![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-162_GCTACGCT-CTCTCTAT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2254610 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
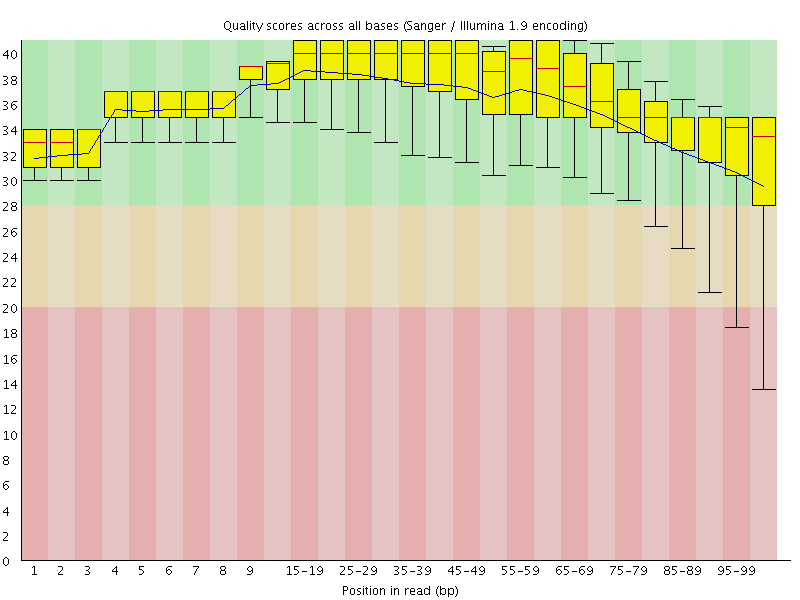
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
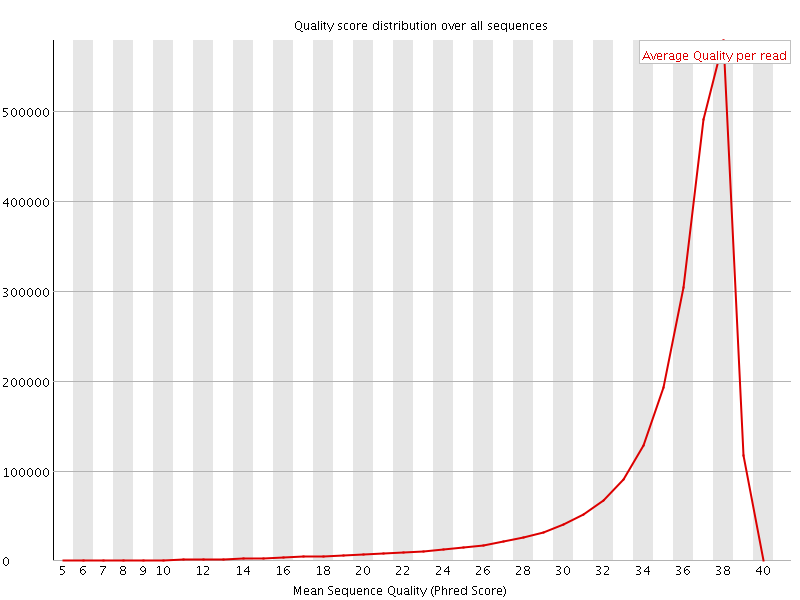
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
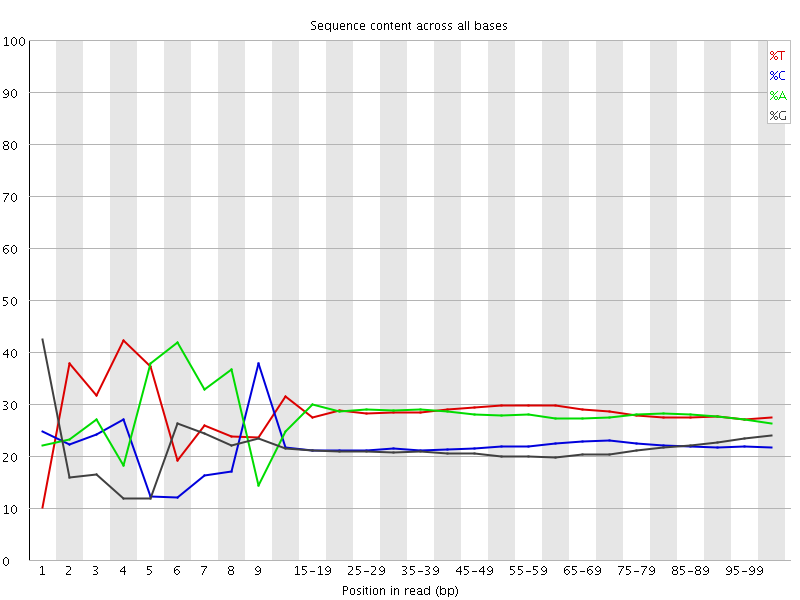
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
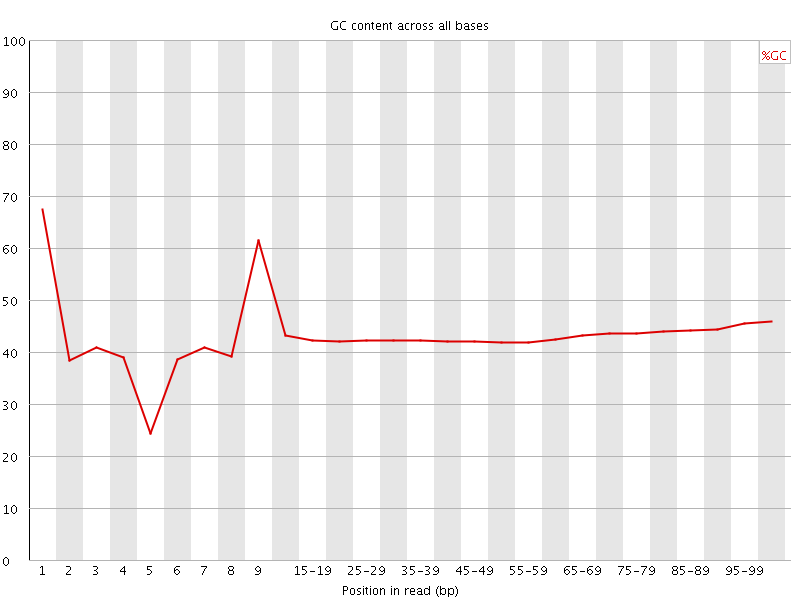
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
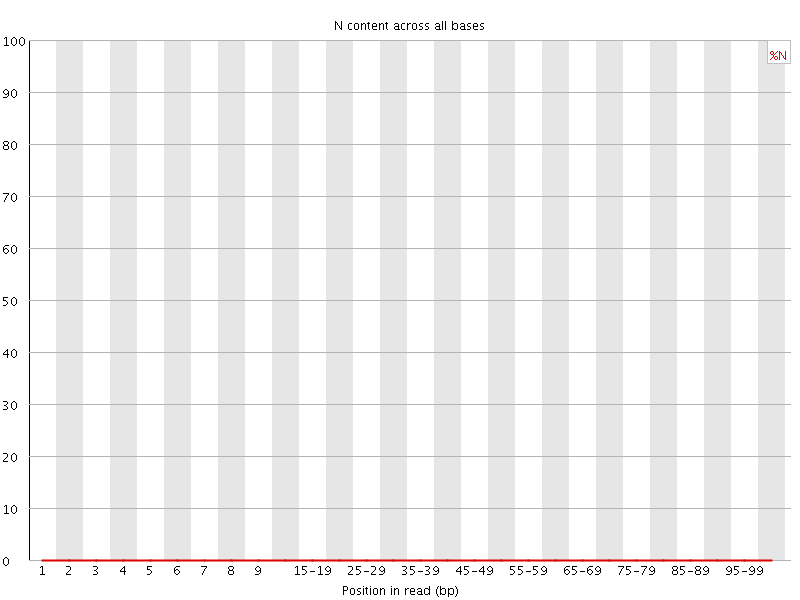
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
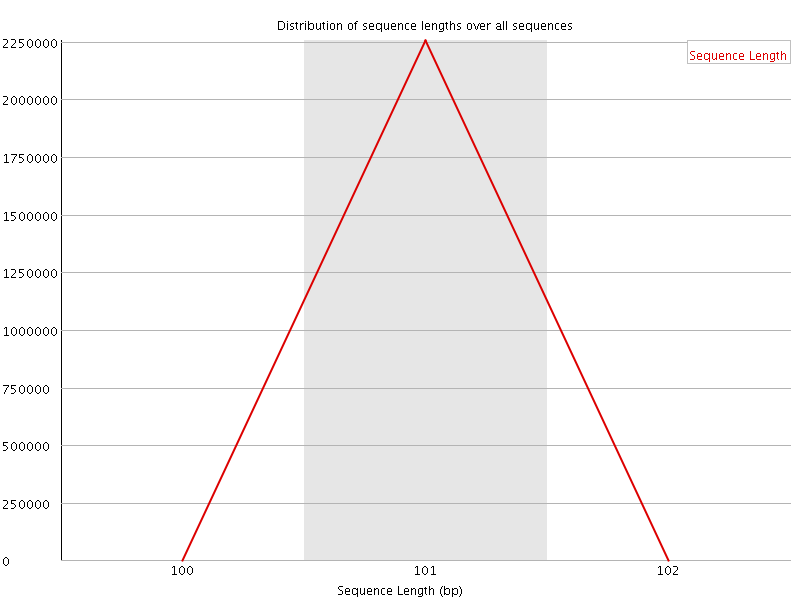
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
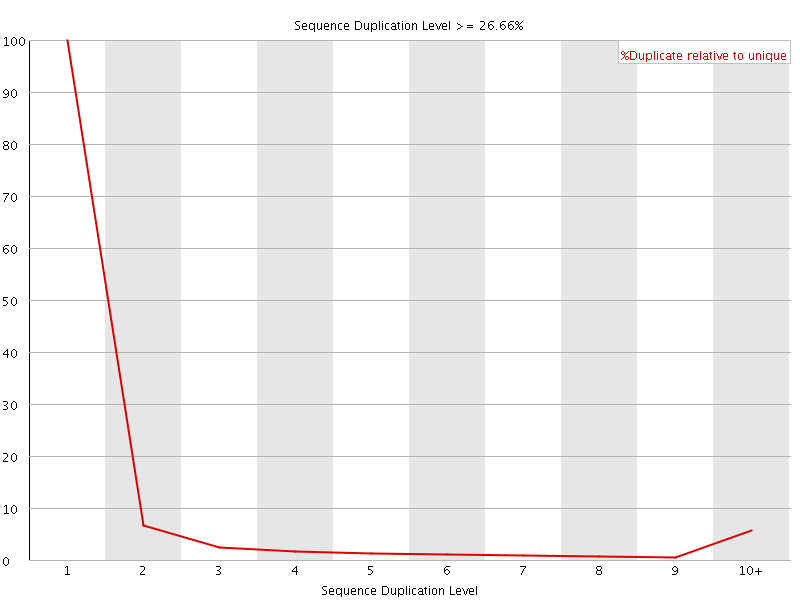
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[FAIL]](Icons/error.png) Kmer Content
Kmer Content
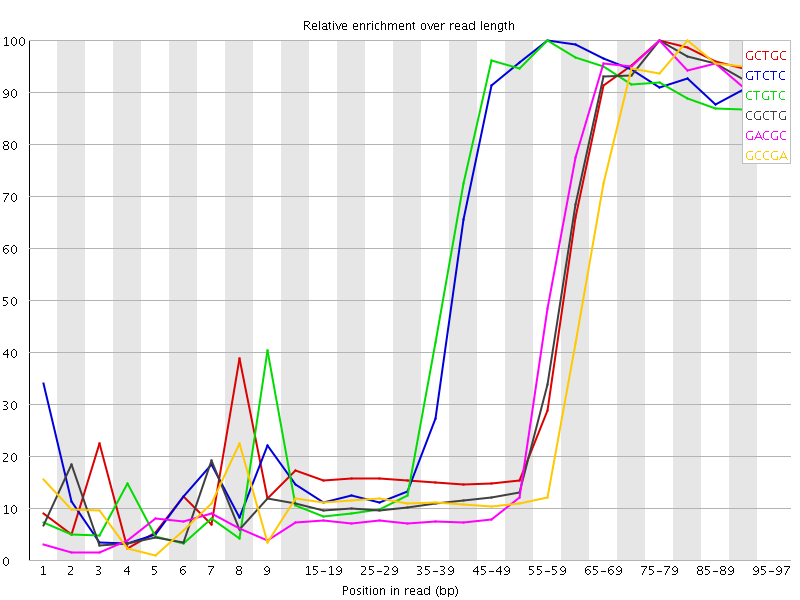
