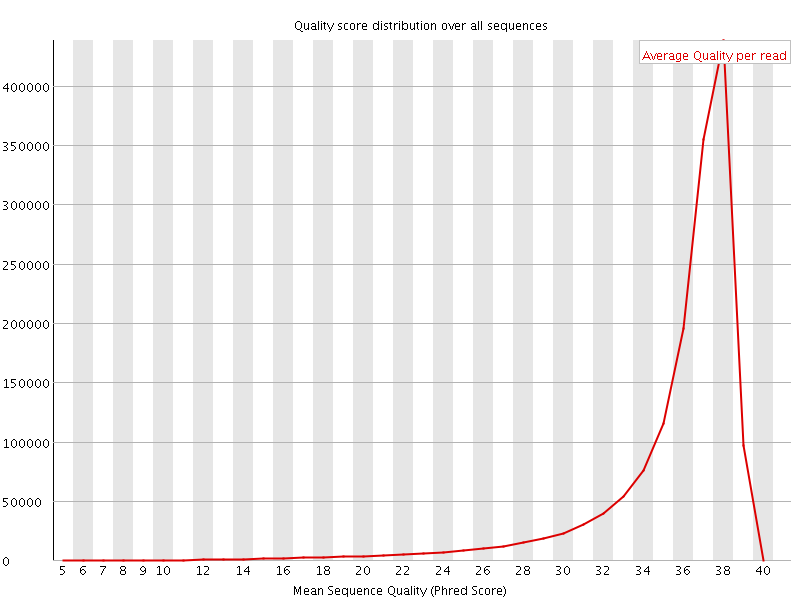
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
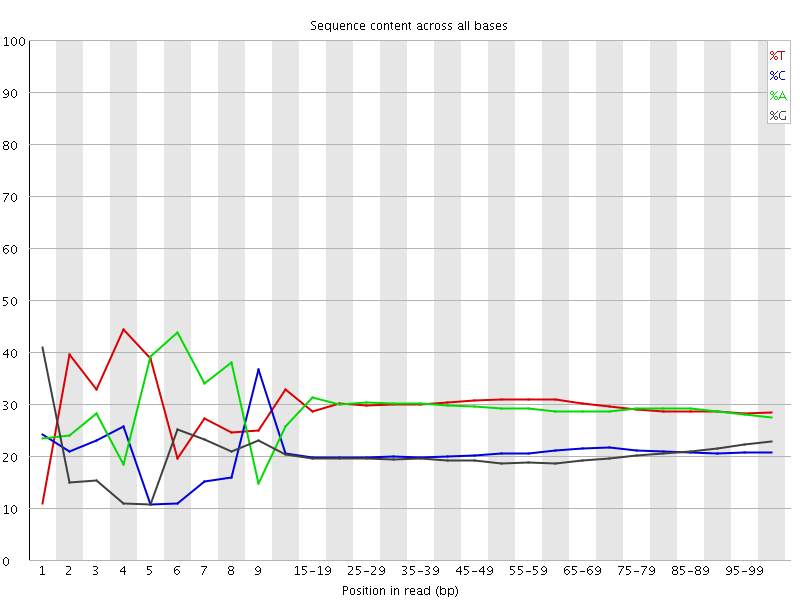
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
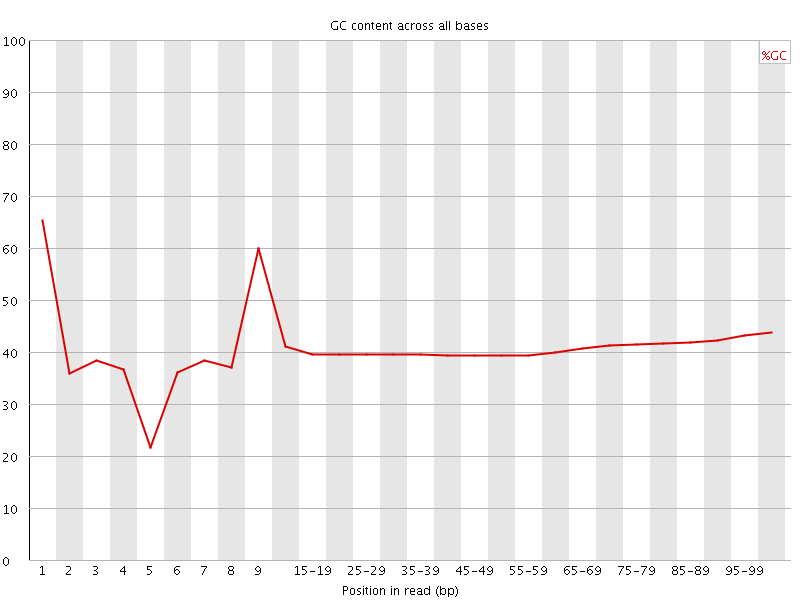
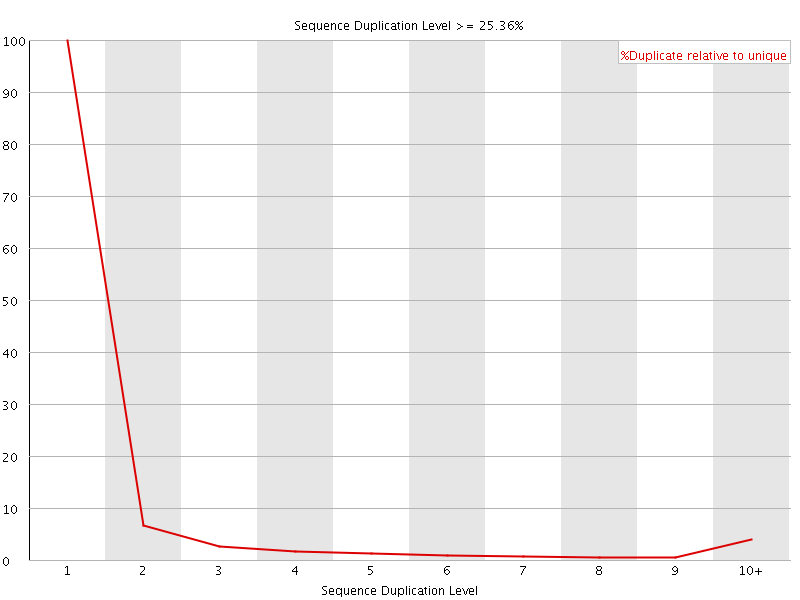
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
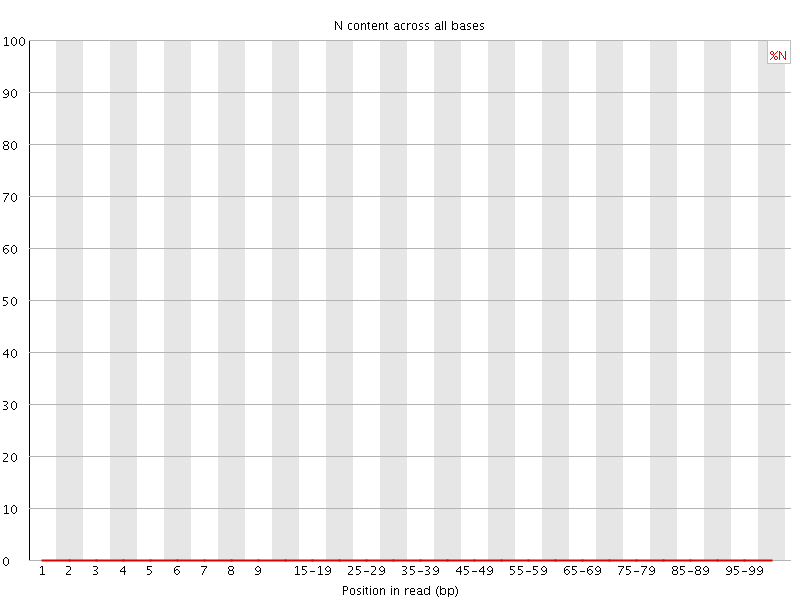
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
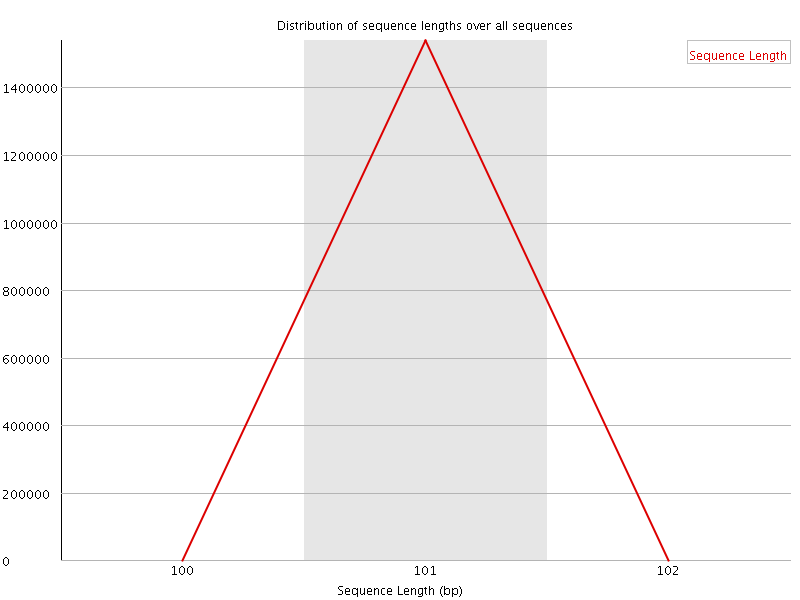
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
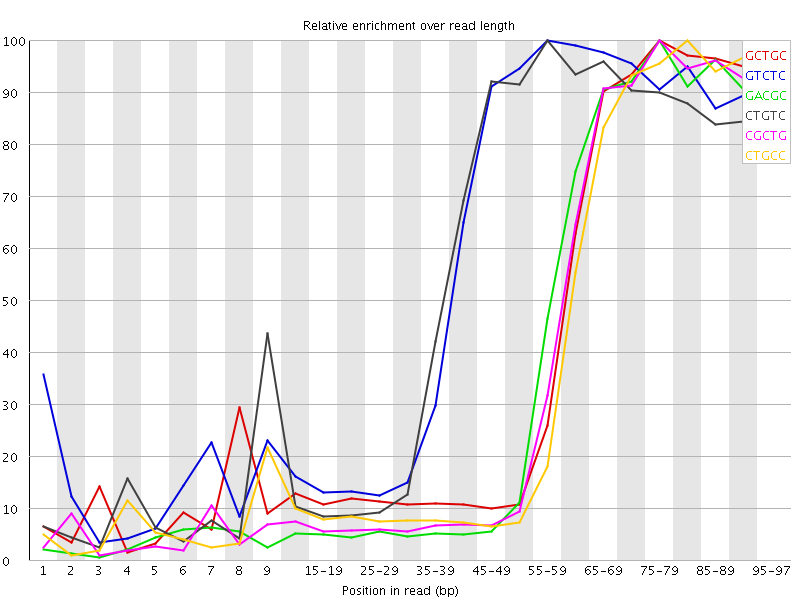
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 370335 | 4.815521 | 11.190461 | 75-79 |
| GTCTC | 540340 | 4.716291 | 7.7338176 | 55-59 |
| GACGC | 345900 | 4.592648 | 11.197219 | 75-79 |
| CTGTC | 516730 | 4.510214 | 7.6787877 | 55-59 |
| CGCTG | 344655 | 4.4816 | 11.068112 | 75-79 |
