![[OK]](Icons/tick.png) Basic Statistics
Basic Statistics
| Measure | Value |
|---|---|
| Filename | Nextera-159_CAGAGAGG-AAGGAGTA_L005_R1_001.fastq.gz |
| File type | Conventional base calls |
| Encoding | Sanger / Illumina 1.9 |
| Total Sequences | 1732499 |
| Filtered Sequences | 0 |
| Sequence length | 101 |
| %GC | 41 |
![[OK]](Icons/tick.png) Per base sequence quality
Per base sequence quality
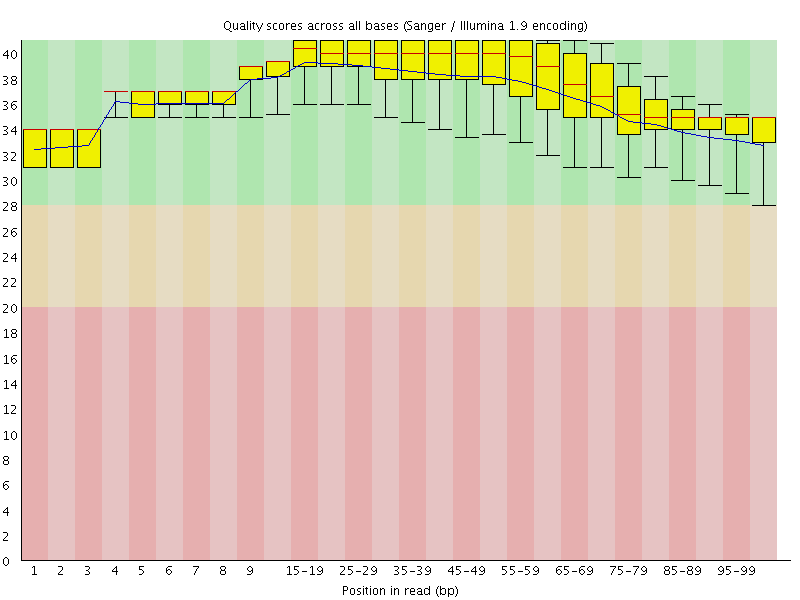
![[OK]](Icons/tick.png) Per sequence quality scores
Per sequence quality scores
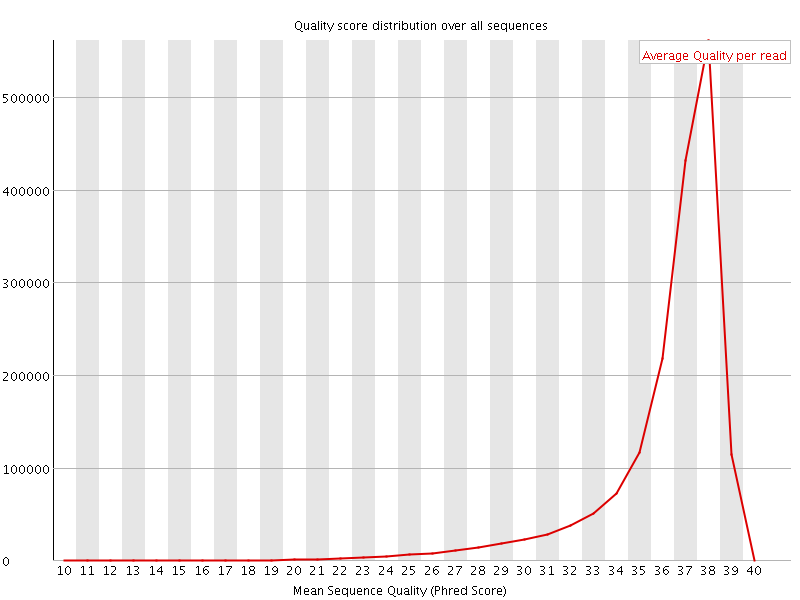
![[FAIL]](Icons/error.png) Per base sequence content
Per base sequence content
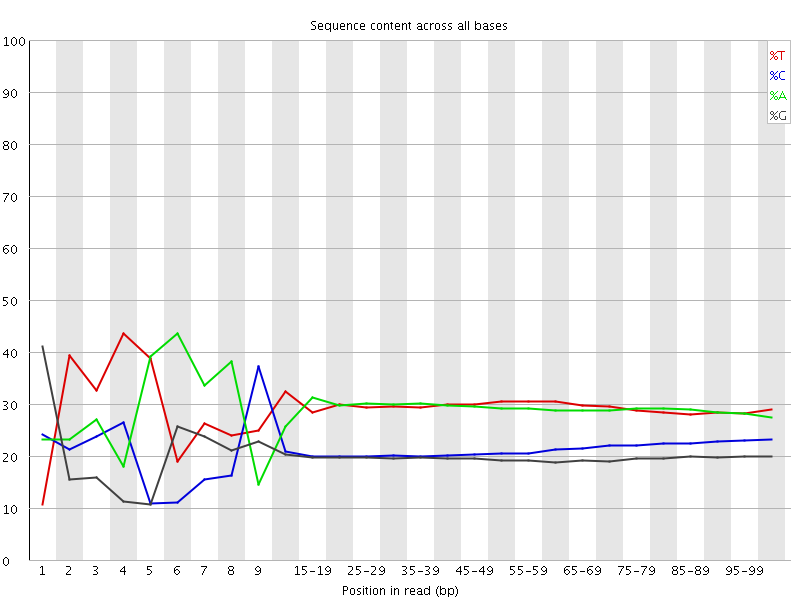
![[FAIL]](Icons/error.png) Per base GC content
Per base GC content
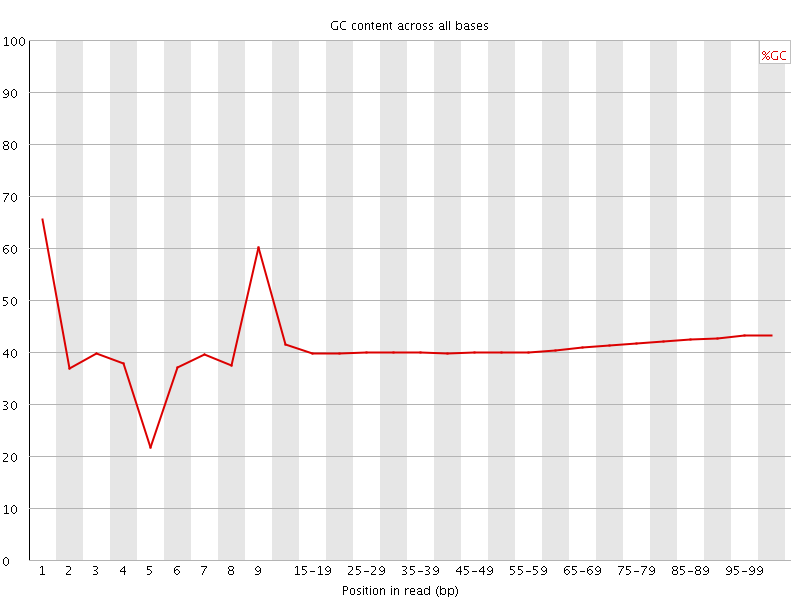
![[WARN]](Icons/warning.png) Per sequence GC content
Per sequence GC content
![[OK]](Icons/tick.png) Per base N content
