![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-162_GCTACGCT-CTCTCTAT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 2254610 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 43 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
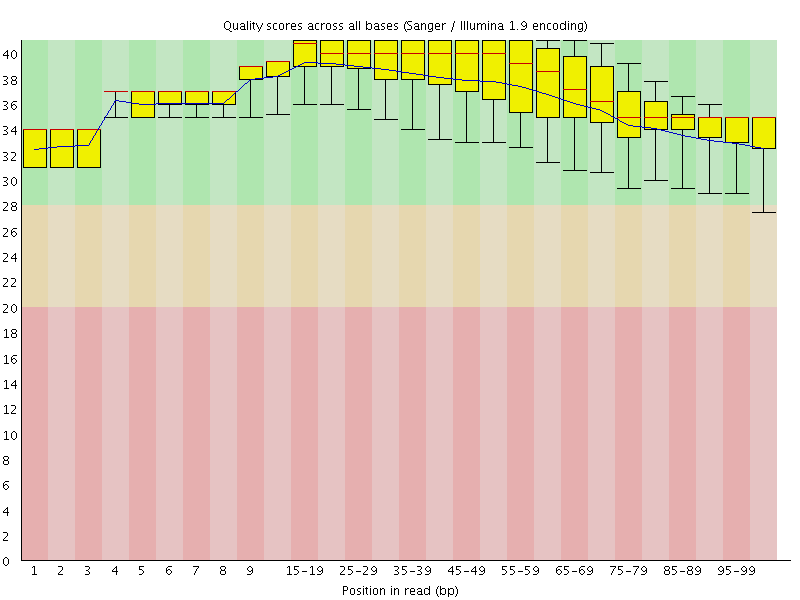
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
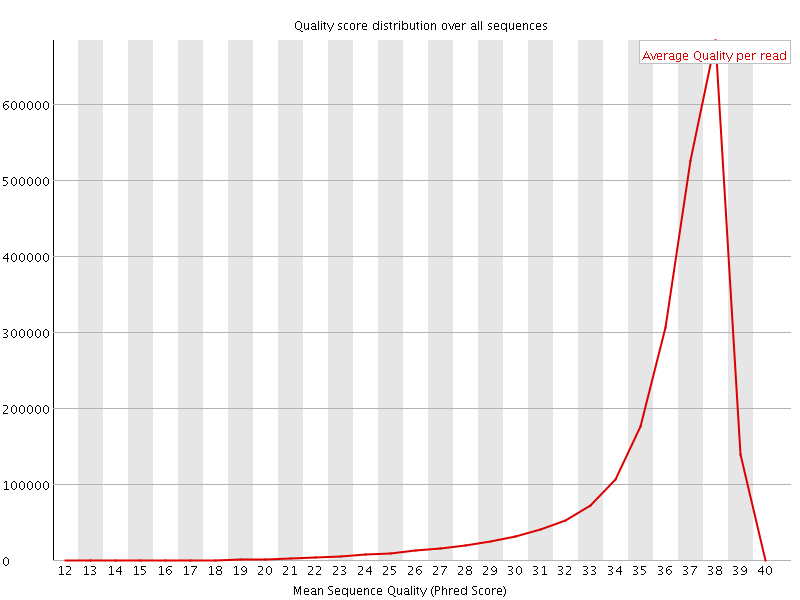
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
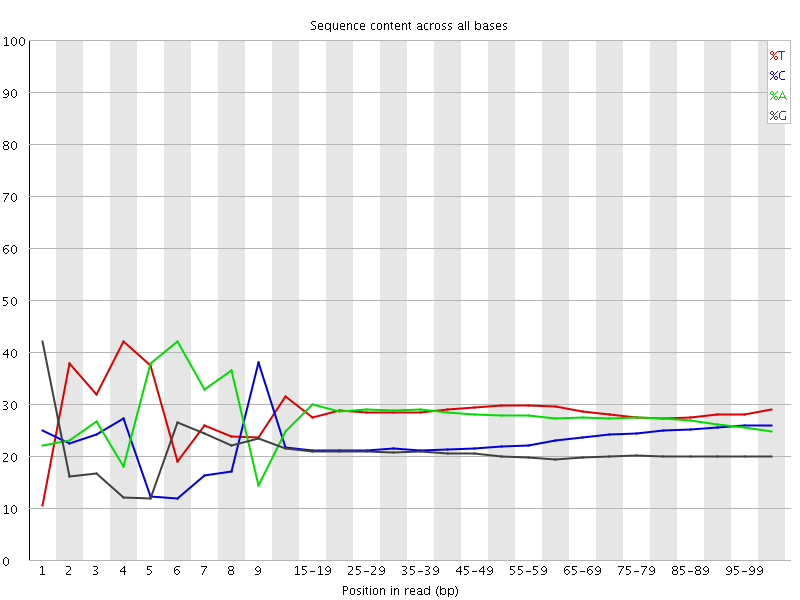
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
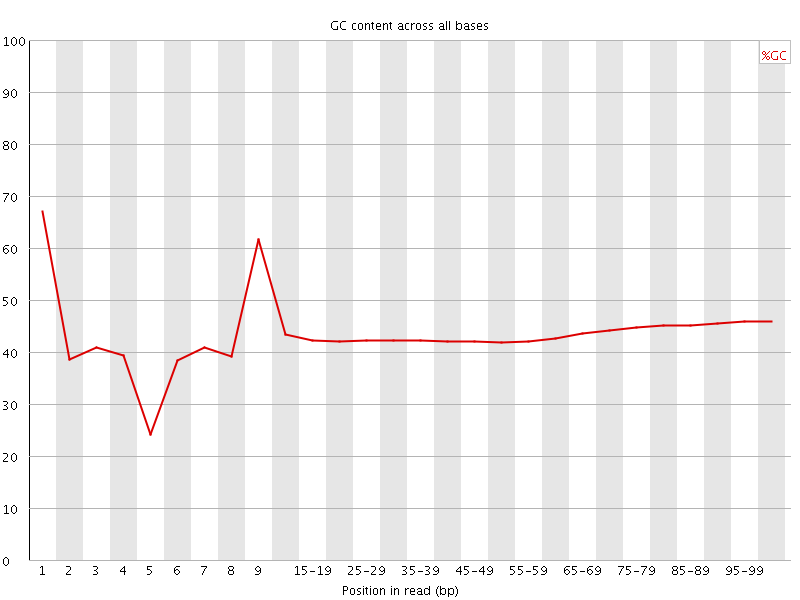
![[FAIL]](Icons/error.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
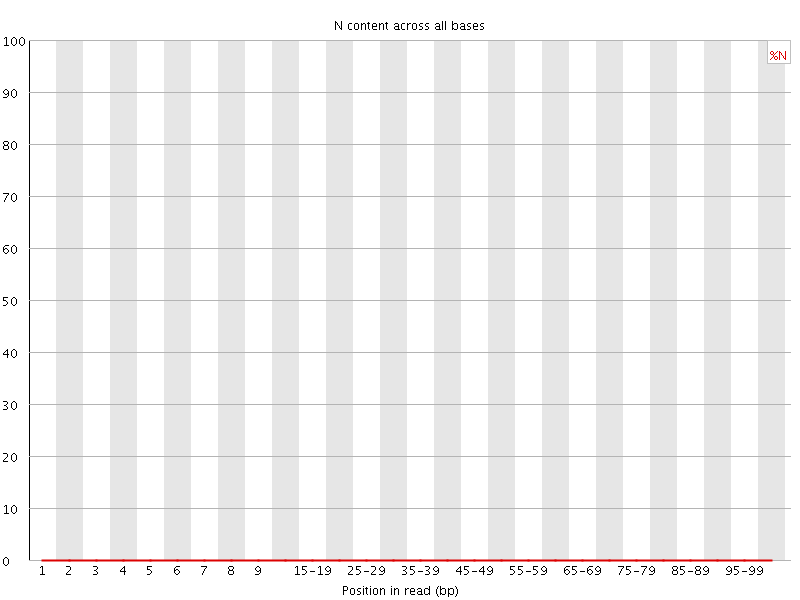
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
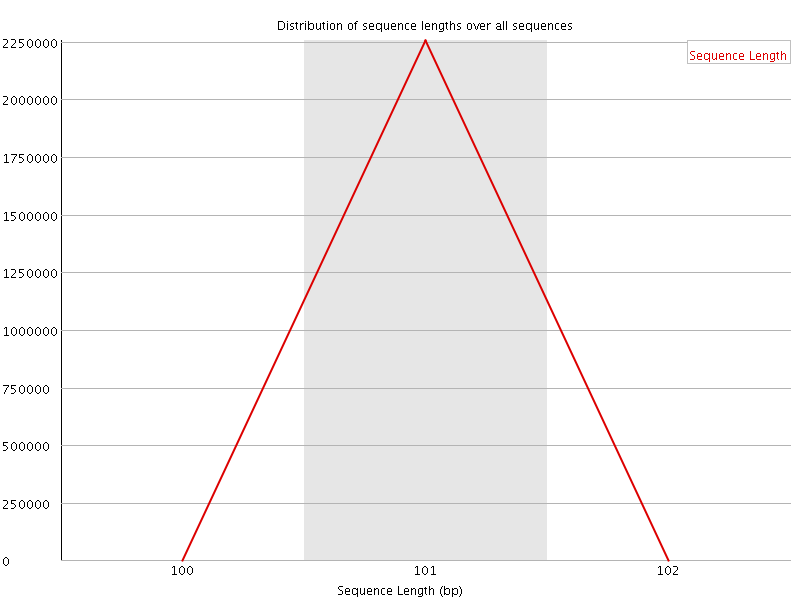
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
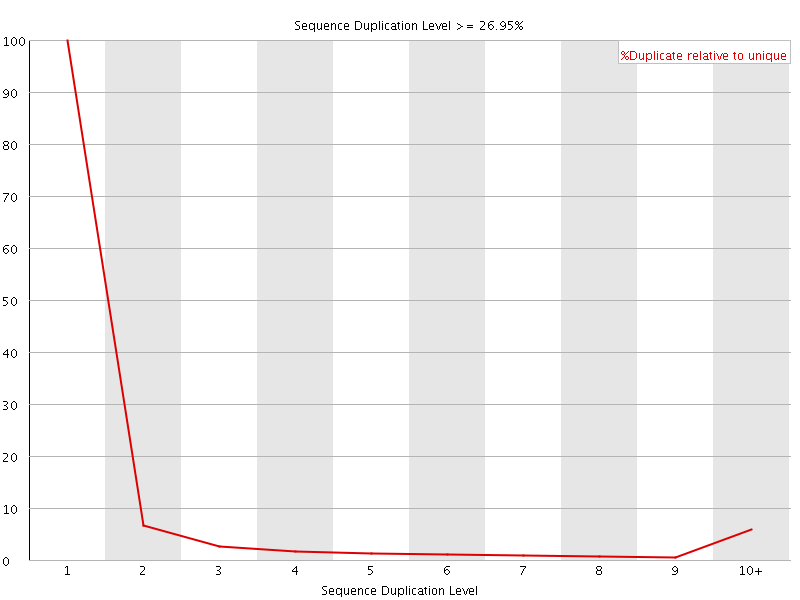
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
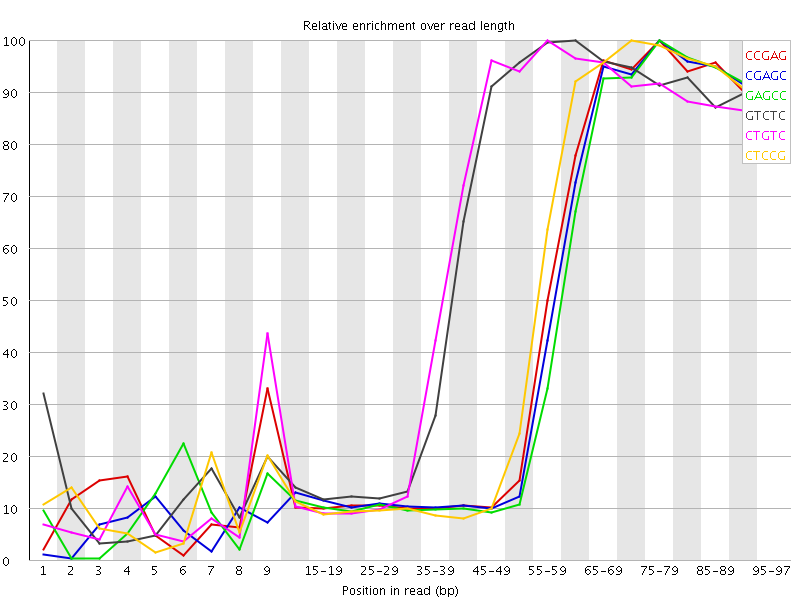
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| CCGAG | 604970 | 4.4774985 | 9.953426 | 75-79 |
| CGAGC | 588960 | 4.359005 | 9.928305 | 75-79 |
