![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-159_CAGAGAGG-AAGGAGTA_L005_R2_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1732499 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 40 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
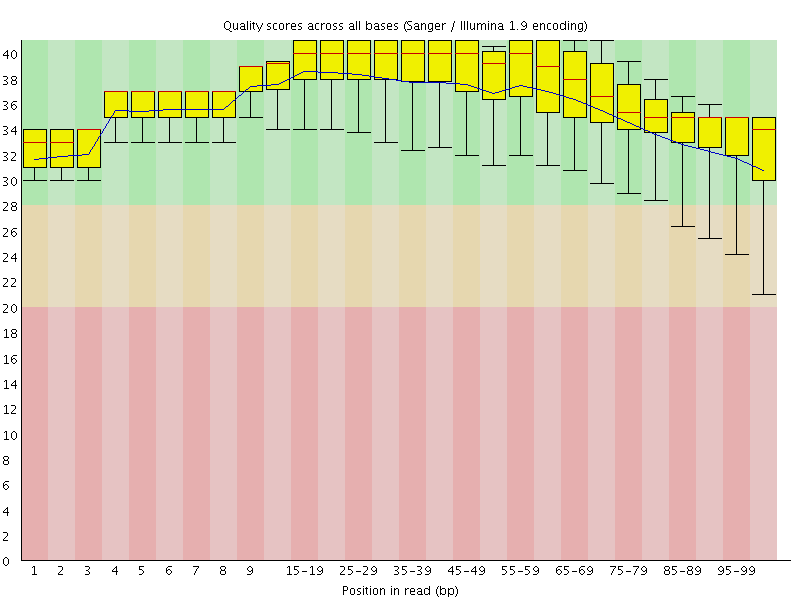
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
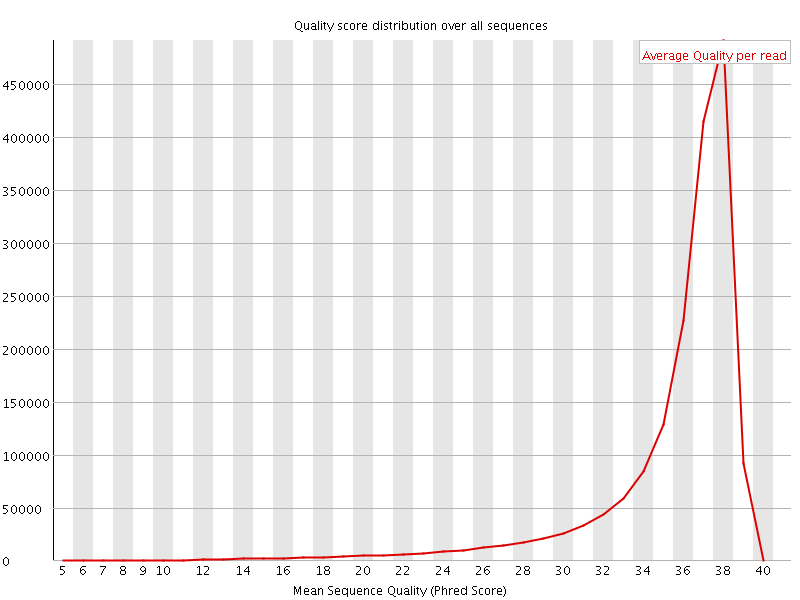
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
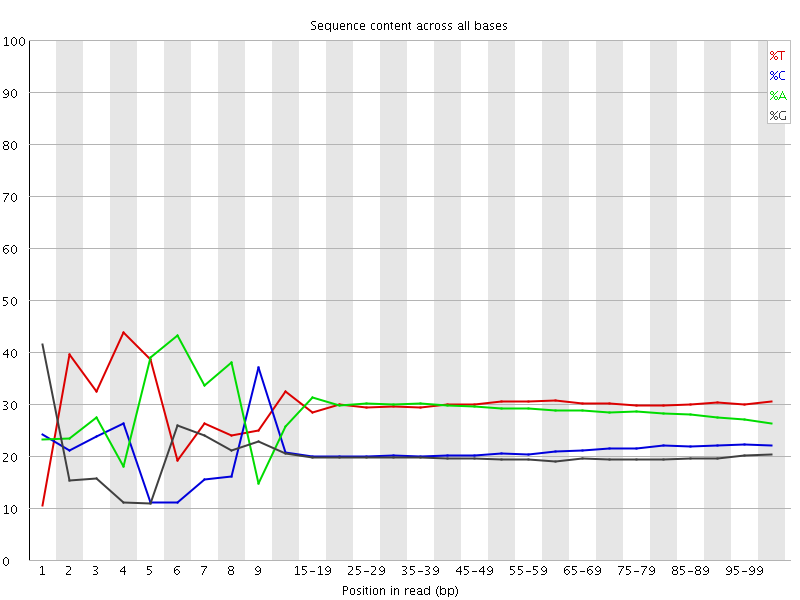
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
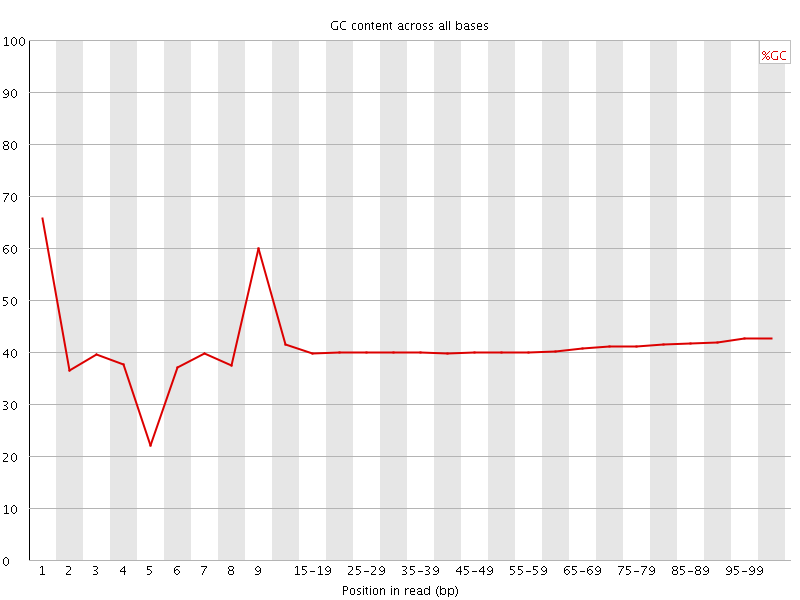
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
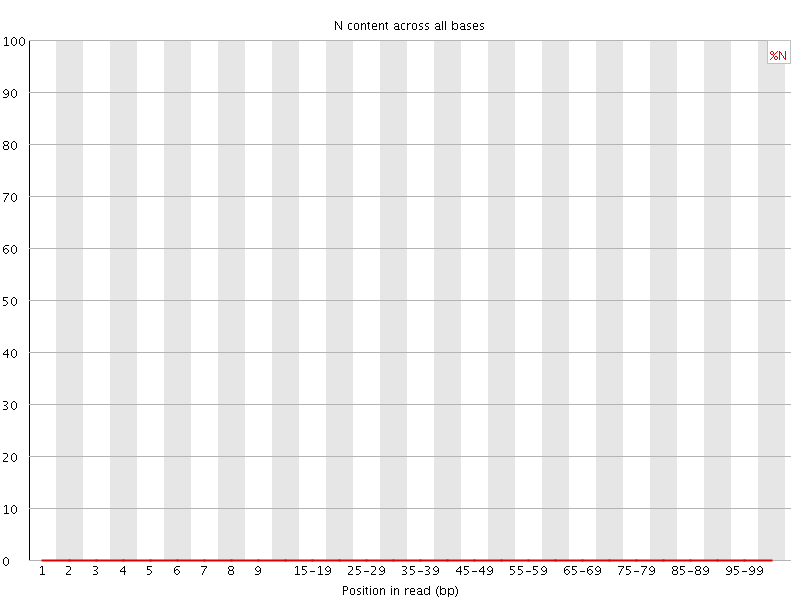
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
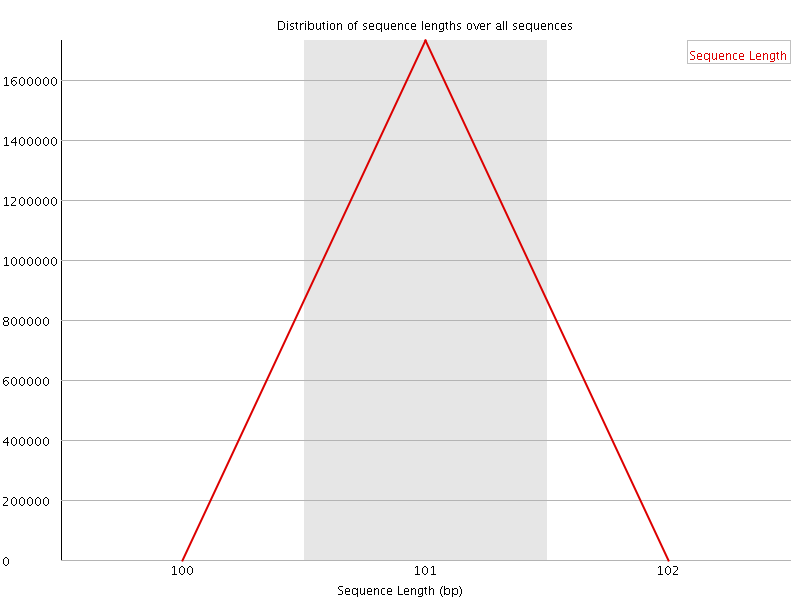
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
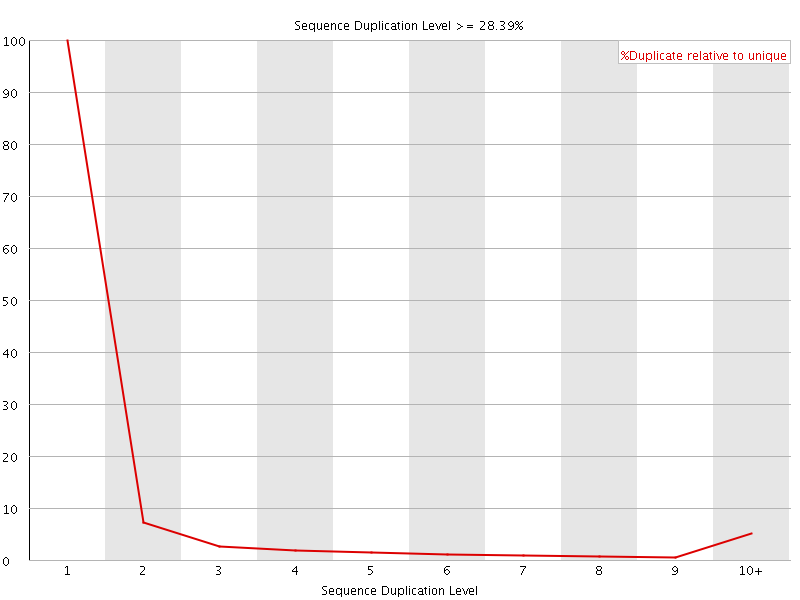
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