Per base N content
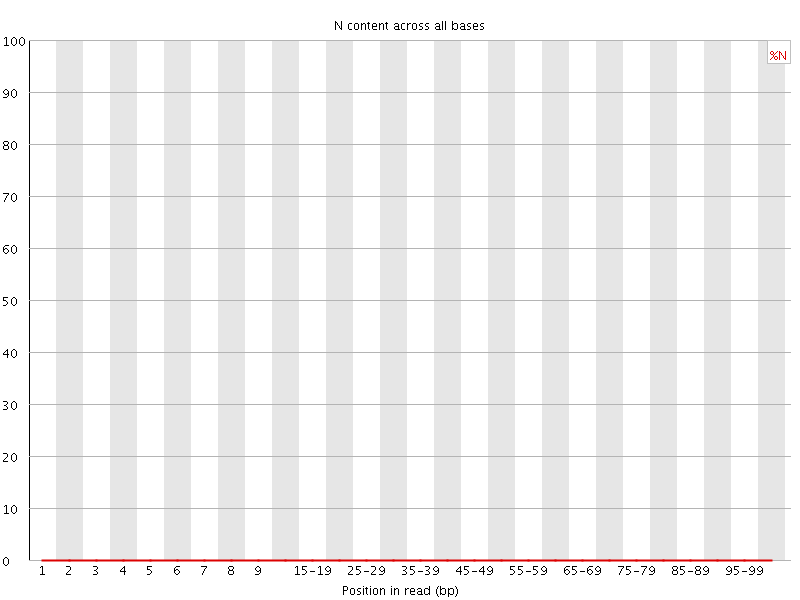
![[OK]](Icons/tick.png) Sequence Length Distribution
Sequence Length Distribution
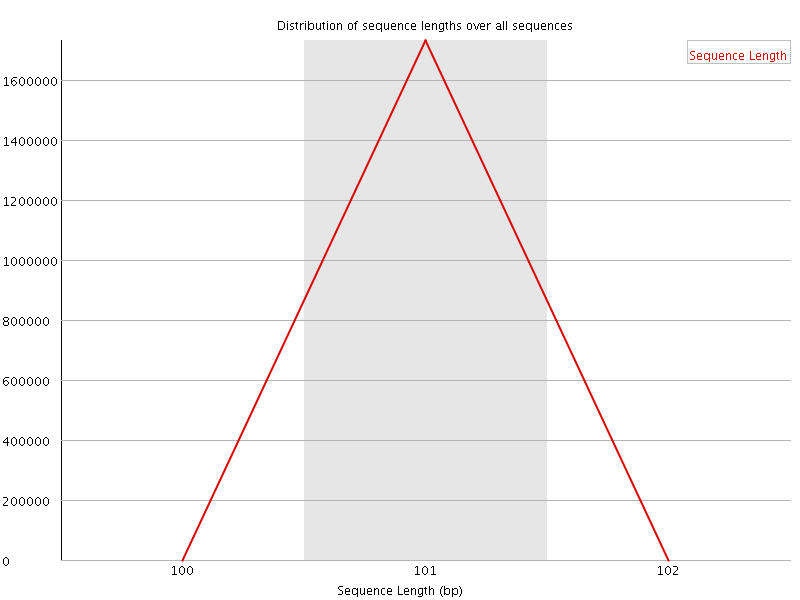
![[WARN]](Icons/warning.png) Sequence Duplication Levels
Sequence Duplication Levels
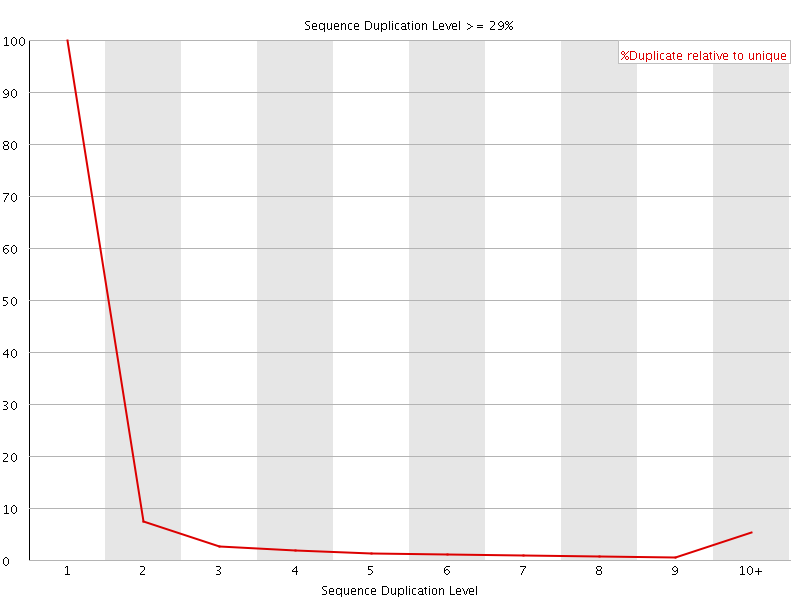
![[OK]](Icons/tick.png) Overrepresented sequences
Overrepresented sequences
No overrepresented sequences
![[WARN]](Icons/warning.png) Kmer Content
Kmer Content
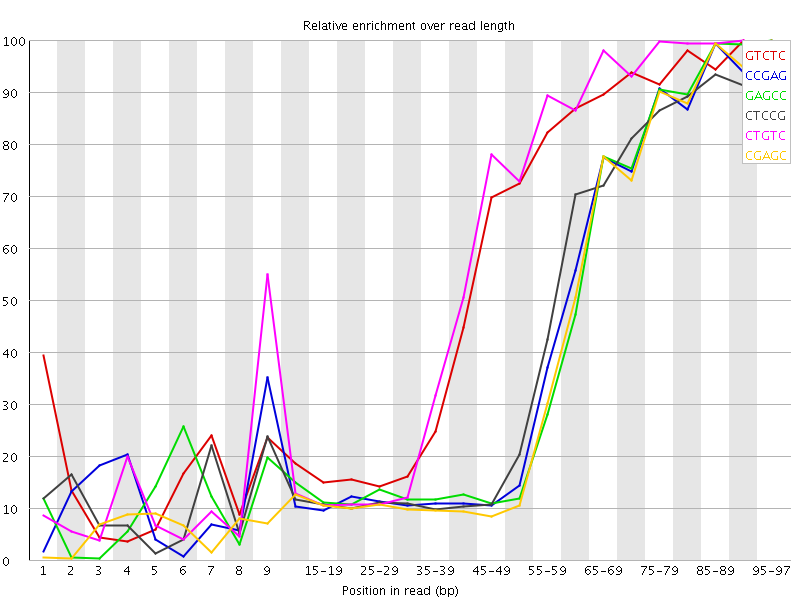
| Sequence | Count | Obs/Exp Overall | Obs/Exp Max | Max Obs/Exp Position |
|---|---|---|---|---|
| GTCTC | 514480 | 3.8886364 | 6.779349 | 90-94 |
| CCGAG | 337645 | 3.8526318 | 9.338907 | 95-97 |
| GAGCC | 327860 | 3.740982 | 9.09732 | 95-97 |
| CTCCG | 352430 | 3.709722 | 8.839806 | 95-97 |
| CTGTC | 485750 | 3.671484 | 6.256733 | 90-94 |
| CGAGC | 314725 | 3.5911078 | 9.078878 | 95-97 |
| TCTCT | 697155 | 3.532445 | 5.351533 | 90-94 |