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GCTGC | 633990 | 4.6524706 | 10.155498 | 75-79 |
| GTCTC | 820660 | 4.4626803 | 7.3928747 | 55-59 |
| CTGTC | 803350 | 4.3685503 | 7.300042 | 55-59 |
| CGCTG | 594580 | 4.363264 | 10.04659 | 75-79 |
| GACGC | 565240 | 4.235147 | 9.830426 | 75-79 |
| GCCGA | 543620 | 4.0731554 | 10.123642 | 80-84 |
| TCTCT | 998815 | 4.0248427 | 6.348052 | 60-64 |
| CTGCC | 560925 | 3.9981122 | 9.565556 | 80-84 |
| TGCCG | 538500 | 3.951727 | 9.764289 | 80-84 |
| CGACG | 518095 | 3.8819056 | 9.883036 | 80-84 |
| CTCTT | 949095 | 3.82449 | 6.138939 | 60-64 |
| TGTCT | 914130 | 3.7924757 | 5.9806566 | 55-59 |
| ACACA | 860320 | 3.690005 | 6.6008306 | 70-74 |
| CCGAC | 496590 | 3.6139536 | 9.5781145 | 80-84 |
| TACAC | 813670 | 3.4180696 | 8.0977125 | 5 |
| CTGAC | 607385 | 3.3723369 | 7.2671986 | 70-74 |
| TGACG | 588905 | 3.36638 | 7.551452 | 75-79 |
| CACAT | 789565 | 3.3168092 | 5.9136233 | 60-64 |
