![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-160_CAGAGAGG-CTAAGCCT_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1531303 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
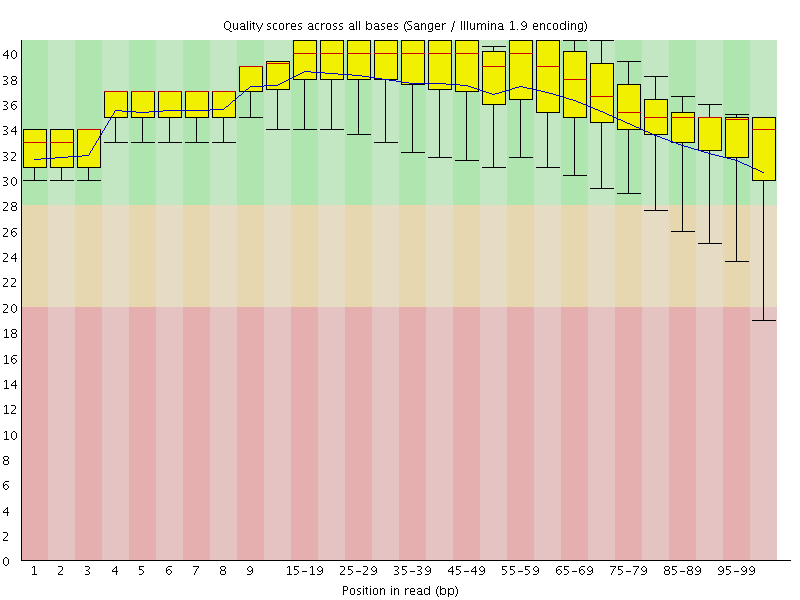
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
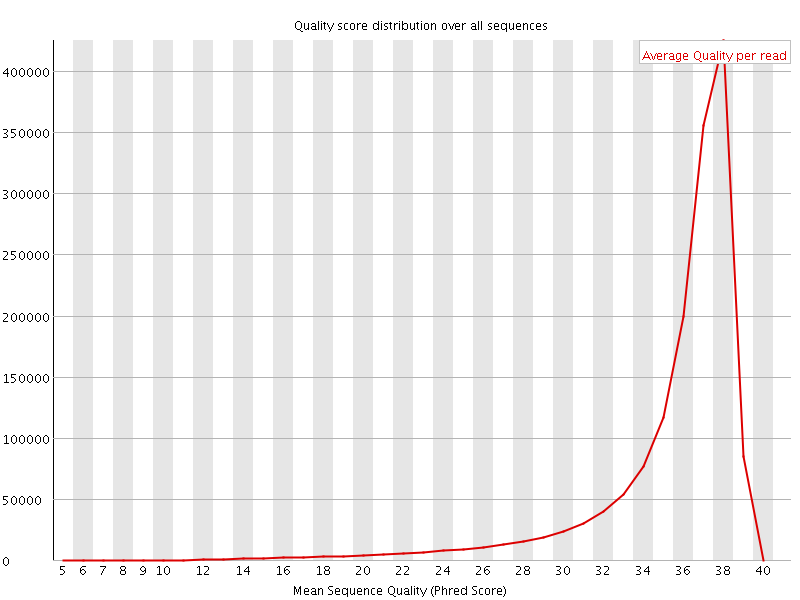
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
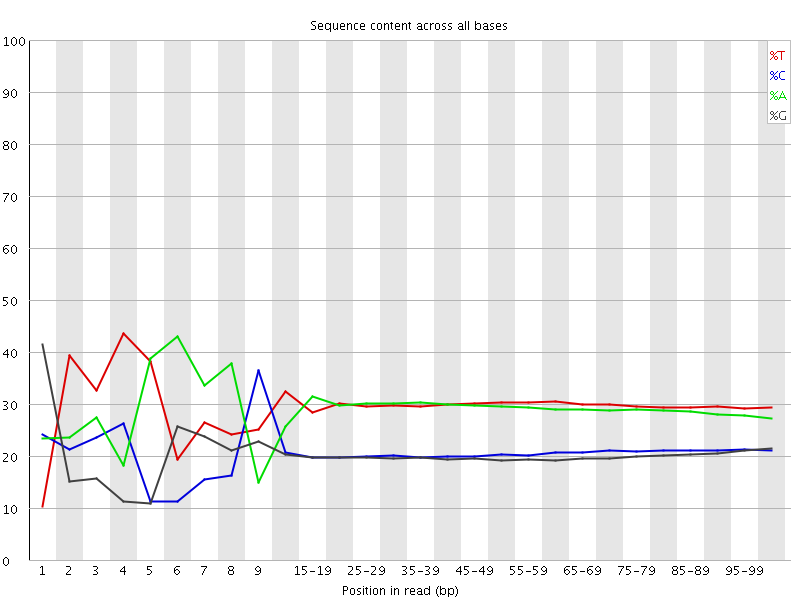
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
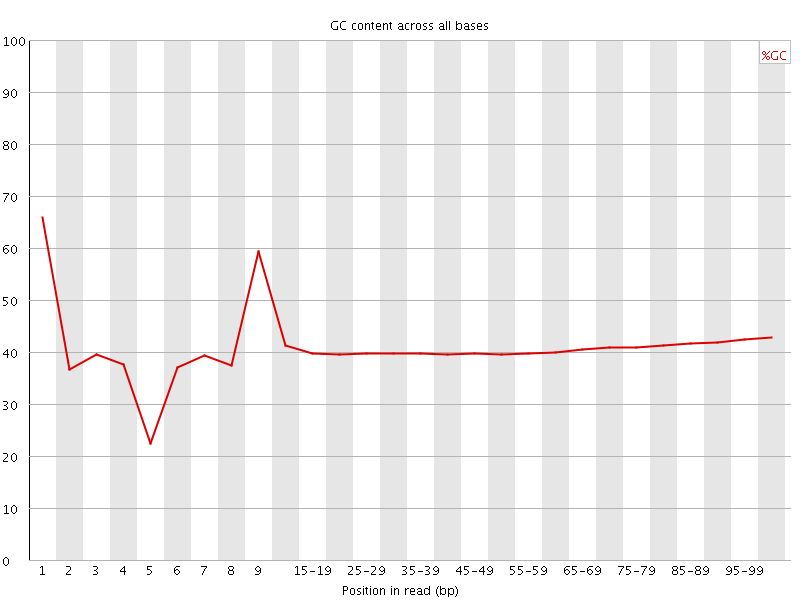
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
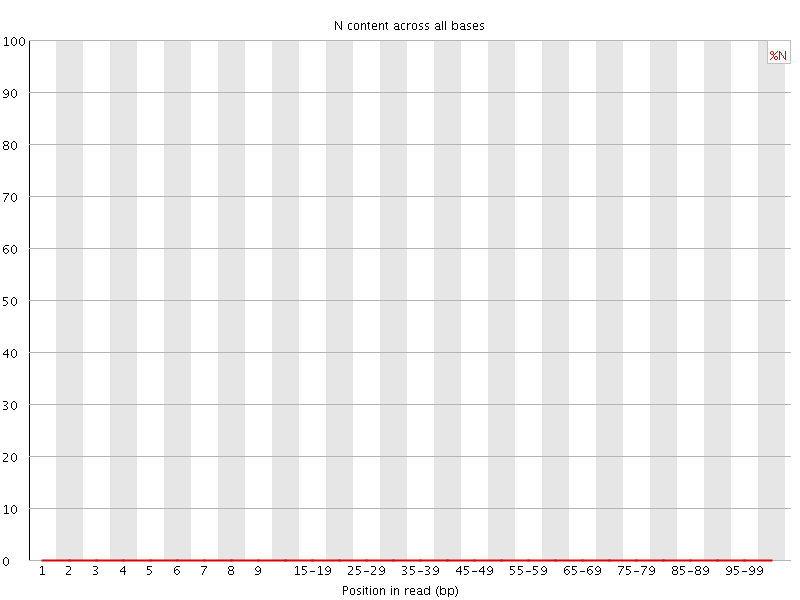
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
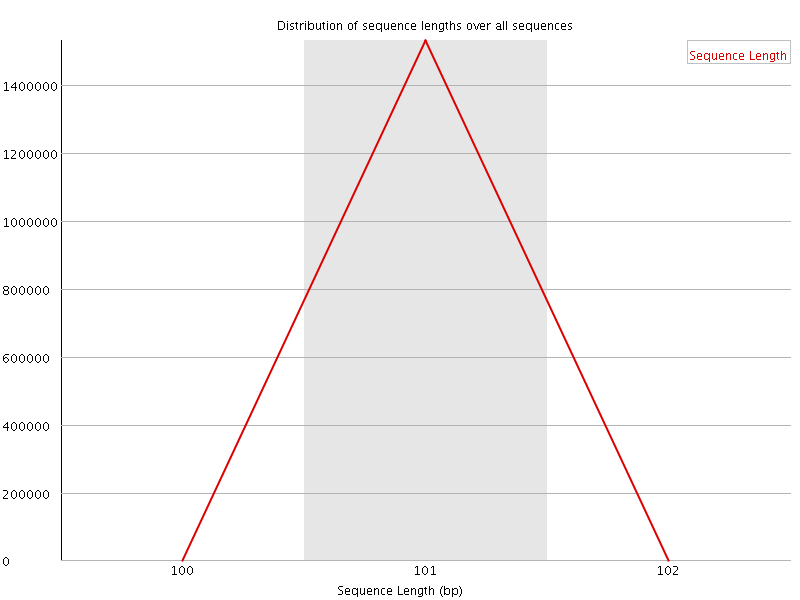
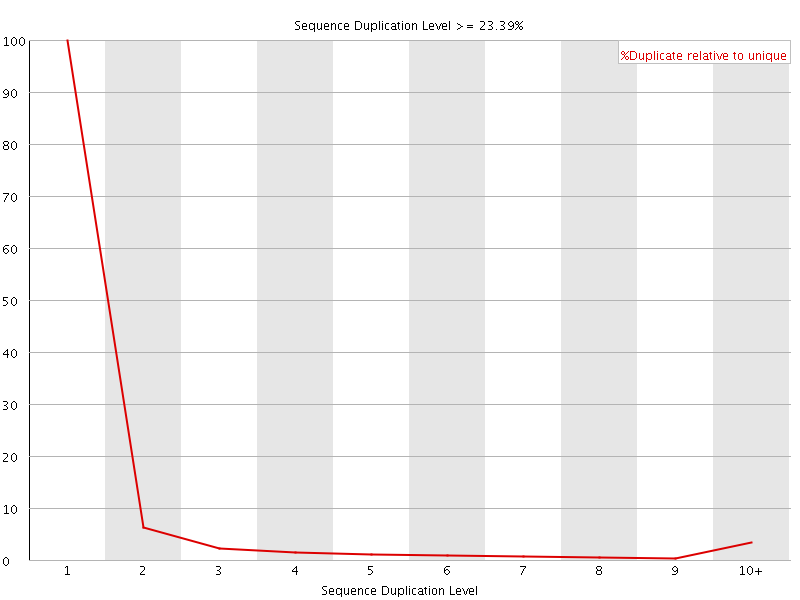