| GAGCC | 577455 | 4.2738547 | 9.965628 | 75-79 |
| GTCTC | 820645 | 4.2055364 | 6.979087 | 60-64 |
| CTGTC | 806350 | 4.132279 | 6.902351 | 55-59 |
| CTCCG | 611870 | 3.9275157 | 8.384816 | 70-74 |
| TGTCT | 915560 | 3.745934 | 5.8861976 | 55-59 |
| ATCTC | 977555 | 3.7101483 | 9.407478 | 95-97 |
| TCTCT | 1003270 | 3.6817827 | 5.7985654 | 60-64 |
| GCCCA | 553765 | 3.6761575 | 8.814378 | 80-84 |
| AGCCC | 553360 | 3.6734688 | 8.66247 | 75-79 |
| ACGCT | 675235 | 3.5787456 | 13.153814 | 90-94 |
| CTCTT | 952750 | 3.496385 | 5.626499 | 60-64 |
| ACACA | 859120 | 3.4875746 | 6.2554817 | 70-74 |
| CCACG | 509865 | 3.384728 | 8.682406 | 80-84 |
| TCTCC | 724970 | 3.332369 | 6.5268054 | 70-74 |
| CGCTA | 622750 | 3.3005748 | 13.010927 | 90-94 |
| TCCGA | 622545 | 3.2994878 | 7.1184063 | 75-79 |
| CGAGA | 527890 | 3.225983 | 8.297682 | 85-89 |
| TACAC | 816575 | 3.205206 | 7.56498 | 5 |
| GAGAC | 516175 | 3.1543918 | 8.374728 | 85-89 |
| CCCAC | 528495 | 3.1468532 | 7.844828 | 80-84 |
| GACGC | 422505 | 3.1270397 | 9.641702 | 85-89 |
| ACGAG | 509475 | 3.1134472 | 8.062568 | 80-84 |
| CACAT | 788555 | 3.0952225 | 5.4941545 | 60-64 |