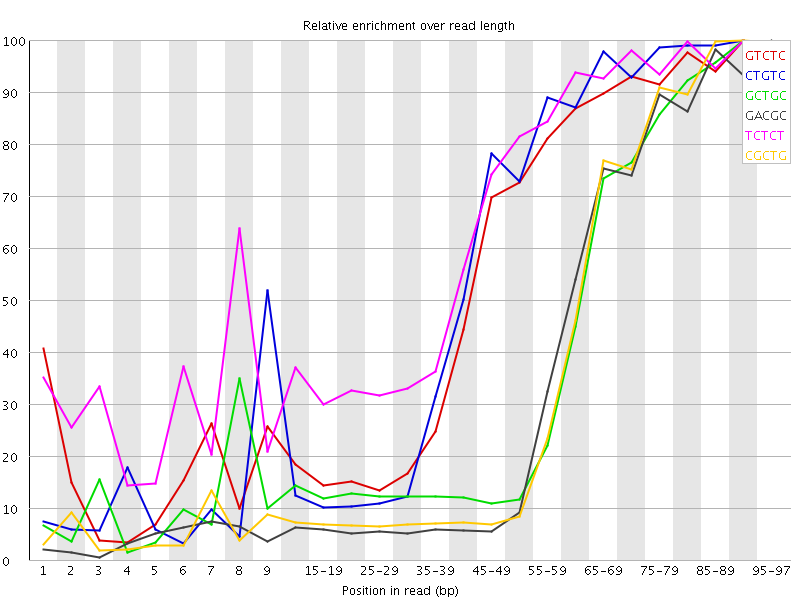
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 512655 | 3.8435378 | 6.714375 | 90-94 |
| CTGTC | 484355 | 3.6313636 | 6.206779 | 90-94 |
| GCTGC | 319605 | 3.6212046 | 9.011771 | 90-94 |
| GACGC | 295795 | 3.4716845 | 9.25223 | 95-97 |
| TCTCT | 691930 | 3.4326904 | 5.194116 | 90-94 |
| CGCTG | 291355 | 3.301125 | 8.686468 | 90-94 |
| GCCGA | 272030 | 3.1927595 | 9.274787 | 90-94 |
| CTCTT | 643505 | 3.1924522 | 4.9512925 | 80-84 |
| CTGCC | 288655 | 3.0977926 | 8.461886 | 90-94 |
| CGACG | 263820 | 3.0964005 | 9.343309 | 95-97 |
| TGTCT | 588560 | 3.0826864 | 5.117966 | 90-94 |
| ACACA | 554255 | 3.0564113 | 7.383334 | 6 |
| TGCCG | 262140 | 2.9701116 | 8.787576 | 90-94 |
| CCGAC | 260710 | 2.898284 | 8.576635 | 95-97 |
| TACAC | 511905 | 2.7250953 | 9.136309 | 5 |
| AGAAG | 442435 | 2.7194684 | 5.1463103 | 5 |
| CTGAC | 344595 | 2.6762388 | 6.517788 | 95-97 |