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
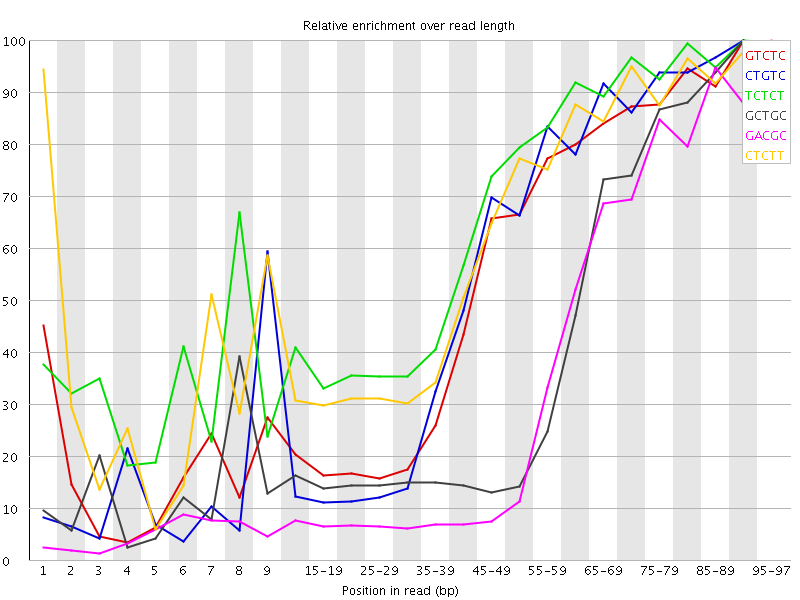
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 410320 | 3.582811 | 6.4352117 | 90-94 |
| CTGTC | 384115 | 3.3539956 | 5.990557 | 90-94 |
| TCTCT | 573810 | 3.3517845 | 5.00244 | 90-94 |
| GCTGC | 256750 | 3.351235 | 8.128152 | 90-94 |
| GACGC | 231365 | 3.0966263 | 8.493312 | 95-97 |
| CTCTT | 528785 | 3.0887809 | 4.9042497 | 95-97 |
| CGCTG | 228605 | 2.9838717 | 7.776393 | 95-97 |
| GCCGA | 217745 | 2.914334 | 8.23341 | 90-94 |
| CTGCC | 227370 | 2.8841894 | 7.7627125 | 90-94 |
| CGACG | 210860 | 2.822184 | 8.504131 | 95-97 |
| ACACA | 445940 | 2.8085015 | 6.7739754 | 6 |
| TGCCG | 206230 | 2.6918218 | 7.8420215 | 90-94 |