| CTGCC | 338330 | 4.2818336 | 10.72611 | 80-84 |
| GCCGA | 319120 | 4.2370796 | 11.337944 | 80-84 |
| CGACG | 311970 | 4.1421466 | 11.029569 | 95-97 |
| TCTCT | 699705 | 4.099523 | 6.239374 | 60-64 |
| TGCCG | 312450 | 4.062834 | 11.023027 | 80-84 |
| CCGAC | 307025 | 3.967594 | 10.854539 | 80-84 |
| CTCTT | 654610 | 3.8353145 | 6.110938 | 60-64 |
| GAAGA | 556740 | 3.665907 | 7.404259 | 85-89 |
| TGTCT | 603605 | 3.6335437 | 5.7296925 | 55-59 |
| ACACA | 558550 | 3.4839566 | 6.572948 | 6 |
| CTGAC | 388055 | 3.4585242 | 7.5127697 | 70-74 |
| TGACG | 373420 | 3.4194343 | 7.7526355 | 75-79 |
| GAGGA | 350155 | 3.3638787 | 8.902914 | 90-94 |
| TACAC | 518810 | 3.1692386 | 7.6662154 | 5 |
| ATCTG | 513695 | 3.1575265 | 6.1091213 | 70-74 |
| AGAGG | 328220 | 3.1531527 | 8.653176 | 90-94 |
| CACAT | 513335 | 3.1357934 | 5.4864497 | 65-69 |
