![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-160_CAGAGAGG-CTAAGCCT_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1531303 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
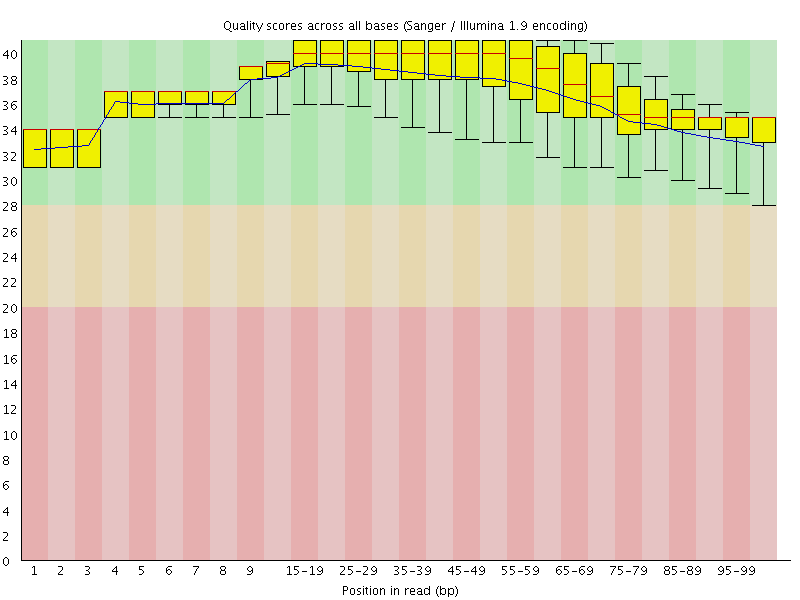
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
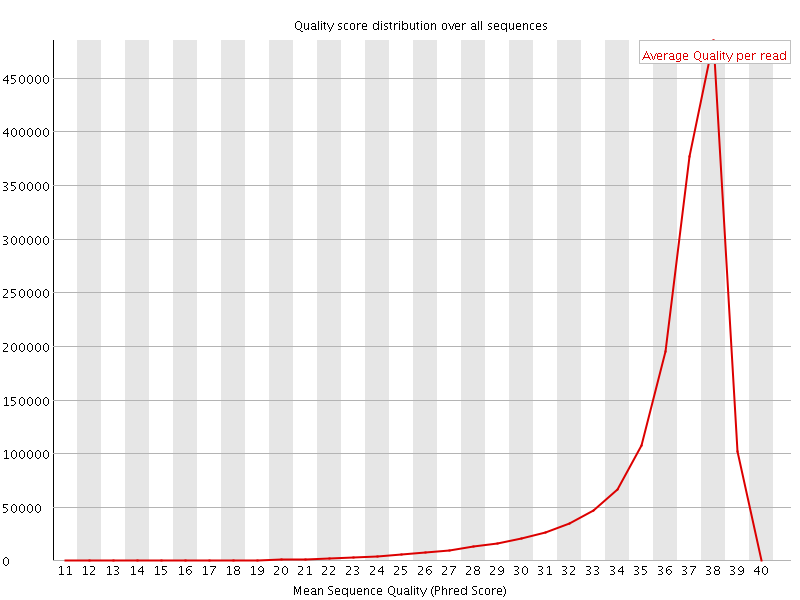
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
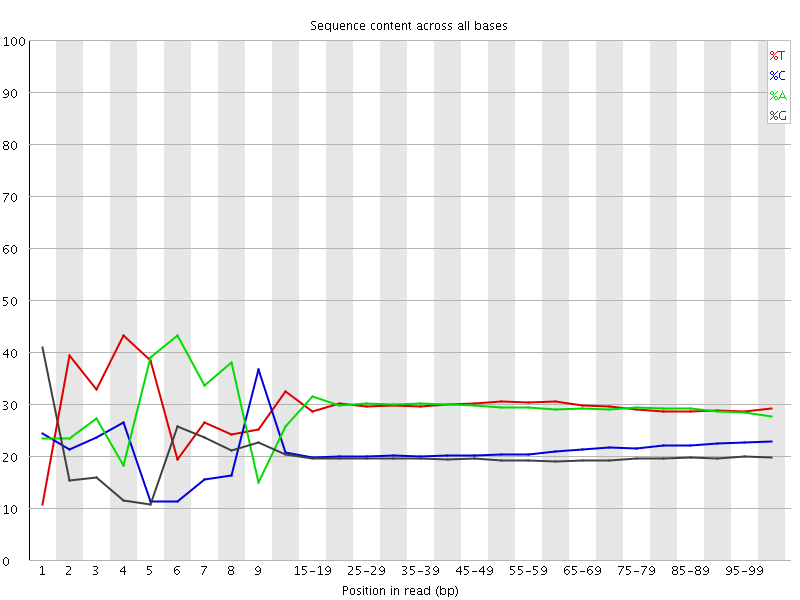
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
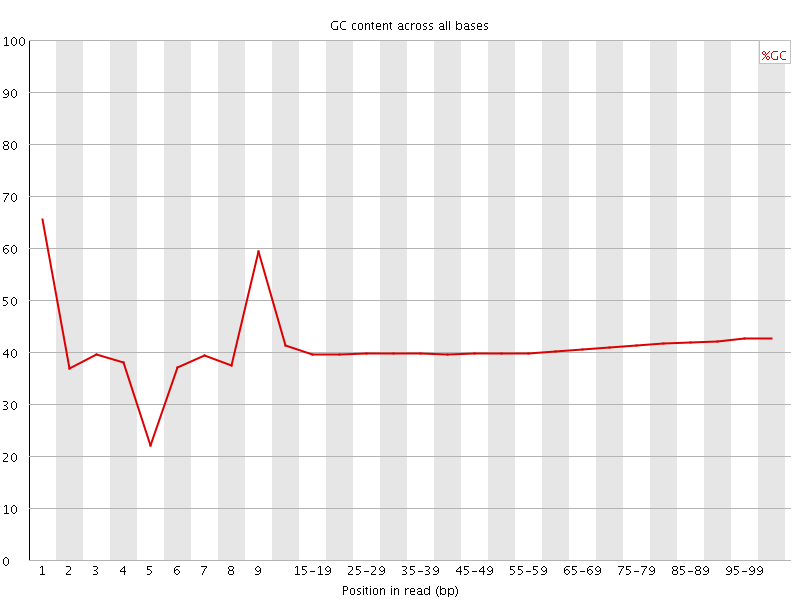
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
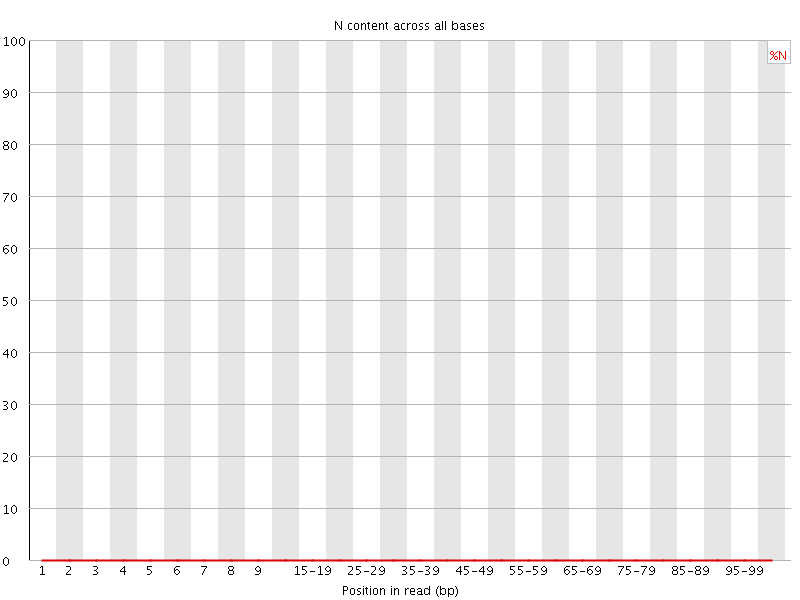
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
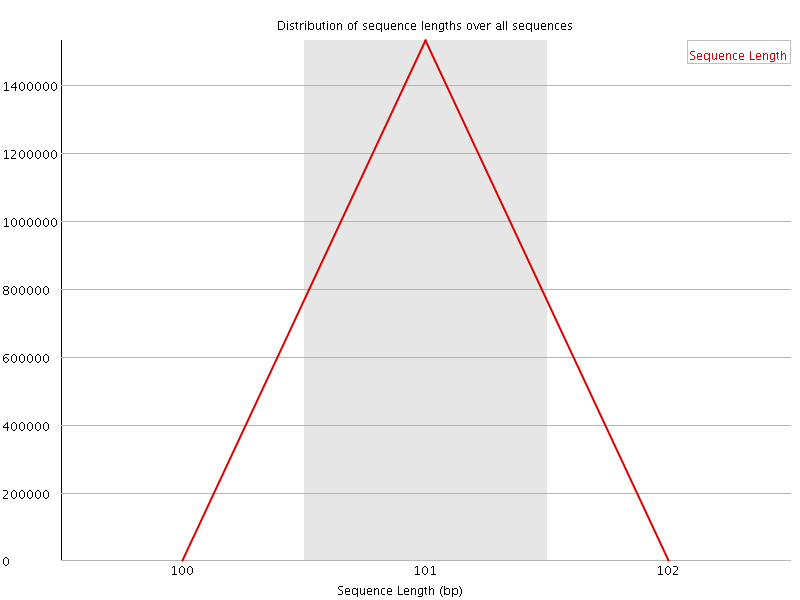
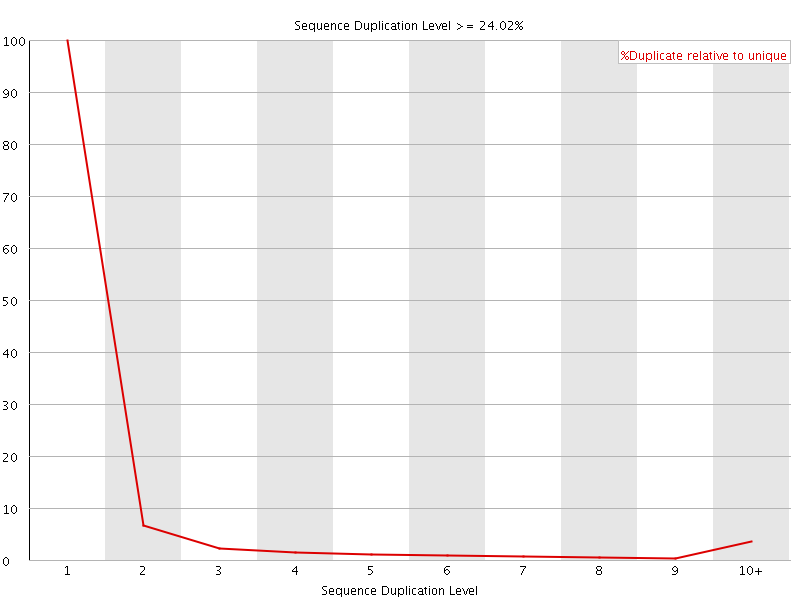
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
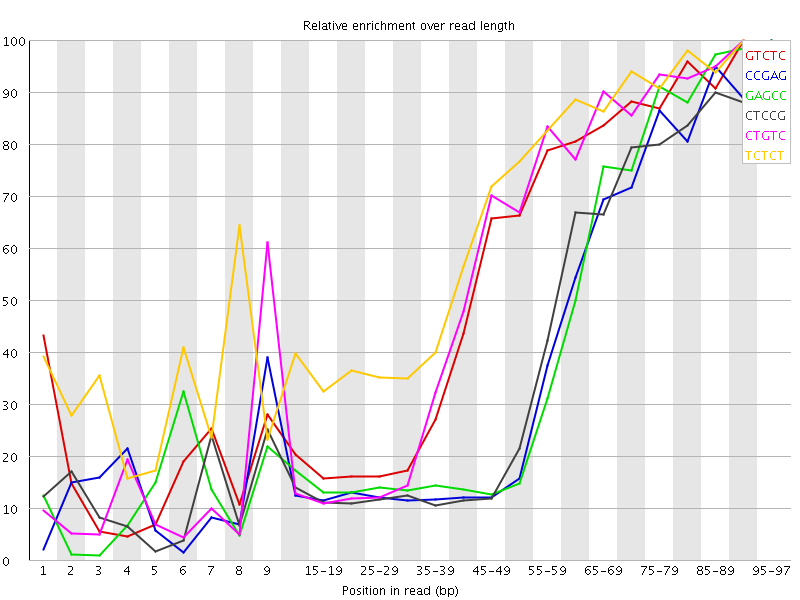