| CCGAC | 205830 | 2.6772938 | 7.61907 | 95-97 |
| GAAGG | 274755 | 2.6708403 | 6.1340475 | 95-97 |
| TACAC | 403305 | 2.4770515 | 8.148068 | 5 |
| CTGAC | 271365 | 2.4296951 | 6.052355 | 95-97 |
| TGACG | 257805 | 2.3751614 | 6.051944 | 95-97 |
| GACGA | 235485 | 2.2246504 | 6.269861 | 95-97 |
| GGCTT | 247160 | 2.220666 | 5.33968 | 90-94 |
| CGAAG | 222625 | 2.1031606 | 5.799431 | 95-97 |
| ACGCT | 233825 | 2.0935767 | 5.3984704 | 85-89 |
| AAGGC | 210250 | 1.9862528 | 5.1040635 | 90-94 |
| AGGCT | 213640 | 1.9682685 | 5.2653456 | 90-94 |
| CTTTG | 317105 | 1.9059647 | 7.6135015 | 9 |
| AGAGA | 280715 | 1.8718518 | 5.193389 | 8 |
| TATAC | 420445 | 1.7775477 | 5.5882325 | 5 |
| GGTGG | 127865 | 1.7670698 | 5.30878 | 95-97 |
| GAGAG | 179820 | 1.7479955 | 5.680455 | 7 |
| GTGTA | 217015 | 1.376265 | 9.471914 | 1 |
| AAGAC | 202675 | 1.3134158 | 5.680101 | 5 |
| CCATG | 144380 | 1.2927216 | 5.3233852 | 9 |
| GAGTC | 137420 | 1.2660524 | 6.3399177 | 9 |
| GTCTT | 202625 | 1.217881 | 5.9361567 | 1 |
| TATGA | 270920 | 1.1785743 | 5.410405 | 4 |
| GATTG | 184615 | 1.170791 | 5.3081956 | 7 |
| GTGTT | 188190 | 1.1638906 | 5.6371307 | 1 |
| GACTT | 188780 | 1.1634951 | 5.6279817 | 7 |
| TCATA | 271320 | 1.1470805 | 5.4037557 | 2 |
| GTTCT | 190545 | 1.1452739 | 5.0789714 | 1 |
| TGGAC | 112825 | 1.0394583 | 6.533487 | 5 |
| GTCCA | 115225 | 1.0316793 | 7.7570252 | 1 |
| AGACT | 152285 | 0.9624151 | 5.369369 | 6 |
| GGACT | 102050 | 0.94018817 | 5.8750477 | 6 |
| GTATA | 166720 | 0.72527647 | 6.809741 | 1 |