| CATCT | 522645 | 3.1267219 | 5.818257 | 70-74 |
| GACGA | 334315 | 3.1259117 | 8.253136 | 85-89 |
| ACGCT | 348095 | 3.1023822 | 7.5645566 | 75-79 |
| ACATC | 496165 | 3.0309076 | 5.690314 | 70-74 |
| TCTGA | 484405 | 2.97749 | 5.7561445 | 70-74 |
| CGAAG | 315965 | 2.9543355 | 8.230462 | 85-89 |
| AAGAG | 434545 | 2.8613024 | 6.358734 | 85-89 |
| AGAAG | 357090 | 2.3512926 | 5.069253 | 5 |
| TATAC | 542605 | 2.2859907 | 5.718425 | 5 |
| ACGAA | 348580 | 2.2339442 | 5.783574 | 85-89 |
| AGGAT | 331400 | 2.1370635 | 5.8247166 | 90-94 |
| GGATA | 293605 | 1.8933389 | 5.5676336 | 90-94 |
| GGTGG | 135835 | 1.8645694 | 7.5700784 | 95-97 |
| GGGGG | 89010 | 1.8202066 | 5.104827 | 95-97 |
| AGAGA | 270525 | 1.7812973 | 5.1028132 | 8 |
| CTTTG | 293960 | 1.7695621 | 7.517098 | 9 |
| GATAG | 266810 | 1.7205489 | 5.3620796 | 95-97 |
| GAGAG | 172855 | 1.6605881 | 5.5859547 | 7 |