| ACATC | 764270 | 2.9998996 | 5.5525575 | 70-74 |
| CATCT | 779260 | 2.957553 | 5.5021124 | 70-74 |
| ATACA | 889795 | 2.8838117 | 5.3254867 | 6 |
| CACGA | 505445 | 2.770509 | 7.2577457 | 80-84 |
| AGACG | 450710 | 2.7543292 | 8.006693 | 85-89 |
| TATAC | 815910 | 2.5568748 | 5.5899477 | 5 |
| GCTAC | 449480 | 2.3822436 | 7.360214 | 90-94 |
| CAAAG | 472085 | 2.1365983 | 5.0207605 | 4 |
| CTACG | 366335 | 1.9415752 | 6.810254 | 90-94 |
| CTTTG | 468590 | 1.9171952 | 8.958557 | 9 |
| TACGC | 352715 | 1.8693892 | 6.822101 | 95-97 |
| TCTCG | 339850 | 1.7416198 | 6.572045 | 90-94 |
| GCTTT | 399845 | 1.635931 | 5.061176 | 1 |
| GTATG | 336150 | 1.5858028 | 6.358285 | 90-94 |
| TGCCG | 218930 | 1.5667404 | 8.997598 | 95-97 |
| GCTAT | 362470 | 1.5337522 | 5.4855814 | 95-97 |
| CTCGT | 296085 | 1.5173384 | 6.4756308 | 90-94 |
| TTGAG | 317490 | 1.4977734 | 5.0092306 | 9 |
| GAGAG | 218175 | 1.4864718 | 5.1103754 | 7 |
| ATGGA | 294565 | 1.4371663 | 5.322543 | 4 |
| GCCGT | 197100 | 1.4105173 | 8.561542 | 95-97 |
| GTCTT | 344625 | 1.4100032 | 5.7039905 | 1 |
| CTATC | 365085 | 1.3856196 | 5.0073843 | 95-97 |
| TATGA | 390860 | 1.3655933 | 6.41006 | 4 |
