![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-163_GCTACGCT-TATCCTCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1538541 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
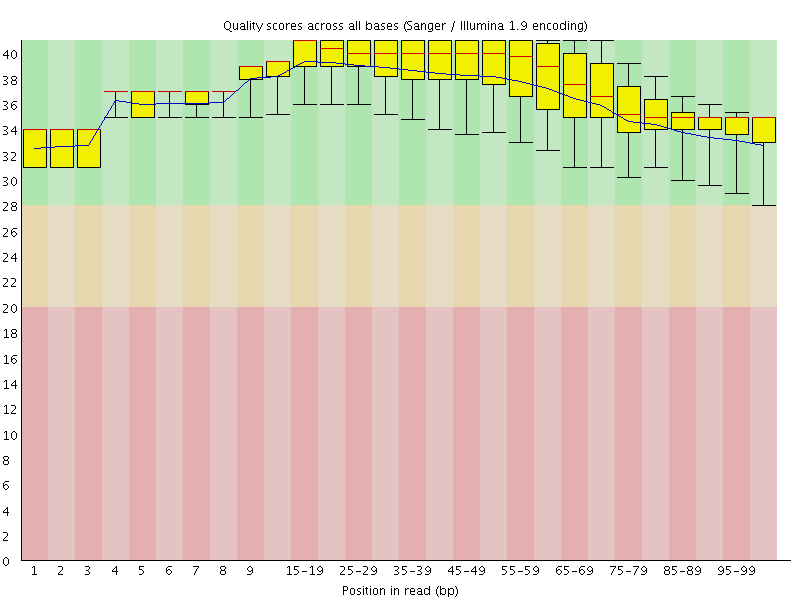
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
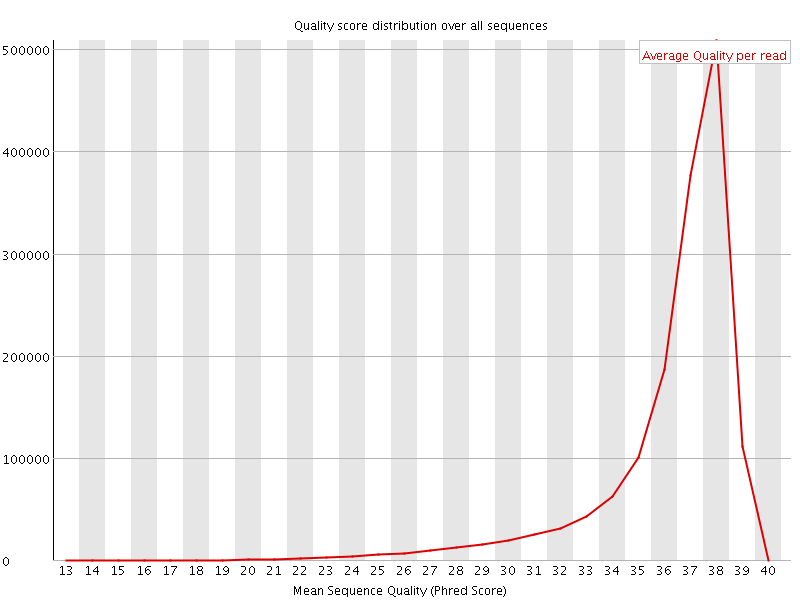
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
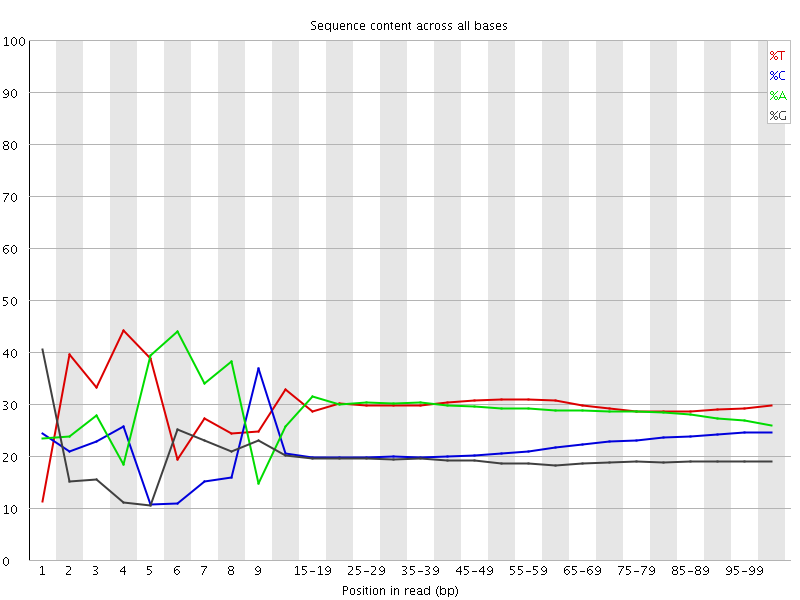
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
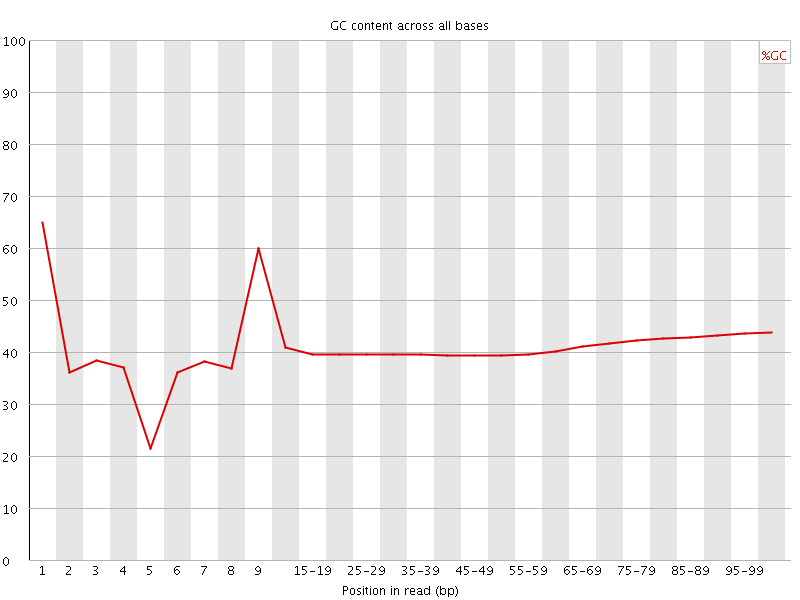
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
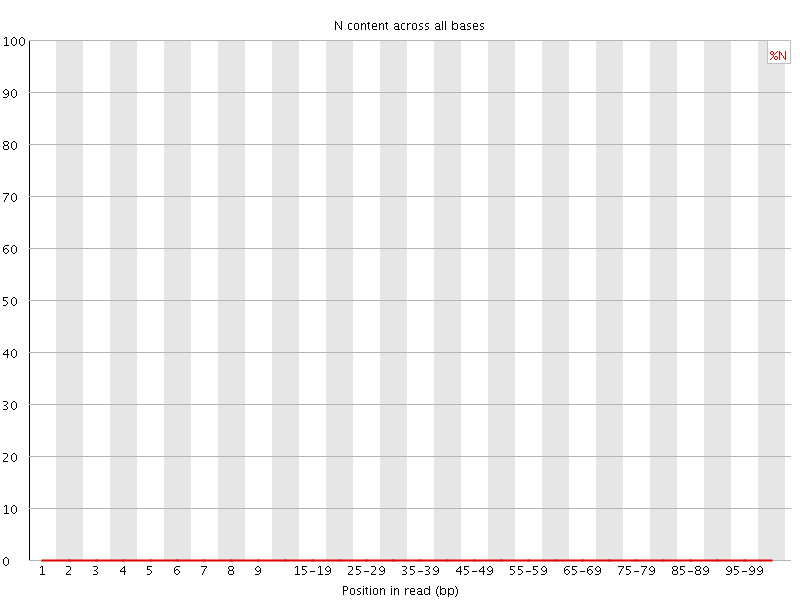
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
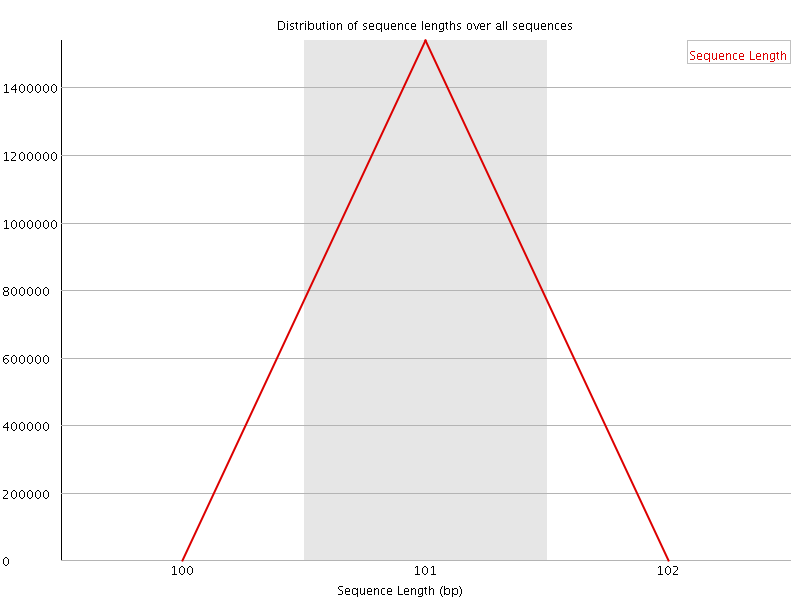
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
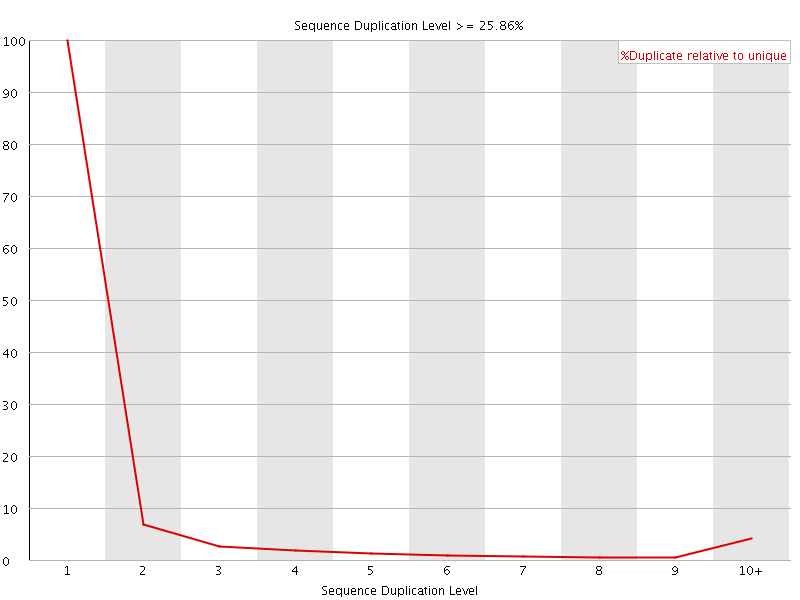
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
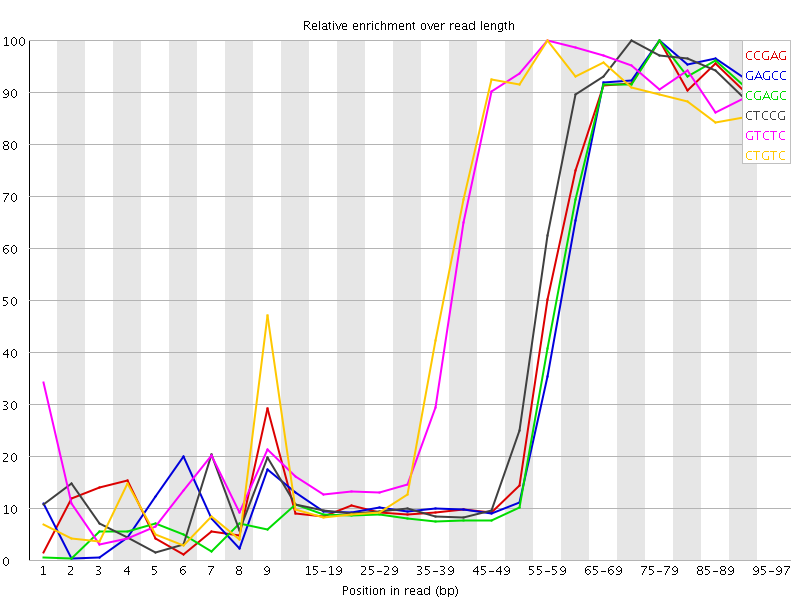
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 380935 | 4.9971933 | 11.461645 | 75-79 |
| GAGCC | 369800 | 4.851122 | 11.314081 | 75-79 |