| ATCTG | 771835 | 3.2694426 | 6.3328595 | 70-74 |
| ACATC | 762785 | 3.2043116 | 5.9466004 | 70-74 |
| CATCT | 778525 | 3.203102 | 5.9538846 | 70-74 |
| GACGA | 529620 | 3.0911257 | 8.0208435 | 85-89 |
| ACGCT | 551185 | 3.0603023 | 7.284565 | 75-79 |
| TCTTA | 966925 | 3.0351021 | 4.722024 | 60-64 |
| TCTGA | 710550 | 3.009843 | 5.9478416 | 70-74 |
| GAGAG | 489105 | 2.9390395 | 8.212436 | 90-94 |
| ATACA | 890205 | 2.9129922 | 5.2493424 | 6 |
| AGAGA | 613525 | 2.7893388 | 6.6677375 | 90-94 |
| AGAGG | 435295 | 2.6156943 | 8.058528 | 95-97 |
| TATAC | 814170 | 2.609336 | 5.5327973 | 5 |
| ACGAA | 525695 | 2.32141 | 6.0381503 | 85-89 |
| CGAAT | 526725 | 2.2780724 | 5.8773417 | 85-89 |
| GAGGT | 352215 | 2.0728924 | 6.705161 | 95-97 |
| TAGAG | 461735 | 2.0560205 | 6.0044 | 90-94 |
| AGGTG | 346595 | 2.039817 | 6.994415 | 95-97 |
| GGTGT | 338450 | 1.9508735 | 5.936583 | 85-89 |
| CTTTG | 464185 | 1.9257767 | 8.920992 | 9 |
| GGTGG | 239255 | 1.8610767 | 7.799576 | 95-97 |