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 411555 | 3.5765522 | 6.4010043 | 90-94 |
| CCGAG | 265705 | 3.5166936 | 8.728042 | 95-97 |
| GAGCC | 257585 | 3.4092226 | 8.090552 | 95-97 |
| CTCCG | 276530 | 3.403762 | 8.296116 | 95-97 |
| CTGTC | 387165 | 3.3645947 | 6.0343895 | 90-94 |
| TCTCT | 576440 | 3.3284254 | 5.058811 | 90-94 |
| CGAGC | 248445 | 3.2882516 | 8.145743 | 85-89 |
| CTCTT | 530430 | 3.062759 | 4.813798 | 95-97 |
| TCTCC | 370195 | 3.0275793 | 6.1096554 | 95-97 |
| AGCCC | 239305 | 2.9806764 | 7.531529 | 90-94 |
| GCCCA | 238815 | 2.9745731 | 7.6305804 | 90-94 |
| TGTCT | 476880 | 2.9259412 | 5.029208 | 90-94 |
| ATCTC | 472500 | 2.760786 | 6.7099123 | 95-97 |
| GAGAG | 274750 | 2.7606428 | 6.372698 | 95-97 |
| ACACA | 446200 | 2.6696405 | 6.779631 | 6 |
| CCCAC | 227260 | 2.663878 | 7.2003403 | 90-94 |
| CCACG | 210595 | 2.6230774 | 7.443349 | 90-94 |