| AGCCC | 308450 | 3.2857828 | 8.360418 | 90-94 |
| CTCTT | 646245 | 3.274487 | 5.108304 | 90-94 |
| GCCCA | 306040 | 3.2601101 | 8.526738 | 90-94 |
| TCTCC | 457150 | 3.2258518 | 6.6262364 | 95-97 |
| TGTCT | 589330 | 3.1985102 | 5.3064036 | 90-94 |
| ATCTC | 585820 | 3.0039716 | 7.555115 | 95-97 |
| GAGAG | 330245 | 2.9330711 | 7.1461453 | 90-94 |
| CCCAC | 293215 | 2.916064 | 7.9855647 | 90-94 |
| ACACA | 554720 | 2.9132426 | 6.992133 | 6 |
| CCACG | 267390 | 2.848389 | 8.269511 | 90-94 |
| TCCGA | 371155 | 2.8390284 | 6.411388 | 95-97 |
| GAGGA | 305450 | 2.7128544 | 7.249079 | 95-97 |
| AGAGA | 445655 | 2.6852696 | 5.471023 | 90-94 |
| AGAAG | 444835 | 2.6803288 | 5.1563234 | 5 |
| TACAC | 513615 | 2.6653543 | 9.012579 | 5 |
| GAGAC | 308695 | 2.5596042 | 7.0692086 | 95-97 |
| CGAGA | 305450 | 2.5326974 | 7.0759087 | 95-97 |
| AGAGG | 272820 | 2.423051 | 6.9792128 | 95-97 |