| TGACG | 325045 | 2.665174 | 6.614788 | 95-97 |
| CACAT | 494445 | 2.632148 | 5.1208563 | 95-97 |
| ATCTG | 474580 | 2.5748863 | 5.260366 | 95-97 |
| GACGA | 292585 | 2.485101 | 7.176203 | 95-97 |
| ACGCT | 301980 | 2.3452766 | 6.0844803 | 85-89 |
| CAAAG | 379230 | 2.2078576 | 5.2049227 | 4 |
| CCTTG | 293360 | 2.1994135 | 5.54052 | 95-97 |
| CTCCT | 300770 | 2.1358674 | 5.4656715 | 90-94 |
| ACTCC | 282575 | 2.0786598 | 5.391361 | 90-94 |
| CTTTG | 371345 | 1.9449848 | 8.153356 | 9 |
| TATAC | 522575 | 1.9434512 | 5.5383954 | 5 |
| AGAGA | 309235 | 1.900742 | 5.0375457 | 8 |
| GGTGG | 145195 | 1.8336798 | 5.9701123 | 95-97 |
| GAGAG | 201495 | 1.8068515 | 6.0488353 | 7 |
| GCTTT | 312145 | 1.6349144 | 5.1704836 | 1 |
| GTGTA | 275210 | 1.5764455 | 10.51463 | 1 |
| TTGAG | 258455 | 1.4804702 | 5.544572 | 9 |
| CTCGG | 122650 | 1.3896551 | 5.905652 | 95-97 |
| AAGAC | 237390 | 1.3820724 | 5.8686 | 5 |