| TAGTG | 262915 | 1.6604133 | 5.088979 | 95-97 |
| CTCGG | 124925 | 1.6244183 | 7.4737735 | 95-97 |
| GTGTA | 246145 | 1.5545039 | 8.324147 | 1 |
| CGGTG | 97885 | 1.3077472 | 6.590247 | 95-97 |
| CGCCG | 64855 | 1.2227787 | 5.294879 | 95-97 |
| AAGAC | 187685 | 1.2028166 | 5.5027666 | 5 |
| GAGTC | 128075 | 1.1727922 | 5.6885457 | 9 |
| GTCTT | 188300 | 1.1335166 | 5.8113885 | 1 |
| TATGA | 253625 | 1.0978472 | 5.201323 | 4 |
| GTTCT | 182190 | 1.096736 | 5.0988364 | 1 |
| TCATA | 255595 | 1.0768198 | 5.262599 | 2 |
| GTGTT | 173660 | 1.0740796 | 5.817867 | 1 |
| GACTT | 173850 | 1.0686029 | 5.472838 | 7 |
| TGGAC | 106195 | 0.9724354 | 6.290106 | 5 |
| GTCCA | 106125 | 0.9458347 | 6.6237755 | 1 |
| AGACT | 142895 | 0.8968561 | 5.092036 | 6 |
| GGACT | 95795 | 0.8772018 | 5.6485057 | 6 |
| GTATA | 156515 | 0.6774945 | 6.8372827 | 1 |