| ACGAG | 290610 | 2.4096487 | 6.7850995 | 95-97 |
| CAGAG | 288325 | 2.390702 | 6.514781 | 90-94 |
| GACCA | 294515 | 2.2798562 | 6.1530957 | 95-97 |
| CACGA | 280720 | 2.1730683 | 6.1118083 | 95-97 |
| ACCAG | 277505 | 2.1481807 | 5.827657 | 90-94 |
| CAAAG | 379635 | 2.1355624 | 5.223013 | 4 |
| AGACC | 272055 | 2.105992 | 6.306986 | 95-97 |
| GGATC | 255790 | 2.095758 | 6.2927303 | 95-97 |
| CCAGA | 267410 | 2.070035 | 5.9004755 | 90-94 |
| CTTTG | 371710 | 2.0174067 | 8.31539 | 9 |
| TATAC | 525310 | 1.9574617 | 5.8247724 | 5 |
| GCTTT | 313165 | 1.6996614 | 5.3556395 | 1 |
| TTGAG | 257755 | 1.5164393 | 5.585177 | 9 |
| GTCTT | 254920 | 1.3835444 | 6.3057065 | 1 |
| GAGTC | 164970 | 1.3516446 | 6.5784597 | 9 |
| AAGAC | 237025 | 1.3333375 | 5.978508 | 5 |
| GTATG | 226070 | 1.3300283 | 5.7962613 | 3 |
| GACTC | 172275 | 1.3177612 | 5.2774687 | 7 |
| CCATG | 169645 | 1.297644 | 5.544495 | 9 |