| TCCGA | 289780 | 2.548304 | 5.93845 | 95-97 |
| GAGGA | 252890 | 2.5409968 | 6.5091157 | 95-97 |
| TACAC | 405765 | 2.3991184 | 8.416999 | 5 |
| GAGAC | 248585 | 2.3505836 | 6.2280226 | 95-97 |
| CGAGA | 244580 | 2.3127127 | 6.26776 | 95-97 |
| AGAGG | 224530 | 2.2560403 | 5.9342093 | 95-97 |
| CAGAG | 237065 | 2.241652 | 5.7406197 | 90-94 |
| ACGAG | 230865 | 2.1830258 | 6.010997 | 95-97 |
| GACCA | 240535 | 2.140461 | 5.323165 | 95-97 |
| ACCAG | 223035 | 1.9847327 | 5.0572014 | 90-94 |
| CACGA | 221500 | 1.9710732 | 5.3490543 | 95-97 |
| CTTTG | 317380 | 1.9473143 | 8.002634 | 9 |
| GGATC | 208295 | 1.9464061 | 5.6351013 | 95-97 |
| AGACC | 217090 | 1.9318297 | 5.5245275 | 95-97 |
| CCAGA | 216920 | 1.9303169 | 5.244474 | 90-94 |
| TATAC | 420775 | 1.7564945 | 5.4548125 | 5 |
| TGGAG | 167135 | 1.6595632 | 5.1370635 | 5 |