| CGAGC | 359630 | 4.7177095 | 11.244116 | 75-79 |
| CTCCG | 394310 | 4.5033326 | 9.718653 | 70-74 |
| GTCTC | 540110 | 4.4611893 | 7.3656974 | 55-59 |
| CTGTC | 517885 | 4.2776165 | 7.2727356 | 55-59 |
| AGCCC | 352095 | 4.137047 | 9.836469 | 75-79 |
| GCCCA | 347675 | 4.085112 | 9.981296 | 80-84 |
| TCTCT | 702175 | 3.7570012 | 5.717496 | 60-64 |
| CCACG | 314380 | 3.6939025 | 9.8901415 | 80-84 |
| ACGCT | 426000 | 3.6200318 | 13.513362 | 90-94 |
| TGTCT | 604065 | 3.6084805 | 5.7362347 | 55-59 |
| TCTCC | 485625 | 3.5927312 | 6.9778347 | 70-74 |
| ATCTC | 652295 | 3.5906603 | 8.865671 | 95-97 |
| CCCAC | 335665 | 3.5325787 | 8.996226 | 80-84 |
| CTCTT | 655155 | 3.5054197 | 5.522186 | 60-64 |
| TCCGA | 408580 | 3.4720013 | 7.4493456 | 75-79 |
| CGCTA | 398195 | 3.383752 | 13.415295 | 90-94 |
| CGAGA | 342305 | 3.3411431 | 8.632897 | 85-89 |
| GAGAC | 341925 | 3.337434 | 8.811803 | 85-89 |
| GACGC | 253890 | 3.3305876 | 10.914199 | 85-89 |
| ACACA | 560320 | 3.264638 | 6.0537534 | 6 |
| ACGAG | 328995 | 3.211228 | 8.293489 | 80-84 |