| TATGA | 319730 | 1.2761571 | 6.419076 | 4 |
| GATTG | 211290 | 1.2430737 | 6.4295144 | 7 |
| GTGTT | 212370 | 1.234598 | 5.7760644 | 1 |
| GACTT | 223225 | 1.2260766 | 6.295476 | 7 |
| TCATA | 323690 | 1.2061654 | 6.188649 | 2 |
| GTTCT | 217335 | 1.1795568 | 5.181943 | 1 |
| ACTCA | 220495 | 1.1442369 | 5.0019565 | 8 |
| GTGTA | 192590 | 1.1330568 | 10.775133 | 1 |
| ACATG | 195960 | 1.0892508 | 5.5194616 | 8 |
| TGGAC | 132040 | 1.0818402 | 7.22995 | 5 |
| GCCGT | 92870 | 1.0470973 | 6.365248 | 95-97 |
| CCGTC | 97840 | 1.029876 | 5.1208267 | 95-97 |
| GTCCA | 131645 | 1.0069753 | 6.976875 | 1 |
| GGACT | 121090 | 0.9921237 | 6.48312 | 6 |
| GAGTA | 166510 | 0.99138844 | 5.410418 | 1 |
| TGCCG | 87880 | 0.9908357 | 6.7861967 | 95-97 |
| AGACT | 175440 | 0.9751895 | 5.6380434 | 6 |
| ACACT | 183320 | 0.95132107 | 5.0422144 | 6 |
| GGATA | 148870 | 0.8863612 | 5.08995 | 1 |
| GTATA | 189115 | 0.7548258 | 7.0333686 | 1 |