| GTGTA | 385485 | 1.681155 | 9.058525 | 1 |
| GCTTT | 397795 | 1.6503428 | 5.3470836 | 1 |
| CTCGG | 203900 | 1.4962991 | 6.6641016 | 95-97 |
| CGGTG | 181115 | 1.3683797 | 6.356609 | 95-97 |
| AAGAC | 292030 | 1.2895715 | 5.661311 | 5 |
| TATGA | 387510 | 1.2786418 | 6.013247 | 4 |
| TCATA | 388890 | 1.2463548 | 6.0924244 | 2 |
| GAGTC | 207410 | 1.1856257 | 6.877765 | 9 |
| GTCTT | 285435 | 1.184192 | 6.210425 | 1 |
| GACTT | 278685 | 1.1804914 | 6.5200963 | 7 |
| ATGAG | 256665 | 1.1428818 | 5.361782 | 7 |
| TGAGT | 258425 | 1.1270282 | 5.7929626 | 8 |
| GTGTT | 263355 | 1.1248834 | 5.604589 | 1 |
| GATTG | 253350 | 1.1048955 | 5.410017 | 7 |
| TGGAC | 179020 | 1.0233388 | 6.8987465 | 5 |
| GTCCA | 176225 | 0.97844046 | 7.4791684 | 1 |
| GGACT | 157880 | 0.90249544 | 6.4728413 | 6 |
| AGACT | 207515 | 0.89749724 | 5.2497015 | 6 |
| TATAA | 334625 | 0.8353914 | 5.2350016 | 2 |
| GTATA | 212810 | 0.70219547 | 6.860072 | 1 |