| TACAC | 519395 | 2.941456 | 7.0814066 | 5 |
| CACAT | 515695 | 2.920502 | 5.128378 | 65-69 |
| CATCT | 521790 | 2.8722749 | 5.343515 | 70-74 |
| AGACG | 291255 | 2.8428586 | 8.403823 | 85-89 |
| CACGA | 322075 | 2.8157501 | 7.4783735 | 80-84 |
| ACATC | 496995 | 2.8145995 | 5.181663 | 70-74 |
| AGAAG | 358135 | 2.6009688 | 5.711404 | 5 |
| GCTAC | 283565 | 2.409658 | 7.674713 | 90-94 |
| TATAC | 545225 | 2.2331228 | 5.8780355 | 5 |
| CTACG | 234485 | 1.9925894 | 7.200039 | 90-94 |
| CAAAG | 298035 | 1.9387014 | 5.0241556 | 4 |
| AGAGA | 266590 | 1.93612 | 5.686756 | 8 |
| TACGC | 221080 | 1.8786775 | 7.0978527 | 90-94 |
| GAGAG | 168785 | 1.8393333 | 6.6943254 | 7 |
| TCTCG | 221700 | 1.8311932 | 6.644382 | 90-94 |
| CTTTG | 293420 | 1.7527921 | 7.640453 | 9 |
| AAGAG | 230465 | 1.6737605 | 5.123361 | 7 |
| CTCGT | 193110 | 1.5950462 | 6.5666847 | 90-94 |
| AGGAG | 139480 | 1.5199822 | 5.531933 | 7 |
| GTATG | 219280 | 1.5045921 | 5.7704377 | 90-94 |
| GCTAT | 243705 | 1.4977506 | 5.3816824 | 95-97 |