| GAGTC | 165410 | 1.3562628 | 7.069791 | 9 |
| CCATG | 168155 | 1.3059474 | 5.551446 | 9 |
| TATGA | 320905 | 1.2599916 | 6.5138516 | 4 |
| AGAAC | 213775 | 1.2445871 | 5.111726 | 5 |
| GTCTT | 233615 | 1.2235997 | 5.947962 | 1 |
| GACTT | 222415 | 1.206737 | 6.3067126 | 7 |
| CGGTG | 100535 | 1.2026048 | 5.683797 | 95-97 |
| TCATA | 322980 | 1.2011594 | 6.205871 | 2 |
| GATTG | 209055 | 1.1974994 | 6.2083864 | 7 |
| GTGTT | 211990 | 1.1722499 | 5.7672853 | 1 |
| GTTCT | 217715 | 1.1403207 | 5.139994 | 1 |
| ACATG | 195955 | 1.1013234 | 5.2603364 | 8 |
| TGGAC | 131480 | 1.078057 | 7.346315 | 5 |
| GTCCA | 132385 | 1.0281458 | 7.263585 | 1 |
| AGACT | 176320 | 0.990969 | 5.399169 | 6 |
| GAGTA | 166255 | 0.98650527 | 5.5726037 | 1 |
| GGACT | 119575 | 0.9804433 | 6.45334 | 6 |
| ACACT | 182185 | 0.9698507 | 5.0313907 | 6 |
| GTATG | 164840 | 0.944229 | 5.579484 | 3 |
| GTATA | 189265 | 0.7431243 | 6.919659 | 1 |