| GCTTT | 267030 | 1.6383871 | 5.004432 | 1 |
| GTCTT | 218670 | 1.3416698 | 6.027931 | 1 |
| AAGAC | 203095 | 1.291202 | 5.678174 | 5 |
| GAGTC | 137585 | 1.2856587 | 5.740575 | 9 |
| CCATG | 145035 | 1.2754271 | 5.155055 | 9 |
| GTGTT | 188975 | 1.2320621 | 5.643374 | 1 |
| GATTG | 184745 | 1.2188412 | 5.607645 | 7 |
| TATGA | 270735 | 1.200917 | 5.542519 | 4 |
| GACTT | 189425 | 1.1760885 | 5.780003 | 7 |
| GTTCT | 190320 | 1.1677259 | 5.2662582 | 1 |
| TCATA | 273275 | 1.1407665 | 5.3643255 | 2 |
| GTGTA | 161745 | 1.0671003 | 9.690496 | 1 |
| TGGAC | 113005 | 1.0559717 | 6.905001 | 5 |
| GCCGT | 80110 | 1.0477927 | 5.6634774 | 95-97 |
| TGCCG | 76575 | 1.0015569 | 5.817801 | 95-97 |
| GTCCA | 113675 | 0.99964964 | 6.6993303 | 1 |
| AGACT | 151795 | 0.9536879 | 5.1695743 | 6 |
| GGACT | 101440 | 0.9479029 | 6.139289 | 6 |
| GTATA | 165095 | 0.73232275 | 6.8337364 | 1 |