| TGCCG | 109475 | 1.3959051 | 9.894109 | 95-97 |
| GCCGT | 107625 | 1.372316 | 9.712754 | 95-97 |
| GTCTT | 223775 | 1.3367563 | 5.689046 | 1 |
| ATGCC | 154580 | 1.3135786 | 6.779254 | 95-97 |
| CCGTC | 109145 | 1.2465222 | 8.020288 | 95-97 |
| AAGAC | 189715 | 1.2340856 | 5.749551 | 5 |
| GAGTC | 128625 | 1.2203178 | 6.0765066 | 9 |
| GATTG | 175495 | 1.2041608 | 5.4791975 | 7 |
| TATGA | 253670 | 1.1599804 | 5.323256 | 4 |
| GTGTT | 170865 | 1.1395636 | 5.8923845 | 1 |
| GACTT | 175785 | 1.0803311 | 5.7121654 | 7 |
| TCATA | 255420 | 1.0461447 | 5.060501 | 2 |
| TGGAC | 106650 | 1.011832 | 6.9596934 | 5 |
| GTGTA | 147080 | 1.009191 | 8.790425 | 1 |
| CGTCT | 114730 | 0.9476446 | 5.305183 | 95-97 |
| GAGTA | 130260 | 0.9195284 | 5.4186645 | 1 |
| GGACT | 95905 | 0.90988964 | 6.140905 | 6 |
| GTCCA | 105965 | 0.90046155 | 6.271547 | 1 |
| AGACT | 141845 | 0.8968574 | 5.0030303 | 6 |
| GAATA | 188685 | 0.88767356 | 5.0005035 | 1 |
| GGATA | 121175 | 0.85539573 | 5.1619363 | 1 |
| GTATA | 156340 | 0.71491045 | 7.060117 | 1 |