| ATGCC | 255985 | 1.3567203 | 6.7758436 | 95-97 |
| AAGAC | 290745 | 1.3158758 | 5.8217115 | 5 |
| ATGAG | 256635 | 1.252108 | 5.736519 | 7 |
| GAGTC | 211310 | 1.2486162 | 7.1022544 | 9 |
| TCATA | 390270 | 1.2230164 | 6.1255846 | 2 |
| TGAGT | 259055 | 1.2221037 | 6.212361 | 8 |
| GATTG | 253590 | 1.1963226 | 5.782345 | 7 |
| GTGTT | 261110 | 1.1910497 | 5.801927 | 1 |
| CCGTC | 184275 | 1.182838 | 7.054703 | 95-97 |
| GACTT | 279230 | 1.1815312 | 6.583603 | 7 |
| TCGTA | 265500 | 1.1234342 | 5.2643833 | 90-94 |
| GTGTA | 236575 | 1.1160535 | 9.5714 | 1 |
| TATGC | 262885 | 1.1123692 | 5.3788934 | 95-97 |
| ATGTG | 227910 | 1.0751758 | 5.1030107 | 7 |
| TGGAC | 176970 | 1.0457034 | 7.6351376 | 5 |
| ACATG | 238510 | 1.0437572 | 5.045632 | 8 |
| CGTAT | 246135 | 1.0414933 | 5.200394 | 95-97 |
| GTCCA | 178570 | 0.94642085 | 6.872343 | 1 |
| GGACT | 157565 | 0.9310406 | 7.082199 | 6 |
| AGACT | 206905 | 0.9054488 | 5.2472024 | 6 |
| CGTCT | 176410 | 0.9040433 | 5.1991124 | 95-97 |
| TATAA | 331700 | 0.85828143 | 5.341405 | 2 |
| GTATA | 210355 | 0.73494184 | 7.1512694 | 1 |
| GTATT | 192765 | 0.6512062 | 5.0029597 | 1 |